|
Retroelements
Retrotransposons (also called Class I transposable elements or transposons via RNA intermediates) are a type of genetic component that copy and paste themselves into different genomic locations (transposon) by converting RNA back into DNA through the reverse transcription process using an RNA transposition intermediate. Through reverse transcription, retrotransposons amplify themselves quickly to become abundant in eukaryotic genomes such as maize (49–78%) and humans (42%). They are only present in eukaryotes but share features with retroviruses such as HIV, for example, discontinuous reverse transcriptase-mediated extrachromosomal recombination. These retrotransposons are regulated by a family of short non-coding RNAs termed as PIWI -element induced wimpy testisinteracting RNAs (piRNAs). piRNA is a recently discovered class of ncRNAs, which are in the length range of ~24-32 nucleotides. Initially, piRNAs were described as repeat-associated siRNAs (rasiRNAs) because of their ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Retrotransposons
Retrotransposons (also called Class I transposable elements or transposons via RNA intermediates) are a type of genetic component that copy and paste themselves into different genomic locations ( transposon) by converting RNA back into DNA through the reverse transcription process using an RNA transposition intermediate. Through reverse transcription, retrotransposons amplify themselves quickly to become abundant in eukaryotic genomes such as maize (49–78%) and humans (42%). They are only present in eukaryotes but share features with retroviruses such as HIV, for example, discontinuous reverse transcriptase-mediated extrachromosomal recombination. These retrotransposons are regulated by a family of short non-coding RNAs termed as PIWI -element induced wimpy testisinteracting RNAs (piRNAs). piRNA is a recently discovered class of ncRNAs, which are in the length range of ~24-32 nucleotides. Initially, piRNAs were described as repeat-associated siRNAs (rasiRNAs) because of the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Restriction Enzyme
A restriction enzyme, restriction endonuclease, REase, ENase or'' restrictase '' is an enzyme that cleaves DNA into fragments at or near specific recognition sites within molecules known as restriction sites. Restriction enzymes are one class of the broader endonuclease group of enzymes. Restriction enzymes are commonly classified into five types, which differ in their structure and whether they cut their DNA substrate at their recognition site, or if the recognition and cleavage sites are separate from one another. To cut DNA, all restriction enzymes make two incisions, once through each sugar-phosphate backbone (i.e. each strand) of the DNA double helix. These enzymes are found in bacteria and archaea and provide a defense mechanism against invading viruses. Inside a prokaryote, the restriction enzymes selectively cut up ''foreign'' DNA in a process called ''restriction digestion''; meanwhile, host DNA is protected by a modification enzyme (a methyltransferase) that modi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Retrovirus
A retrovirus is a type of virus that inserts a DNA copy of its RNA genome into the DNA of a host cell that it invades, thus changing the genome of that cell. Once inside the host cell's cytoplasm, the virus uses its own reverse transcriptase enzyme to produce DNA from its RNA genome, the reverse of the usual pattern, thus ''retro'' (backwards). The new DNA is then incorporated into the host cell genome by an integrase enzyme, at which point the retroviral DNA is referred to as a provirus. The host cell then treats the viral DNA as part of its own genome, transcribing and translating the viral genes along with the cell's own genes, producing the proteins required to assemble new copies of the virus. Although retroviruses have different subfamilies, they have three basic groups: the oncoretroviruses (oncogenic retroviruses), the lentiviruses (slow retroviruses) and the spumaviruses (foamy viruses). The oncoretroviruses are able to cause cancer in some species, the lentivi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribonuclease
Ribonuclease (commonly abbreviated RNase) is a type of nuclease that catalyzes the degradation of RNA into smaller components. Ribonucleases can be divided into endoribonucleases and exoribonucleases, and comprise several sub-classes within the EC 2.7 (for the phosphorolytic enzymes) and 3.1 (for the hydrolytic enzymes) classes of enzymes. Function All organisms studied contain many RNases of two different classes, showing that RNA degradation is a very ancient and important process. As well as clearing of cellular RNA that is no longer required, RNases play key roles in the maturation of all RNA molecules, both messenger RNAs that carry genetic material for making proteins and non-coding RNAs that function in varied cellular processes. In addition, active RNA degradation systems are the first defense against RNA viruses and provide the underlying machinery for more advanced cellular immune strategies such as RNAi. Some cells also secrete copious quantities of non-specific ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
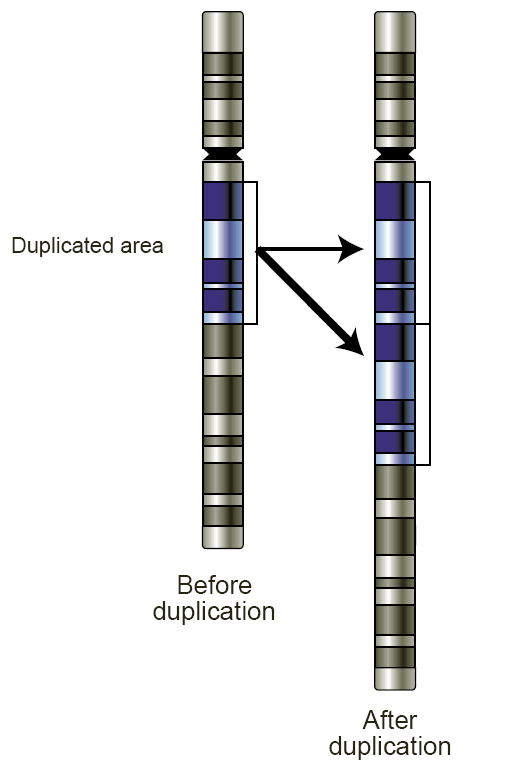
Copy-number Variation
Copy number variation (CNV) is a phenomenon in which sections of the genome are repeated and the number of repeats in the genome varies between individuals. Copy number variation is a type of structural variation: specifically, it is a type of duplication or deletion event that affects a considerable number of base pairs. Approximately two-thirds of the entire human genome may be composed of repeats and 4.8–9.5% of the human genome can be classified as copy number variations. In mammals, copy number variations play an important role in generating necessary variation in the population as well as disease phenotype. Copy number variations can be generally categorized into two main groups: short repeats and long repeats. However, there are no clear boundaries between the two groups and the classification depends on the nature of the loci of interest. Short repeats include mainly dinucleotide repeats (two repeating nucleotides e.g. A-C-A-C-A-C...) and trinucleotide repeats. Long r ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Non-coding RNA
A non-coding RNA (ncRNA) is a functional RNA molecule that is not translated into a protein. The DNA sequence from which a functional non-coding RNA is transcribed is often called an RNA gene. Abundant and functionally important types of non-coding RNAs include transfer RNAs (tRNAs) and ribosomal RNAs (rRNAs), as well as small RNAs such as microRNAs, siRNAs, piRNAs, snoRNAs, snRNAs, exRNAs, scaRNAs and the long ncRNAs such as Xist and HOTAIR. The number of non-coding RNAs within the human genome is unknown; however, recent transcriptomic and bioinformatic studies suggest that there are thousands of non-coding transcripts. Many of the newly identified ncRNAs have not been validated for their function. There is no consensus in the literature on how much of non-coding transcription is functional. Some researchers have argued that many ncRNAs are non-functional (sometimes referred to as "junk RNA"), spurious transcriptions. Others, however, disagree, arguing instead ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA Interference
RNA interference (RNAi) is a biological process in which RNA molecules are involved in sequence-specific suppression of gene expression by double-stranded RNA, through translational or transcriptional repression. Historically, RNAi was known by other names, including ''co-suppression'', ''post-transcriptional gene silencing'' (PTGS), and ''quelling''. The detailed study of each of these seemingly different processes elucidated that the identity of these phenomena were all actually RNAi. Andrew Fire and Craig C. Mello shared the 2006 Nobel Prize in Physiology or Medicine for their work on RNAi in the nematode worm ''Caenorhabditis elegans'', which they published in 1998. Since the discovery of RNAi and its regulatory potentials, it has become evident that RNAi has immense potential in suppression of desired genes. RNAi is now known as precise, efficient, stable and better than antisense therapy for gene suppression. Antisense RNA produced intracellularly by an expression vector ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alu Element
An Alu element is a short stretch of DNA originally characterized by the action of the '' Arthrobacter luteus (Alu)'' restriction endonuclease. ''Alu'' elements are the most abundant transposable elements, containing over one million copies dispersed throughout the human genome. ''Alu'' elements were thought to be selfish or parasitic DNA, because their sole known function is self reproduction. However, they are likely to play a role in evolution and have been used as genetic markers. They are derived from the small cytoplasmic 7SL RNA, a component of the signal recognition particle. ''Alu'' elements are highly conserved within primate genomes and originated in the genome of an ancestor of Supraprimates. ''Alu'' insertions have been implicated in several inherited human diseases and in various forms of cancer. The study of Alu elements has also been important in elucidating human population genetics and the evolution of primates, including the evolution of humans. Alu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Long Interspersed Nuclear Element
Long interspersed nuclear elements (LINEs) (also known as long interspersed nucleotide elements or long interspersed elements) are a group of non-LTR ( long terminal repeat) retrotransposons that are widespread in the genome of many eukaryotes. They make up around 21.1% of the human genome. LINEs make up a family of transposons, where each LINE is about 7,000 base pairs long. LINEs are transcribed into mRNA and translated into protein that acts as a reverse transcriptase. The reverse transcriptase makes a DNA copy of the LINE RNA that can be integrated into the genome at a new site. The only abundant LINE in humans is LINE1. The human genome contains an estimated 100,000 truncated and 4,000 full-length LINE-1 elements. Due to the accumulation of random mutations, the sequence of many LINEs has degenerated to the extent that they are no longer transcribed or translated. Comparisons of LINE DNA sequences can be used to date transposon insertion in the genome. History of di ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
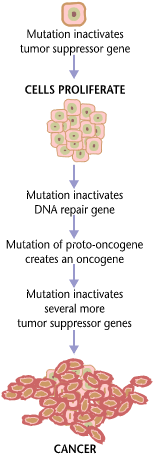
Tumorigenesis
Carcinogenesis, also called oncogenesis or tumorigenesis, is the formation of a cancer, whereby normal cells are transformed into cancer cells. The process is characterized by changes at the cellular, genetic, and epigenetic levels and abnormal cell division. Cell division is a physiological process that occurs in almost all tissues and under a variety of circumstances. Normally, the balance between proliferation and programmed cell death, in the form of apoptosis, is maintained to ensure the integrity of tissues and organs. According to the prevailing accepted theory of carcinogenesis, the somatic mutation theory, mutations in DNA and epimutations that lead to cancer disrupt these orderly processes by interfering with the programming regulating the processes, upsetting the normal balance between proliferation and cell death. This results in uncontrolled cell division and the evolution of those cells by natural selection in the body. Only certain mutations lead to can ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Direct Repeat
Direct repeats are a type of genetic sequence that consists of two or more repeats of a specific sequence. In other words, the direct repeats are nucleotide sequences present in multiple copies in the genome. Generally, a direct repeat occurs when a sequence is repeated with the same pattern downstream. There is and no reverse complement associated with a direct repeat. It may or may not have intervening nucleotides. The nucleotide sequence written in bold characters signifies the repeated sequence. ::: ::: Linguistically, a typical direct repeat is comparable to saying "bye-bye". Types There are several types of repeated sequences : *Interspersed (or dispersed) DNA repeats (interspersed repetitive sequences) are copies of transposable elements interspersed throughout the genome. *Flanking (or terminal) repeats (terminal repeat sequences) are sequences that are repeated on both ends of a sequence, for example, the long terminal repeats (LTRs) on retroviruses. Direct terminal r ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |