|
Subtractive DNA Hybridization
Subtractive hybridization is a technology that allows for PCR-based amplification of only cDNA fragments that differ between a control (driver) and experimental transcriptome. cDNA is produced from mRNA. Differences in relative abundance of transcripts are highlighted, as are genetic differences between species. The technique relies on the removal of dsDNA formed by hybridization between a control and test sample, thus eliminating cDNAs or gDNAs of similar abundance, and retaining differentially expressed, or variable in sequence, transcripts or genomic sequences. Suppression subtractive hybridization has also been successfully used to identify strain- or species-specific DNA sequences in a variety of bacteria including ''Vibrio'' species (Metagenomics). See also * Representational difference analysis Representational difference analysis (RDA) is a technique used in biological research to find sequence differences in two genomic or cDNA samples. Genomes or cDNA sequences from ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polymerase Chain Reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) to a large enough amount to study in detail. PCR was invented in 1983 by the American biochemist Kary Mullis at Cetus Corporation; Mullis and biochemist Michael Smith (chemist), Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993. PCR is fundamental to many of the procedures used in genetic testing and research, including analysis of Ancient DNA, ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory research for a broad variety of applications ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Complementary DNA
In genetics, complementary DNA (cDNA) is DNA synthesized from a single-stranded RNA (e.g., messenger RNA (mRNA) or microRNA (miRNA)) template in a reaction catalyzed by the enzyme reverse transcriptase. cDNA is often used to express a specific protein in a cell that does not normally express that protein (i.e., heterologous expression), or to sequence or quantify mRNA molecules using DNA based methods (qPCR, RNA-seq). cDNA that codes for a specific protein can be transferred to a recipient cell for expression, often bacterial or yeast expression systems. cDNA is also generated to analyze transcriptomic profiles in bulk tissue, single cells, or single nuclei in assays such as microarrays, qPCR, and RNA-seq. cDNA is also produced naturally by retroviruses (such as HIV-1, HIV-2, simian immunodeficiency virus, etc.) and then integrated into the host's genome, where it creates a provirus. The term ''cDNA'' is also used, typically in a bioinformatics context, to refer to a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transcriptome
The transcriptome is the set of all RNA transcripts, including coding and non-coding, in an individual or a population of cells. The term can also sometimes be used to refer to all RNAs, or just mRNA, depending on the particular experiment. The term ''transcriptome'' is a portmanteau of the words ''transcript'' and ''genome''; it is associated with the process of transcript production during the biological process of transcription. The early stages of transcriptome annotations began with cDNA libraries published in the 1980s. Subsequently, the advent of high-throughput technology led to faster and more efficient ways of obtaining data about the transcriptome. Two biological techniques are used to study the transcriptome, namely DNA microarray, a hybridization-based technique and RNA-seq, a sequence-based approach. RNA-seq is the preferred method and has been the dominant transcriptomics technique since the 2010s. Single-cell transcriptomics allows tracking of transcript changes ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Messenger RNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein. mRNA is created during the process of transcription, where an enzyme (RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and, utilising amino acids carried by transfer RNA (tRNA), the ribosome creates the protein. This process is known as translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genetic information in a biological system. As in DNA, genetic inf ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
GDNA
Genomic deoxyribonucleic acid (abbreviated as gDNA) is chromosomal DNA, in contrast to extra-chromosomal DNAs like plasmids. Most organisms have the same genomic DNA in every cell; however, only certain genes are active in each cell to allow for cell function and differentiation within the body. The genome of an organism (encoded by the genomic DNA) is the (biological) information of heredity which is passed from one generation of organism to the next. That genome is transcribed to produce various RNAs, which are necessary for the function of the organism. Precursor mRNA (pre-mRNA) is transcribed by RNA polymerase II in the nucleus. pre-mRNA is then processed by splicing to remove introns, leaving the exons in the mature messenger RNA (mRNA). Additional processing includes the addition of a 5' cap and a poly(A) tail to the pre-mRNA. The mature mRNA may then be transported to the cytosol and translated by the ribosome into a protein. Other types of RNA include ribosomal RNA (rRN ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
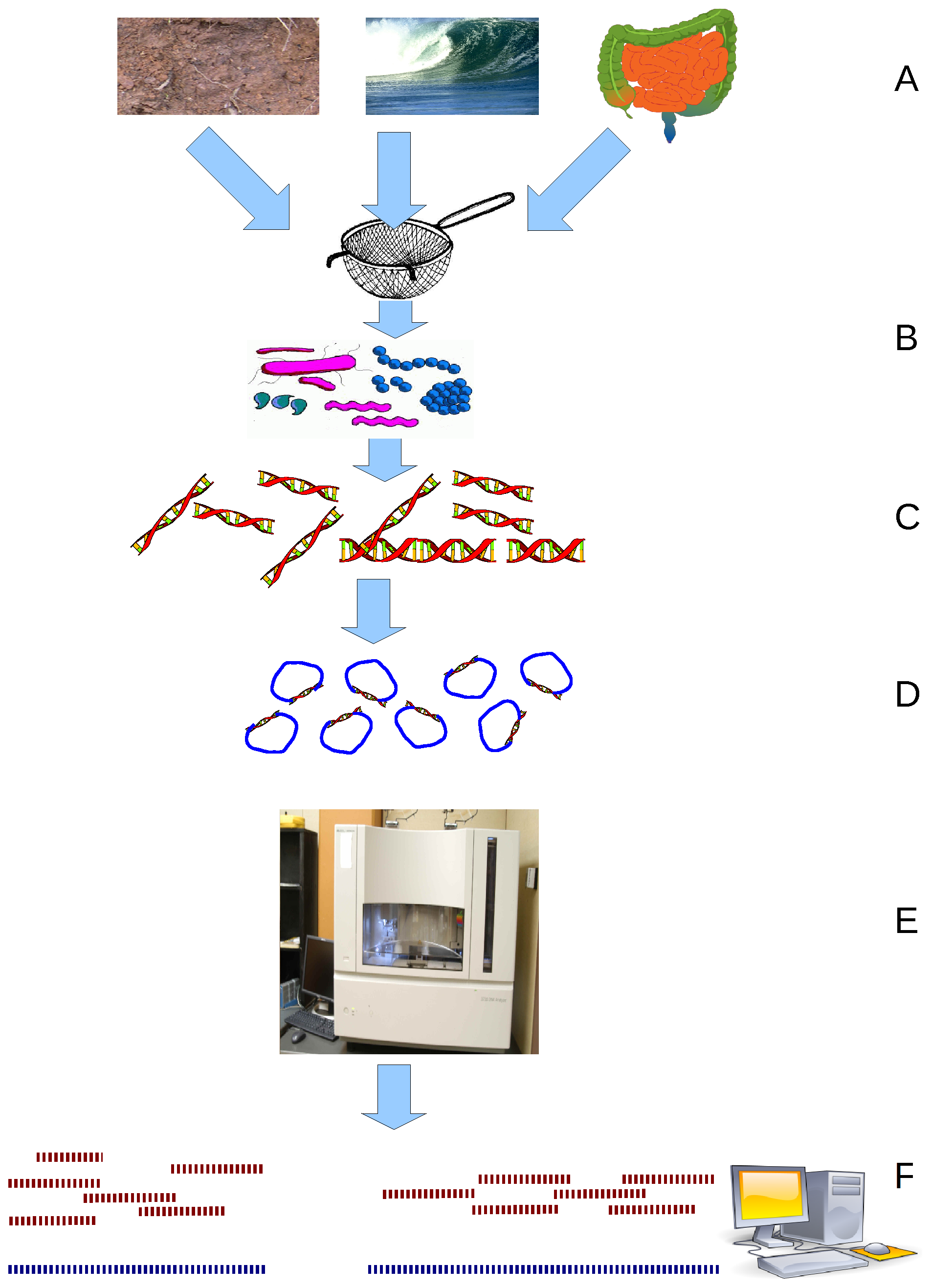
Metagenomics
Metagenomics is the study of genetic material recovered directly from environmental or clinical samples by a method called sequencing. The broad field may also be referred to as environmental genomics, ecogenomics, community genomics or microbiomics. While traditional microbiology and microbial genome sequencing and genomics rely upon cultivated clonal cultures, early environmental gene sequencing cloned specific genes (often the 16S rRNA gene) to produce a profile of diversity in a natural sample. Such work revealed that the vast majority of microbial biodiversity had been missed by cultivation-based methods. Because of its ability to reveal the previously hidden diversity of microscopic life, metagenomics offers a powerful lens for viewing the microbial world that has the potential to revolutionize understanding of the entire living world. As the price of DNA sequencing continues to fall, metagenomics now allows microbial ecology to be investigated at a much greater scale ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Representational Difference Analysis
Representational difference analysis (RDA) is a technique used in biological research to find sequence differences in two genomic or cDNA samples. Genomes or cDNA sequences from two samples (i.e. cancer sample and a normal sample) are PCR amplified and differences analyzed using subtractive DNA hybridization. This technology has been further enhanced through the development of representation oligonucleotide microarray analysis (ROMA), which uses array An array is a systematic arrangement of similar objects, usually in rows and columns. Things called an array include: {{TOC right Music * In twelve-tone and serial composition, the presentation of simultaneous twelve-tone sets such that the ... technology to perform such analyses. This method may also be adapted to detect DNA methylation differences, as seen in methylation-sensitive representational difference analysis (MS-RDA). Theory This method relies on PCR to differentially amplify non-homologous DNA regions between ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |