|
RNA-sequencing
RNA-Seq (named as an abbreviation of RNA sequencing) is a DNA sequencing, sequencing technique which uses next-generation sequencing (NGS) to reveal the presence and quantity of RNA in a biological sample at a given moment, analyzing the continuously changing cellular transcriptome. Specifically, RNA-Seq facilitates the ability to look at Alternative splicing, alternative gene spliced transcripts, post-transcriptional modifications, gene fusion, mutations/single nucleotide polymorphism, SNPs and changes in gene expression over time, or differences in gene expression in different groups or treatments. In addition to mRNA transcripts, RNA-Seq can look at different populations of RNA to include total RNA, small RNA, such as miRNA, tRNA, and Ribosome profiling, ribosomal profiling. RNA-Seq can also be used to determine exon/intron boundaries and verify or amend previously Gene annotation, annotated Directionality (molecular biology)#5.E2.80.B2-end, 5' and Directionality (molecular bi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Single-cell Transcriptomics
Single-cell transcriptomics examines the gene expression level of individual cells in a given population by simultaneously measuring the RNA concentration (conventionally only messenger RNA (mRNA)) of hundreds to thousands of genes. Single-cell transcriptomics makes it possible to unravel heterogeneous cell populations, reconstruct cellular developmental pathways, and model transcriptional dynamics — all previously masked in bulk RNA sequencing. Background The development of high-throughput RNA sequencing (RNA-seq) and microarrays has made gene expression analysis a routine. RNA analysis was previously limited to tracing individual transcripts by Northern blots or quantitative PCR. Higher throughput and speed allow researchers to frequently characterize the expression profiles of populations of thousands of cells. The data from bulk assays has led to identifying genes differentially expressed in distinct cell populations, and biomarker discovery. These studies are limited as ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transcriptomics
Transcriptomics technologies are the techniques used to study an organism's transcriptome, the sum of all of its RNA transcripts. The information content of an organism is recorded in the DNA of its genome and expressed through transcription. Here, mRNA serves as a transient intermediary molecule in the information network, whilst non-coding RNAs perform additional diverse functions. A transcriptome captures a snapshot in time of the total transcripts present in a cell. Transcriptomics technologies provide a broad account of which cellular processes are active and which are dormant. A major challenge in molecular biology is to understand how a single genome gives rise to a variety of cells. Another is how gene expression is regulated. The first attempts to study whole transcriptomes began in the early 1990s. Subsequent technological advances since the late 1990s have repeatedly transformed the field and made transcriptomics a widespread discipline in biological sciences. There ar ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Microarray
A DNA microarray (also commonly known as DNA chip or biochip) is a collection of microscopic DNA spots attached to a solid surface. Scientists use DNA microarrays to measure the expression levels of large numbers of genes simultaneously or to genotype multiple regions of a genome. Each DNA spot contains picomoles (10−12 moles) of a specific DNA sequence, known as '' probes'' (or ''reporters'' or '' oligos''). These can be a short section of a gene or other DNA element that are used to hybridize a cDNA or cRNA (also called anti-sense RNA) sample (called ''target'') under high-stringency conditions. Probe-target hybridization is usually detected and quantified by detection of fluorophore-, silver-, or chemiluminescence-labeled targets to determine relative abundance of nucleic acid sequences in the target. The original nucleic acid arrays were macro arrays approximately 9 cm × 12 cm and the first computerized image based analysis was published in 1981. It was inv ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA Integrity Number
The RNA integrity number (RIN) is an algorithm for assigning integrity values to RNA measurements. The integrity of RNA is a major concern for gene expression studies and traditionally has been evaluated using the 28S to 18S rRNA ratio, a method that has been shown to be inconsistent. This inconsistency arises because subjective, human interpretation is necessary to compare the 28S and 18S gel images. The RIN algorithm was devised to overcome this issue. The RIN algorithm is applied to electrophoretic RNA measurements, typically obtained using capillary gel electrophoresis, and based on a combination of different features that contribute information about the RNA integrity to provide a more universal measure. RIN has been demonstrated to be robust and reproducible in studies comparing it to other RNA integrity calculation algorithms, cementing its position as a preferred method of determining the quality of RNA to be analyzed. A major criticism to RIN is when using with plants o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Capillary Electrophoresis
Capillary electrophoresis (CE) is a family of electrokinetic separation methods performed in submillimeter diameter capillaries and in micro- and nanofluidic channels. Very often, CE refers to capillary zone electrophoresis (CZE), but other electrophoretic techniques including capillary gel electrophoresis (CGE), capillary isoelectric focusing (CIEF), capillary isotachophoresis and micellar electrokinetic chromatography (MEKC) belong also to this class of methods. In CE methods, analytes migrate through electrolyte solutions under the influence of an electric field. Analytes can be separated according to ionic mobility and/or partitioning into an alternate phase via non-covalent interactions. Additionally, analytes may be concentrated or "focused" by means of gradients in conductivity and pH. Instrumentation The instrumentation needed to perform capillary electrophoresis is relatively simple. A basic schematic of a capillary electrophoresis system is shown in ''figure 1''. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gel Electrophoresis
Gel electrophoresis is a method for separation and analysis of biomacromolecules ( DNA, RNA, proteins, etc.) and their fragments, based on their size and charge. It is used in clinical chemistry to separate proteins by charge or size (IEF agarose, essentially size independent) and in biochemistry and molecular biology to separate a mixed population of DNA and RNA fragments by length, to estimate the size of DNA and RNA fragments or to separate proteins by charge. Nucleic acid molecules are separated by applying an electric field to move the negatively charged molecules through a matrix of agarose or other substances. Shorter molecules move faster and migrate farther than longer ones because shorter molecules migrate more easily through the pores of the gel. This phenomenon is called sieving. Proteins are separated by the charge in agarose because the pores of the gel are too small to sieve proteins. Gel electrophoresis can also be used for the separation of nanoparticles. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Deoxyribonuclease
Deoxyribonuclease (DNase, for short) refers to a group of glycoprotein endonucleases which are enzymes that catalyze the hydrolytic cleavage of phosphodiester linkages in the DNA backbone, thus degrading DNA. The role of the DNase enzyme in cells includes breaking down extracellular DNA (ecDNA) excreted by apoptosis, necrosis, and neutrophil extracellular traps (NET) of cells to help reduce inflammatory responses that otherwise are elicited. A wide variety of deoxyribonucleases are known and fall into one of two families (DNase I or DNase II), which differ in their substrate specificities, chemical mechanisms, and biological functions. Laboratory applications of DNase include purifying proteins when extracted from prokaryotic organisms. Additionally, DNase has been applied as a treatment for diseases that are caused by ecDNA in the blood plasma. Assays of DNase are emerging in the research field as well. Types The two main types of DNase found in metazoans are known as deoxyr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA Extraction
RNA extraction is the purification of RNA from biological samples. This procedure is complicated by the ubiquitous presence of ribonuclease enzymes in cells and tissues, which can rapidly degrade RNA. Several methods are used in molecular biology to isolate RNA from samples, the most common of these is guanidinium thiocyanate-phenol-chloroform extraction. The filter paper based lysis and elution method features high throughput capacity. RNA extraction in liquid nitrogen, commonly using a mortar and pestle (or specialized steel devices known as tissue pulverizers) is also useful in preventing ribonuclease activity. RNase contamination The extraction of RNA in molecular biology experiments is greatly complicated by the presence of ubiquitous and hardy RNases that degrade RNA samples. Certain RNases can be extremely hardy and inactivating them is difficult compared to neutralizing DNases. In addition to the cellular RNases that are released there are several RNases that are present in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Journal
A journal, from the Old French ''journal'' (meaning "daily"), may refer to: *Bullet journal, a method of personal organization *Diary, a record of what happened over the course of a day or other period *Daybook, also known as a general journal, a daily record of financial transactions * Logbook, a record of events important to the operation of a vehicle, facility, or otherwise *Record (other) *Transaction log, a chronological record of data processing *Travel journal In publishing, ''journal'' can refer to various periodicals or serials: *Academic journal, an academic or scholarly periodical **Scientific journal, an academic journal focusing on science **Medical journal, an academic journal focusing on medicine **Law review, a professional journal focusing on legal interpretation *Magazine, non-academic or scholarly periodicals in general **Trade magazine, a magazine of interest to those of a particular profession or trade **Literary magazine, a magazine devoted to litera ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Complementary DNA
In genetics, complementary DNA (cDNA) is DNA synthesized from a single-stranded RNA (e.g., messenger RNA (mRNA) or microRNA (miRNA)) template in a reaction catalyzed by the enzyme reverse transcriptase. cDNA is often used to express a specific protein in a cell that does not normally express that protein (i.e., heterologous expression), or to sequence or quantify mRNA molecules using DNA based methods (qPCR, RNA-seq). cDNA that codes for a specific protein can be transferred to a recipient cell for expression, often bacterial or yeast expression systems. cDNA is also generated to analyze transcriptomic profiles in bulk tissue, single cells, or single nuclei in assays such as microarrays, qPCR, and RNA-seq. cDNA is also produced naturally by retroviruses (such as HIV-1, HIV-2, simian immunodeficiency virus, etc.) and then integrated into the host's genome, where it creates a provirus. The term ''cDNA'' is also used, typically in a bioinformatics context, to refer to a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Next-gen Sequencing
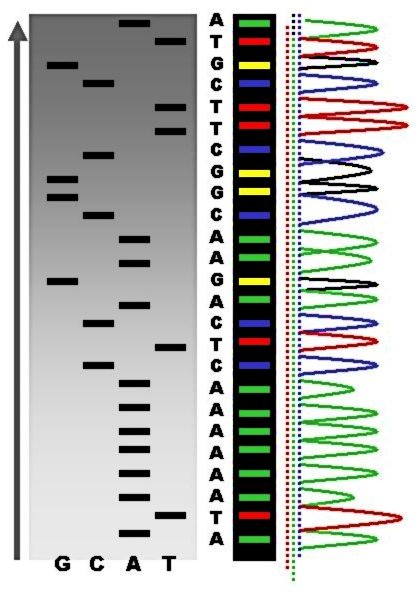
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery. Knowledge of DNA sequences has become indispensable for basic biological research, DNA Genographic Projects and in numerous applied fields such as medical diagnosis, biotechnology, forensic biology, virology and biological systematics. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers, characterize antibody repertoire, and can be used to guide patient treatment. Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged. The rapid speed of sequencing attained with modern DNA ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Serial Analysis Of Gene Expression
Serial Analysis of Gene Expression (SAGE) is a transcriptomic technique used by molecular biologists to produce a snapshot of the messenger RNA population in a sample of interest in the form of small tags that correspond to fragments of those transcripts. Several variants have been developed since, most notably a more robust version, LongSAGE, RL-SAGE and the most recent SuperSAGE. Many of these have improved the technique with the capture of longer tags, enabling more confident identification of a source gene. Overview Briefly, SAGE experiments proceed as follows: # The mRNA of an input sample (e.g. a tumour) is isolated and a reverse transcriptase and biotinylated primers are used to synthesize cDNA from mRNA. # The cDNA is bound to Streptavidin beads via interaction with the biotin attached to the primers, and is then cleaved using a restriction endonuclease called an anchoring enzyme (AE). The location of the cleavage site and thus the length of the remaining cDNA bound ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
_B2.jpg)