|
Read Length
In DNA sequencing, a read is an inferred sequence of base pairs (or base pair probabilities) corresponding to all or part of a single DNA fragment. A typical sequencing experiment involves fragmentation of the genome into millions of molecules, which are size-selected and ligated to adapters. The set of fragments is referred to as a sequencing library, which is sequenced to produce a set of reads. Read length Sequencing technologies vary in the length of reads produced. Reads of length 20-40 base pairs (bp) are referred to as ultra-short. Typical sequencers produce read lengths in the range of 100-500 bp. However, Pacific Biosciences platforms produce read lengths of approximately 1500 bp. Read length is a factor which can affect the results of biological studies. For example, longer read lengths improve the resolution of ''de novo'' genome assembly and detection of structural variants. It is estimated that read lengths greater than 100 kilobases (kb) will be required for rout ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Sequencing
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery. Knowledge of DNA sequences has become indispensable for basic biological research, DNA Genographic Projects and in numerous applied fields such as medical diagnosis, biotechnology, forensic biology, virology and biological systematics. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers, characterize antibody repertoire, and can be used to guide patient treatment. Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged. The rapid speed of sequencing attained with modern D ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Base Pair
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA and RNA. Dictated by specific hydrogen bonding patterns, "Watson–Crick" (or "Watson–Crick–Franklin") base pairs (guanine–cytosine and adenine–thymine) allow the DNA helix to maintain a regular helical structure that is subtly dependent on its nucleotide sequence. The Complementarity (molecular biology), complementary nature of this based-paired structure provides a redundant copy of the genetic information encoded within each strand of DNA. The regular structure and data redundancy provided by the DNA double helix make DNA well suited to the storage of genetic information, while base-pairing between DNA and incoming nucleotides provides the mechanism through which DNA polymerase replicates DNA and RNA polymerase transcribes DNA in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Fragmentation
DNA fragmentation is the separation or breaking of DNA strands into pieces. It can be done intentionally by laboratory personnel or by cells, or can occur spontaneously. Spontaneous or accidental DNA fragmentation is fragmentation that gradually accumulates in a cell. It can be measured by e.g. the Comet assay or by the TUNEL assay. Its main units of measurement is the DNA Fragmentation Index (DFI). A DFI of 20% or more significantly reduces the success rates after ICSI. DNA fragmentation was first documented by Williamson in 1970 when he observed discrete oligomeric fragments occurring during cell death in primary neonatal liver cultures. He described the cytoplasmic DNA isolated from mouse liver cells after culture as characterized by DNA fragments with a molecular weight consisting of multiples of 135 kDa. This finding was consistent with the hypothesis that these DNA fragments were a specific degradation product of nuclear DNA. Intentional DNA fragmentation is often necessar ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ligation (molecular Biology)
In molecular biology, ligation is the joining of two nucleic acid fragments through the action of an enzyme. It is an essential laboratory procedure in the molecular cloning of DNA, whereby DNA fragments are joined to create recombinant DNA molecules (such as when a foreign DNA fragment is inserted into a plasmid). The ends of DNA fragments are joined by the formation of phosphodiester bonds between the 3'-hydroxyl of one DNA terminus with the 5'-phosphoryl of another. RNA may also be ligated similarly. A co-factor is generally involved in the reaction, and this is usually ATP or NAD+. Ligation in the laboratory is normally performed using T4 DNA ligase. However, procedures for ligation without the use of standard DNA ligase are also popular. Ligation reaction The mechanism of the ligation reaction was first elucidated in the laboratory of I. Robert Lehman. Two fragments of DNA may be joined by DNA ligase which catalyzes the formation of a phosphodiester bond between the 3 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Adapter (genetics)
An adapter or adaptor, or a linker in genetic engineering is a short, chemically synthesized, single-stranded or double-stranded oligonucleotide that can be ligated to the ends of other DNA or RNA molecules. Double stranded adapters can be synthesized to have blunt ends to both terminals or to have sticky end at one end and blunt end at the other. For instance, a double stranded DNA adapter can be used to link the ends of two other DNA molecules (i.e., ends that do not have "sticky ends", that is complementary protruding single strands by themselves). It may be used to add sticky ends to cDNA allowing it to be ligated into the plasmid much more efficiently. Two adapters could base pair to each other to form dimers. A conversion adapter is used to join a DNA insert cut with one restriction enzyme A restriction enzyme, restriction endonuclease, REase, ENase or'' restrictase '' is an enzyme that cleaves DNA into fragments at or near specific recognition sites within molecules k ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Pacific Biosciences
Pacific Biosciences of California, Inc. (aka PacBio) is an American biotechnology company founded in 2004 that develops and manufactures systems for gene sequencing and some novel real time biological observation. PacBio describes its platform as single-molecule real-time sequencing (SMRT), based on the properties of zero-mode waveguides. History The company was founded based on research done at Cornell University that combined semiconductor processing and photonics with biotechnology research. Three graduate students in the lab of Professors Watt W. Webb — Jonas Korlach — and Harold Craighead — Steve Turner and Mathieu Foquet — became the first employees. It began under the name Nanofluidics, Inc. The company raised nearly in six rounds of primarily venture capital financing, making it one of the most capitalized startups in 2010 leading up to their public offering in October of that year. Key investors included Mohr Davidow Ventures, Kleiner, P ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sanger Sequencing
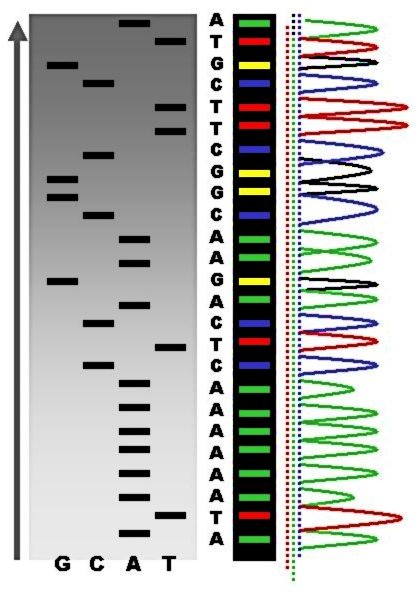
Sanger sequencing is a method of DNA sequencing that involves electrophoresis and is based on the random incorporation of chain-terminating dideoxynucleotides by DNA polymerase during in vitro DNA replication. After first being developed by Frederick Sanger and colleagues in 1977, it became the most widely used sequencing method for approximately 40 years. It was first commercialized by Applied Biosystems in 1986. More recently, higher volume Sanger sequencing has been replaced by next generation sequencing methods, especially for large-scale, automated genome analyses. However, the Sanger method remains in wide use for smaller-scale projects and for validation of deep sequencing results. It still has the advantage over short-read sequencing technologies (like Illumina) in that it can produce DNA sequence reads of > 500 nucleotides and maintains a very low error rate with accuracies around 99.99%. Sanger sequencing is still actively being used in efforts for public health initiative ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Maxam–Gilbert Sequencing
Maxam–Gilbert sequencing is a method of DNA sequencing developed by Allan Maxam and Walter Gilbert in 1976–1977. This method is based on nucleobase-specific partial chemical modification of DNA and subsequent cleavage of the DNA backbone at sites adjacent to the modified nucleotides. Maxam–Gilbert sequencing was the first widely adopted method for DNA sequencing, and, along with the Sanger dideoxy method, represents the first generation of DNA sequencing methods. Maxam–Gilbert sequencing is no longer in widespread use, having been supplanted by next-generation sequencing methods. History Although Maxam and Gilbert published their chemical sequencing method two years after Frederick Sanger and Alan Coulson published their work on plus-minus sequencing, Maxam–Gilbert sequencing rapidly became more popular, since purified DNA could be used directly, while the initial Sanger method required that each read start be cloned for production of single-stranded DNA. However, with ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Massive Parallel Sequencing
Massive parallel sequencing or massively parallel sequencing is any of several high-throughput approaches to DNA sequencing using the concept of massively parallel processing; it is also called next-generation sequencing (NGS) or second-generation sequencing. Some of these technologies emerged between 1994 and 1998 and have been commercially available since 2005. These technologies use miniaturized and parallelized platforms for sequencing of 1 million to 43 billion short reads (50 to 400 bases each) per instrument run. Many NGS platforms differ in engineering configurations and sequencing chemistry. They share the technical paradigm of massive parallel sequencing via spatially separated, clonally amplified DNA templates or single DNA molecules in a flow cell. This design is very different from that of Sanger sequencing—also known as capillary sequencing or first-generation sequencing—which is based on electrophoretic separation of chain-termination products produced in individ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Third-generation Sequencing
Third-generation sequencing (also known as long-read sequencing) is a class of DNA sequencing methods currently under active development. Third generation sequencing technologies have the capability to produce substantially longer reads than second generation sequencing, also known as next-generation sequencing. Such an advantage has critical implications for both genome science and the study of biology in general. However, third generation sequencing data have much higher error rates than previous technologies, which can complicate downstream genome assembly and analysis of the resulting data. These technologies are undergoing active development and it is expected that there will be improvements to the high error rates. For applications that are more tolerant to error rates, such as structural variant calling, third generation sequencing has been found to outperform existing methods, even at a low depth of sequencing coverage. Current technologies Sequencing technologies with a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Reference Genome
A reference genome (also known as a reference assembly) is a digital nucleic acid sequence database, assembled by scientists as a representative example of the set of genes in one idealized individual organism of a species. As they are assembled from the sequencing of DNA from a number of individual donors, reference genomes do not accurately represent the set of genes of any single individual organism. Instead a reference provides a haploid mosaic of different DNA sequences from each donor. For example, the most recent human reference genome (assembly '' GRCh38/hg38'') is derived from >60 genomic clone libraries. There are reference genomes for multiple species of viruses, bacteria, fungus, plants, and animals. Reference genomes are typically used as a guide on which new genomes are built, enabling them to be assembled much more quickly and cheaply than the initial Human Genome Project. Reference genomes can be accessed online at several locations, using dedicated browsers s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Epigenetics
In biology, epigenetics is the study of stable phenotypic changes (known as ''marks'') that do not involve alterations in the DNA sequence. The Greek prefix '' epi-'' ( "over, outside of, around") in ''epigenetics'' implies features that are "on top of" or "in addition to" the traditional genetic basis for inheritance. Epigenetics most often involves changes that affect the regulation of gene expression, but the term can also be used to describe any heritable phenotypic change. Such effects on cellular and physiological phenotypic traits may result from external or environmental factors, or be part of normal development. The term also refers to the mechanism of changes: functionally relevant alterations to the genome that do not involve mutation of the nucleotide sequence. Examples of mechanisms that produce such changes are DNA methylation and histone modification, each of which alters how genes are expressed without altering the underlying DNA sequence. Gene expression c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |