|
Random Amplified Polymorphic DNA
Random amplification of polymorphic DNA (RAPD), pronounced "rapid", is a type of polymerase chain reaction (PCR), but the segments of DNA that are amplified are random. The scientist performing RAPD creates several arbitrary, short primers (10- 12 nucleotides), then proceeds with the PCR using a large template of genomic DNA, hoping that fragments will amplify. By resolving the resulting patterns, a semi-unique profile can be gleaned from an RAPD reaction. No knowledge of the DNA sequence of the targeted genome is required, as the primers will bind somewhere in the sequence, but it is not certain exactly where. This makes the method popular for comparing the DNA of biological systems that have not had the attention of the scientific community, or in a system in which relatively few DNA sequences are compared (it is not suitable for forming a cDNA databank). Because it relies on a large, intact DNA template sequence, it has some limitations in the use of degraded DNA samples. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RAPD
{{dab ...
RAPD may refer to: *Relative afferent pupillary defect *Random amplification of polymorphic DNA Random amplification of polymorphic DNA (RAPD), pronounced "rapid", is a type of polymerase chain reaction (PCR), but the segments of DNA that are amplified are random. The scientist performing RAPD creates several arbitrary, short primers (10- 1 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polymerase Chain Reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) to a large enough amount to study in detail. PCR was invented in 1983 by the American biochemist Kary Mullis at Cetus Corporation; Mullis and biochemist Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993. PCR is fundamental to many of the procedures used in genetic testing and research, including analysis of ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory research for a broad variety of applications including biomedical research ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Short Tandem Repeats
A microsatellite is a tract of repetitive DNA in which certain DNA motifs (ranging in length from one to six or more base pairs) are repeated, typically 5–50 times. Microsatellites occur at thousands of locations within an organism's genome. They have a higher mutation rate than other areas of DNA leading to high genetic diversity. Microsatellites are often referred to as short tandem repeats (STRs) by forensic geneticists and in genetic genealogy, or as simple sequence repeats (SSRs) by plant geneticists. Microsatellites and their longer cousins, the minisatellites, together are classified as VNTR (variable number of tandem repeats) DNA. The name "satellite" DNA refers to the early observation that centrifugation of genomic DNA in a test tube separates a prominent layer of bulk DNA from accompanying "satellite" layers of repetitive DNA. They are widely used for DNA profiling in cancer diagnosis, in kinship analysis (especially paternity testing) and in forensic identificat ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
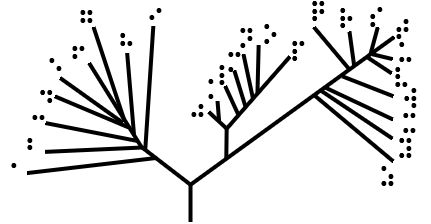
Phylogeny
A phylogenetic tree (also phylogeny or evolutionary tree Felsenstein J. (2004). ''Inferring Phylogenies'' Sinauer Associates: Sunderland, MA.) is a branching diagram or a tree showing the evolutionary relationships among various biological species or other entities based upon similarities and differences in their physical or genetic characteristics. All life on Earth is part of a single phylogenetic tree, indicating common ancestry. In a ''rooted'' phylogenetic tree, each node with descendants represents the inferred most recent common ancestor of those descendants, and the edge lengths in some trees may be interpreted as time estimates. Each node is called a taxonomic unit. Internal nodes are generally called hypothetical taxonomic units, as they cannot be directly observed. Trees are useful in fields of biology such as bioinformatics, systematics, and phylogenetics. ''Unrooted'' trees illustrate only the relatedness of the leaf nodes and do not require the ancestral root ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gel Electrophoresis Of Nucleic Acids
Nucleic acid electrophoresis is an analytical technique used to separate DNA or RNA fragments by size and reactivity. Nucleic acid molecules which are to be analyzed are set upon a viscous medium, the gel, where an electric field induces the nucleic acids (which are negatively charged due to their sugar- phosphate backbone) to migrate toward the anode (which is positively charged because this is an electrolytic rather than galvanic cell). The separation of these fragments is accomplished by exploiting the mobilities with which different sized molecules are able to pass through the gel. Longer molecules migrate more slowly because they experience more resistance within the gel. Because the size of the molecule affects its mobility, smaller fragments end up nearer to the anode than longer ones in a given period. After some time, the voltage is removed and the fragmentation gradient is analyzed. For larger separations between similar sized fragments, either the voltage or run time ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cleaved Amplified Polymorphic Sequence
The cleaved amplified polymorphic sequence (CAPS) method is a technique in molecular biology for the analysis of genetic markers. It is an extension to the restriction fragment length polymorphism (RFLP) method, using polymerase chain reaction (PCR) to more quickly analyse the results. Like RFLP, CAPS works on the principle that genetic differences between individuals can create or abolish restriction endonuclease restriction sites, and that these differences can be detected in the resulting DNA fragment length after digestion. In the CAPS method, PCR amplification is directed across the altered restriction site, and the products digested with the restriction enzyme. When fractionated by agarose or polyacrylamide gel electrophoresis, the digested PCR products will give readily distinguishable patterns of bands. Alternatively, the amplified segment can be analyzed by allele-specific oligonucleotide (ASO) probes, a process that can often be done by a simple dot blot A dot blot (or ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genetic Marker
A genetic marker is a gene or DNA sequence with a known location on a chromosome that can be used to identify individuals or species. It can be described as a variation (which may arise due to mutation or alteration in the genomic loci) that can be observed. A genetic marker may be a short DNA sequence, such as a sequence surrounding a single base-pair change ( single nucleotide polymorphism, SNP), or a long one, like minisatellites. Background For many years, gene mapping was limited to identifying organisms by traditional phenotypes markers. This included genes that encoded easily observable characteristics such as blood types or seed shapes. The insufficient number of these types of characteristics in several organisms limited the mapping efforts that could be done. This prompted the development of gene markers which could identify genetic characteristics that are not readily observable in organisms (such as protein variation). Types Some commonly used types of genetic mark ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Biochemistry Detection Methods
Biochemistry or biological chemistry is the study of chemical processes within and relating to living organisms. A sub-discipline of both chemistry and biology, biochemistry may be divided into three fields: structural biology, enzymology and metabolism. Over the last decades of the 20th century, biochemistry has become successful at explaining living processes through these three disciplines. Almost all areas of the life sciences are being uncovered and developed through biochemical methodology and research.Voet (2005), p. 3. Biochemistry focuses on understanding the chemical basis which allows biological molecules to give rise to the processes that occur within living cells and between cells, Karp (2009), p. 2. in turn relating greatly to the understanding of tissues and organs, as well as organism structure and function. Miller (2012). p. 62. Biochemistry is closely related to molecular biology, which is the study of the molecular mechanisms of biological phenome ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecular Biology
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions. The study of chemical and physical structure of biological macromolecules is known as molecular biology. Molecular biology was first described as an approach focused on the underpinnings of biological phenomena - uncovering the structures of biological molecules as well as their interactions, and how these interactions explain observations of classical biology. In 1945 the term molecular biology was used by physicist William Astbury. In 1953 Francis Crick, James Watson, Rosalind Franklin, and colleagues, working at Medical Research Council unit, Cavendish laboratory, Cambridge (now the MRC Laboratory of Molecular Biology), made a double helix model of DNA which changed the entire research scenario. They proposed the DNA structure based on previous research done by ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |