|
RNA-directed DNA Methylation
RNA-directed DNA methylation (RdDM) is a biological process in which non-coding RNA molecules direct the addition of DNA methylation to specific DNA sequences. The RdDM pathway is unique to plants, although other mechanisms of RNA-directed chromatin modification have also been described in fungi and animals. To date, the RdDM pathway is best characterized within angiosperms (flowering plants), and particularly within the model plant ''Arabidopsis thaliana''. However, conserved RdDM pathway components and associated small RNAs (sRNAs) have also been found in other groups of plants, such as gymnosperms and ferns. The RdDM pathway closely resembles other sRNA pathways, particularly the highly conserved RNAi pathway found in fungi, plants, and animals. Both the RdDM and RNAi pathways produce sRNAs and involve conserved Argonaute, Dicer and RNA-dependent RNA polymerase proteins. RdDM has been implicated in a number of regulatory processes in plants. The DNA methylation added by RdDM ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RdDM Biological Functions
RNA-directed DNA methylation (RdDM) is a biological process in which non-coding RNA molecules direct the addition of DNA methylation to specific DNA sequences. The RdDM pathway is unique to plants, although other mechanisms of RNA-directed chromatin modification have also been described in fungi and animals. To date, the RdDM pathway is best characterized within angiosperms (flowering plants), and particularly within the model plant ''Arabidopsis thaliana''. However, conserved RdDM pathway components and associated small RNAs (sRNAs) have also been found in other groups of plants, such as gymnosperms and ferns. The RdDM pathway closely resembles other sRNA pathways, particularly the highly conserved RNAi pathway found in fungi, plants, and animals. Both the RdDM and RNAi pathways produce sRNAs and involve conserved Argonaute, Dicer and RNA-dependent RNA polymerase proteins. RdDM has been implicated in a number of regulatory processes in plants. The DNA methylation added by RdDM is ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Pathogen
In biology, a pathogen ( el, πάθος, "suffering", "passion" and , "producer of") in the oldest and broadest sense, is any organism or agent that can produce disease. A pathogen may also be referred to as an infectious agent, or simply a germ. The term ''pathogen'' came into use in the 1880s. Typically, the term ''pathogen'' is used to describe an ''infectious'' microorganism or agent, such as a virus, bacterium, protozoan, prion, viroid, or fungus. Small animals, such as helminths and insects, can also cause or transmit disease. However, these animals are usually referred to as parasites rather than pathogens. The scientific study of microscopic organisms, including microscopic pathogenic organisms, is called microbiology, while parasitology refers to the scientific study of parasites and the organisms that host them. There are several pathways through which pathogens can invade a host. The principal pathways have different episodic time frames, but soil has the longest ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Allele
An allele (, ; ; modern formation from Greek ἄλλος ''állos'', "other") is a variation of the same sequence of nucleotides at the same place on a long DNA molecule, as described in leading textbooks on genetics and evolution. ::"The chromosomal or genomic location of a gene or any other genetic element is called a locus (plural: loci) and alternative DNA sequences at a locus are called alleles." The simplest alleles are single nucleotide polymorphisms (SNP). but they can also be insertions and deletions of up to several thousand base pairs. Popular definitions of 'allele' typically refer only to different alleles within genes. For example, the ABO blood grouping is controlled by the ABO gene, which has six common alleles (variants). In population genetics, nearly every living human's phenotype for the ABO gene is some combination of just these six alleles. Most alleles observed result in little or no change in the function of the gene product it codes for. However, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Post-transcriptional Gene Silencing
RNA interference (RNAi) is a biological process in which RNA molecules are involved in sequence-specific suppression of gene expression by double-stranded RNA, through translational or transcriptional repression. Historically, RNAi was known by other names, including ''co-suppression'', ''post-transcriptional gene silencing'' (PTGS), and ''quelling''. The detailed study of each of these seemingly different processes elucidated that the identity of these phenomena were all actually RNAi. Andrew Fire and Craig C. Mello shared the 2006 Nobel Prize in Physiology or Medicine for their work on RNAi in the nematode worm ''Caenorhabditis elegans'', which they published in 1998. Since the discovery of RNAi and its regulatory potentials, it has become evident that RNAi has immense potential in suppression of desired genes. RNAi is now known as precise, efficient, stable and better than antisense therapy for gene suppression. Antisense RNA produced intracellularly by an expression vector may ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Retrotransposon
Retrotransposons (also called Class I transposable elements or transposons via RNA intermediates) are a type of genetic component that copy and paste themselves into different genomic locations (transposon) by converting RNA back into DNA through the reverse transcription process using an RNA transposition intermediate. Through reverse transcription, retrotransposons amplify themselves quickly to become abundant in eukaryotic genomes such as maize (49–78%) and humans (42%). They are only present in eukaryotes but share features with retroviruses such as HIV, for example, discontinuous reverse transcriptase-mediated extrachromosomal recombination. These retrotransposons are regulated by a family of short non-coding RNAs termed as PIWI -element induced wimpy testisinteracting RNAs (piRNAs). piRNA is a recently discovered class of ncRNAs, which are in the length range of ~24-32 nucleotides. Initially, piRNAs were described as repeat-associated siRNAs (rasiRNAs) because of thei ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Post-transcriptional Regulation
Post-transcriptional regulation is the control of gene expression at the RNA level. It occurs once the RNA polymerase has been attached to the gene's promoter and is synthesizing the nucleotide sequence. Therefore, as the name indicates, it occurs between the transcription phase and the translation phase of gene expression. These controls are critical for the regulation of many genes across human tissues. It also plays a big role in cell physiology, being implicated in pathologies such as cancer and neurodegenerative diseases. Mechanism After being produced, the stability and distribution of the different transcripts is regulated (post-transcriptional regulation) by means of RNA binding protein (RBP) that control the various steps and rates controlling events such as alternative splicing, nuclear degradation ( exosome), processing, nuclear export (three alternative pathways), sequestration in P-bodies for storage or degradation and ultimately translation. These proteins ac ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Evolutionary Arms Race
In evolutionary biology, an evolutionary arms race is an ongoing struggle between competing sets of co-evolving genes, phenotypic and behavioral traits that develop escalating adaptations and counter-adaptations against each other, resembling an arms race. These are often described as examples of positive feedback.Dawkins, R. 1996. ''The Blind Watchmaker'' New York: W. W. Norton. Note: This book was also published by Penguin in 1991. While the text is identical, page numbers differ The co-evolving gene sets may be in different species, as in an evolutionary arms race between a predator species and its prey (Vermeij, 1987), or a parasite and its host. Alternatively, the arms race may be between members of the same species, as in the manipulation/sales resistance model of communication (Dawkins & Krebs, 1979) or as in runaway evolution or Red Queen effects. One example of an evolutionary arms race is in sexual conflict between the sexes, often described with the term Fisherian runa ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Helitron (biology)
Helitrons are one of the three groups of eukaryotic class 2 transposable elements (TEs) so far described. They are the eukaryotic rolling-circle transposable elements which are hypothesized to transpose by a rolling circle replication mechanism via a single-stranded DNA intermediate. They were first discovered in plants ('' Arabidopsis thaliana'' and ''Oryza sativa'') and in the nematode '' Caenorhabditis elegans'', and now they have been identified in a diverse range of species, from protists to mammals. Helitrons make up a substantial fraction of many genomes where non-autonomous elements frequently outnumber the putative autonomous partner. Helitrons seem to have a major role in the evolution of host genomes. They frequently capture diverse host genes, some of which can evolve into novel host genes or become essential for Helitron transposition. History Helitrons were the first group of TEs to be discovered by computational analysis of whole genome sequences. The first Helitron ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
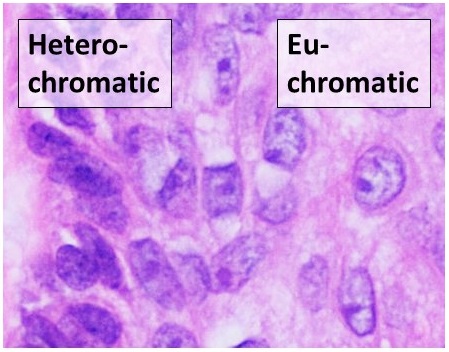
Heterochromatin
Heterochromatin is a tightly packed form of DNA or '' condensed DNA'', which comes in multiple varieties. These varieties lie on a continue between the two extremes of constitutive heterochromatin and facultative heterochromatin. Both play a role in the expression of genes. Because it is tightly packed, it was thought to be inaccessible to polymerases and therefore not transcribed; however, according to Volpe et al. (2002), and many other papers since, much of this DNA is in fact transcribed, but it is continuously turned over via RNA-induced transcriptional silencing (RITS). Recent studies with electron microscopy and OsO4 staining reveal that the dense packing is not due to the chromatin. Constitutive heterochromatin can affect the genes near itself (e.g. position-effect variegation). It is usually repetitive and forms structural functions such as centromeres or telomeres, in addition to acting as an attractor for other gene-expression or repression signals. Facultative hete ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Euchromatin
Euchromatin (also called "open chromatin") is a lightly packed form of chromatin ( DNA, RNA, and protein) that is enriched in genes, and is often (but not always) under active transcription. Euchromatin stands in contrast to heterochromatin, which is tightly packed and less accessible for transcription. 92% of the human genome is euchromatic. In eukaryotes, euchromatin comprises the most active portion of the genome within the cell nucleus. In prokaryotes, euchromatin is the ''only'' form of chromatin present; this indicates that the heterochromatin structure evolved later along with the nucleus, possibly as a mechanism to handle increasing genome size. Structure Euchromatin is composed of repeating subunits known as nucleosomes, reminiscent of an unfolded set of beads on a string, that are approximately 11 nm in diameter. At the core of these nucleosomes are a set of four histone protein pairs: H3, H4, H2A, and H2B. Each core histone protein possesses a 'tail' str ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genome Instability
Genome instability (also genetic instability or genomic instability) refers to a high frequency of mutations within the genome of a cellular lineage. These mutations can include changes in nucleic acid sequences, chromosomal rearrangements or aneuploidy. Genome instability does occur in bacteria. In multicellular organisms genome instability is central to carcinogenesis, and in humans it is also a factor in some neurodegenerative diseases such as amyotrophic lateral sclerosis or the neuromuscular disease myotonic dystrophy. The sources of genome instability have only recently begun to be elucidated. A high frequency of externally caused DNA damage can be one source of genome instability since DNA damage can cause inaccurate translesion DNA synthesis past the damage or errors in repair, leading to mutation. Another source of genome instability may be epigenetic or mutational reductions in expression of DNA repair genes. Because endogenous (metabolically-caused) DNA damage is very fr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Wheat
Wheat is a grass widely cultivated for its seed, a cereal grain that is a worldwide staple food. The many species of wheat together make up the genus ''Triticum'' ; the most widely grown is common wheat (''T. aestivum''). The archaeological record suggests that wheat was first cultivated in the regions of the Fertile Crescent around 9600 BCE. Botanically, the wheat kernel is a type of fruit called a caryopsis. Wheat is grown on more land area than any other food crop (, 2014). World trade in wheat is greater than for all other crops combined. In 2020, world production of wheat was , making it the second most-produced cereal after maize. Since 1960, world production of wheat and other grain crops has tripled and is expected to grow further through the middle of the 21st century. Global demand for wheat is increasing due to the unique viscoelastic and adhesive properties of gluten proteins, which facilitate the production of processed foods, whose consumption is inc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |