|
Ribosomal Protein L10 Leader
This family is a putative ribosomal protein leader autoregulatory structure found in ''B. subtilis'' and other low-GC Gram-positive bacteria. It is located in the 5′ untranslated regions of mRNAs encoding ribosomal proteins L10 and L12 (rplJ-rplL). A Rho Rho (uppercase Ρ, lowercase ρ or ; el, ρο or el, ρω, label=none) is the 17th letter of the Greek alphabet. In the system of Greek numerals it has a value of 100. It is derived from Phoenician letter res . Its uppercase form uses the sa ...-independent transcription terminator structure that is probably involved in regulation is included at the 3′ end. Other ribosomal protein leaders identified in the same study include those of L13, L19, L20 and L21. References External links * Ribosomal protein leader {{molecular-cell-biology-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Secondary Structure
Protein secondary structure is the three dimensional conformational isomerism, form of ''local segments'' of proteins. The two most common Protein structure#Secondary structure, secondary structural elements are alpha helix, alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary structure elements typically spontaneously form as an intermediate before the protein protein folding, folds into its three dimensional protein tertiary structure, tertiary structure. Secondary structure is formally defined by the pattern of hydrogen bonds between the Amine, amino hydrogen and carboxyl oxygen atoms in the peptide backbone chain, backbone. Secondary structure may alternatively be defined based on the regular pattern of backbone Dihedral angle#Dihedral angles of proteins, dihedral angles in a particular region of the Ramachandran plot regardless of whether it has the correct hydrogen bonds. The concept of secondary structure was first introduced by Kaj Ulrik ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequence Conservation
In evolutionary biology, conserved sequences are identical or similar sequences in nucleic acids ( DNA and RNA) or proteins across species ( orthologous sequences), or within a genome ( paralogous sequences), or between donor and receptor taxa ( xenologous sequences). Conservation indicates that a sequence has been maintained by natural selection. A highly conserved sequence is one that has remained relatively unchanged far back up the phylogenetic tree, and hence far back in geological time. Examples of highly conserved sequences include the RNA components of ribosomes present in all domains of life, the homeobox sequences widespread amongst Eukaryotes, and the tmRNA in Bacteria. The study of sequence conservation overlaps with the fields of genomics, proteomics, evolutionary biology, phylogenetics, bioinformatics and mathematics. History The discovery of the role of DNA in heredity, and observations by Frederick Sanger of variation between animal insulins in 1949, promp ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cis-regulatory Element
''Cis''-regulatory elements (CREs) or ''Cis''-regulatory modules (CRMs) are regions of non-coding DNA which regulate the transcription of neighboring genes. CREs are vital components of genetic regulatory networks, which in turn control morphogenesis, the development of anatomy, and other aspects of embryonic development, studied in evolutionary developmental biology. CREs are found in the vicinity of the genes that they regulate. CREs typically regulate gene transcription by binding to transcription factors. A single transcription factor may bind to many CREs, and hence control the expression of many genes ( pleiotropy). The Latin prefix ''cis'' means "on this side", i.e. on the same molecule of DNA as the gene(s) to be transcribed. CRMs are stretches of DNA, usually 100–1000 DNA base pairs in length, where a number of transcription factors can bind and regulate expression of nearby genes and regulate their transcription rates. They are labeled as ''cis'' because they are ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among the first life forms to appear on Earth, and are present in most of its habitats. Bacteria inhabit soil, water, acidic hot springs, radioactive waste, and the deep biosphere of Earth's crust. Bacteria are vital in many stages of the nutrient cycle by recycling nutrients such as the fixation of nitrogen from the atmosphere. The nutrient cycle includes the decomposition of dead bodies; bacteria are responsible for the putrefaction stage in this process. In the biological communities surrounding hydrothermal vents and cold seeps, extremophile bacteria provide the nutrients needed to sustain life by converting dissolved compounds, such as hydrogen sulphide and methane, to energy. Bacteria also live in symbiotic and parasitic relationsh ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribosomal Protein Leader
A ribosomal protein leader is a mechanism used in cells to control the cellular concentration of a protein that forms a part of the ribosome, and to make sure that the concentration is neither too high nor too low. Ribosomal protein leaders are RNA sequences that are a part of the 5' UTR of mRNAs encoding a ribosomal protein. When cellular concentrations of the ribosomal protein are high, excess protein will bind to the mRNA leader. This binding event can lower gene expression via a number of mechanisms; for example, in the protein-bound state, the RNA could form an intrinsic transcription termination stem-loop. When cellular concentrations of the ribosomal protein are not high, they are occupied in the ribosome, and are not available in significant quantities to bind the mRNA leader. This leads to increased expression of the gene, which leads to the synthesis of more copies of the ribosomal protein. Many examples of ribosomal protein leaders are known in bacteria, including r ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gram-positive
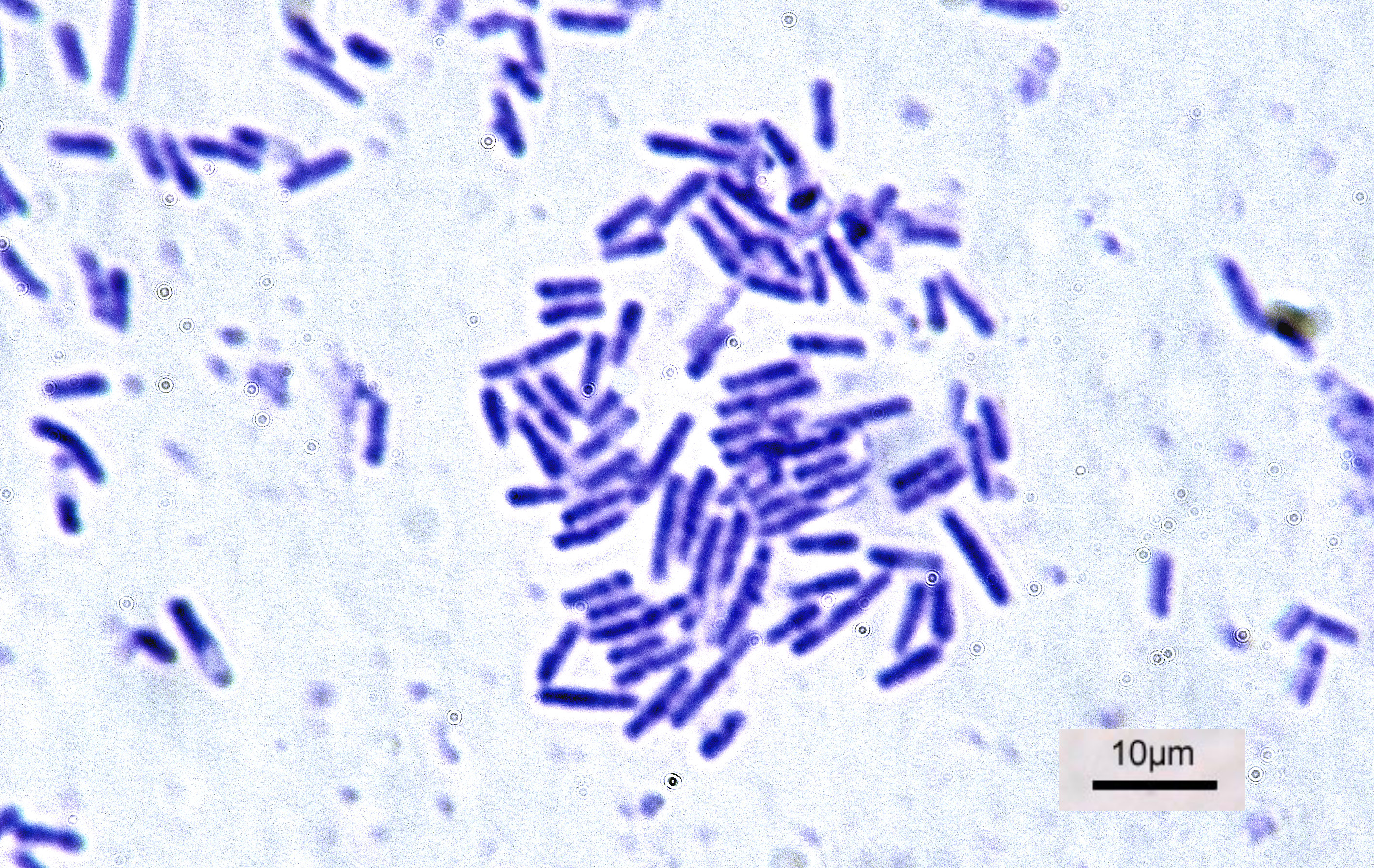
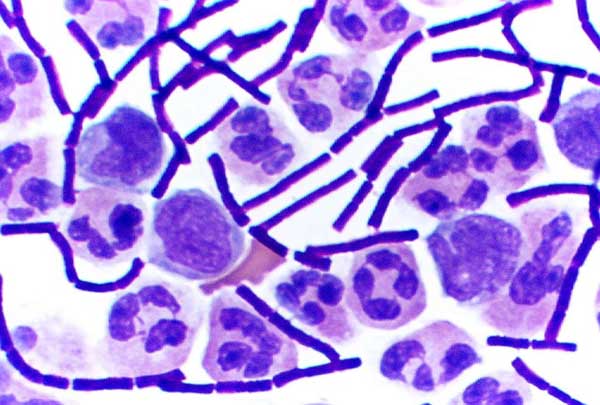
In bacteriology, gram-positive bacteria are bacteria that give a positive result in the Gram stain test, which is traditionally used to quickly classify bacteria into two broad categories according to their type of cell wall. Gram-positive bacteria take up the crystal violet stain used in the test, and then appear to be purple-coloured when seen through an optical microscope. This is because the thick peptidoglycan layer in the bacterial cell wall retains the stain after it is washed away from the rest of the sample, in the decolorization stage of the test. Conversely, gram-negative bacteria cannot retain the violet stain after the decolorization step; alcohol used in this stage degrades the outer membrane of gram-negative cells, making the cell wall more porous and incapable of retaining the crystal violet stain. Their peptidoglycan layer is much thinner and sandwiched between an inner cell membrane and a bacterial outer membrane, causing them to take up the counterstain (saf ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among the first life forms to appear on Earth, and are present in most of its habitats. Bacteria inhabit soil, water, acidic hot springs, radioactive waste, and the deep biosphere of Earth's crust. Bacteria are vital in many stages of the nutrient cycle by recycling nutrients such as the fixation of nitrogen from the atmosphere. The nutrient cycle includes the decomposition of dead bodies; bacteria are responsible for the putrefaction stage in this process. In the biological communities surrounding hydrothermal vents and cold seeps, extremophile bacteria provide the nutrients needed to sustain life by converting dissolved compounds, such as hydrogen sulphide and methane, to energy. Bacteria also live in symbiotic and parasitic relationsh ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein. mRNA is created during the process of Transcription (biology), transcription, where an enzyme (RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and, utilising amino acids carried by transfer RNA (tRNA), the ribosome creates the protein. This process is known as Translation (biology), translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genet ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribosomal Proteins
A ribosomal protein (r-protein or rProtein) is any of the proteins that, in conjunction with rRNA, make up the ribosomal subunits involved in the cellular process of translation. ''E. coli'', other bacteria and Archaea have a 30S small subunit and a 50S large subunit, whereas humans and yeasts have a 40S small subunit and a 60S large subunit. Equivalent subunits are frequently numbered differently between bacteria, Archaea, yeasts and humans. A large part of the knowledge about these organic molecules has come from the study of '' E. coli'' ribosomes. All ribosomal proteins have been isolated and many specific antibodies have been produced. These, together with electronic microscopy and the use of certain reactives, have allowed for the determination of the topography of the proteins in the ribosome. More recently, a near-complete (near)atomic picture of the ribosomal proteins is emerging from the latest high-resolution cryo-EM data (including ). Conservation Ribosomal prot ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Rho Family Of GTPases
The Rho family of GTPases is a family of small (~21 kDa) signaling G proteins, and is a subfamily of the Ras superfamily. The members of the Rho GTPase family have been shown to regulate many aspects of intracellular actin dynamics, and are found in all eukaryotic kingdoms, including yeasts and some plants. Three members of the family have been studied in detail: Cdc42, Rac1, and RhoA. All G proteins are "molecular switches", and Rho proteins play a role in organelle development, cytoskeletal dynamics, cell movement, and other common cellular functions. History Identification of the Rho family of GTPases began in the mid-1980s. The first identified Rho member was RhoA, isolated serendipitously in 1985 from a low stringency cDNA screening. Rac1 and Rac2 were identified next, in 1989 followed by Cdc42 in 1990. Eight additional mammalian Rho members were identified from biological screenings until the late 1990s, a turning point in biology where availability of complete genome seq ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribosomal Protein L13 Leader
L13 ribosomal protein leaders play a role in ribosome biogenesis as part of an autoregulatory mechanism to control the concentration of ribosomal proteins L13. Three structural classes of L13 ribosomal protein leaders were detected by different bioinformatics approaches: in ''B. subtilis'' and other low-GC Gram-positive bacteria., in ''E. coli'' and in Bacteroidia. Although these RNAs are expected to perform the same biological function, they do not appear to be structurally related to one another. The ''E. coli'' example has been experimentally confirmed, though the experiments are not comprehensive. The other two leader structures are thus far not based on experimental support. See also Ribosomal protein leader A ribosomal protein leader is a mechanism used in cells to control the cellular concentration of a protein that forms a part of the ribosome, and to make sure that the concentration is neither too high nor too low. Ribosomal protein leaders are RNA ... Reference ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribosomal Protein L19 Leader
L19 Ribosomal protein leaders are part of the ribosome biogenesis. They are used as an autoregulatory mechanism to control the concentration of ribosomal proteins L19, and are located in the 5′ untranslated regions of mRNAs encoding ribosomal protein L19 (rplS). L19 ribosomal protein leaders have been bioinformatically predicted in ''B. subtilis'' and other low-GC Gram-positive bacteria in the phylum Bacillota. More examples that share a similar structure were predicted in Flavobacteria, also using bioinformatic approaches. See also *Ribosomal protein leader References External links * Ribosomal protein leader {{molecular-cell-biology-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |