|
P-bodies
P-bodies, or processing bodies are distinct foci formed by phase separation within the cytoplasm of the eukaryotic cell consisting of many enzymes involved in mRNA turnover. P-bodies are highly conserved structures and have been observed in somatic cells originating from vertebrates and invertebrates, plants and yeast. To date, P-bodies have been demonstrated to play fundamental roles in general mRNA decay, nonsense-mediated mRNA decay, adenylate-uridylate-rich element mediated mRNA decay, and microRNA (miRNA) induced mRNA silencing. Not all mRNAs which enter P-bodies are degraded, as it has been demonstrated that some mRNAs can exit P-bodies and re-initiate translation. Purification and sequencing of the mRNA from purified processing bodies showed that these mRNAs are largely translationally repressed upstream of translation initiation and are protected from 5' mRNA decay. P-bodies are involved in decapping and degradation of unwanted mRNAs, storing mRNA until needed for tra ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Stress Granules
Stress granules are dense aggregations in the cytosol composed of proteins and RNAs that appear when the cell is under stress. The RNA molecules stored are stalled translation pre-initiation complexes: failed attempts to make protein from mRNA. Stress granules are 100–200 nm in size (when biochemically purified), not surrounded by membrane, and associated with the endoplasmatic reticulum. Note that there are also nuclear stress granules. This article is about the cytosolic variety. Proposed functions The function of stress granules remains largely unknown. Stress granules have long been proposed to have a function to protect RNAs from harmful conditions, thus their appearance under stress. The accumulation of RNAs into dense globules could keep them from reacting with harmful chemicals and safeguard the information coded in their RNA sequence. Stress granules might also function as a decision point for untranslated mRNAs. Molecules can go down one of three paths: further ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Stress Granule
Stress granules are dense aggregations in the cytosol composed of proteins and RNAs that appear when the cell is under stress. The RNA molecules stored are stalled translation pre-initiation complexes: failed attempts to make protein from mRNA. Stress granules are 100–200 nm in size (when biochemically purified), not surrounded by membrane, and associated with the endoplasmatic reticulum. Note that there are also nuclear stress granules. This article is about the cytosolic variety. Proposed functions The function of stress granules remains largely unknown. Stress granules have long been proposed to have a function to protect RNAs from harmful conditions, thus their appearance under stress. The accumulation of RNAs into dense globules could keep them from reacting with harmful chemicals and safeguard the information coded in their RNA sequence. Stress granules might also function as a decision point for untranslated mRNAs. Molecules can go down one of three paths: further ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Messenger RNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein. mRNA is created during the process of transcription, where an enzyme (RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and, utilising amino acids carried by transfer RNA (tRNA), the ribosome creates the protein. This process is known as translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genetic information in a biological system. As in DNA, genetic inf ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nonsense-mediated Decay
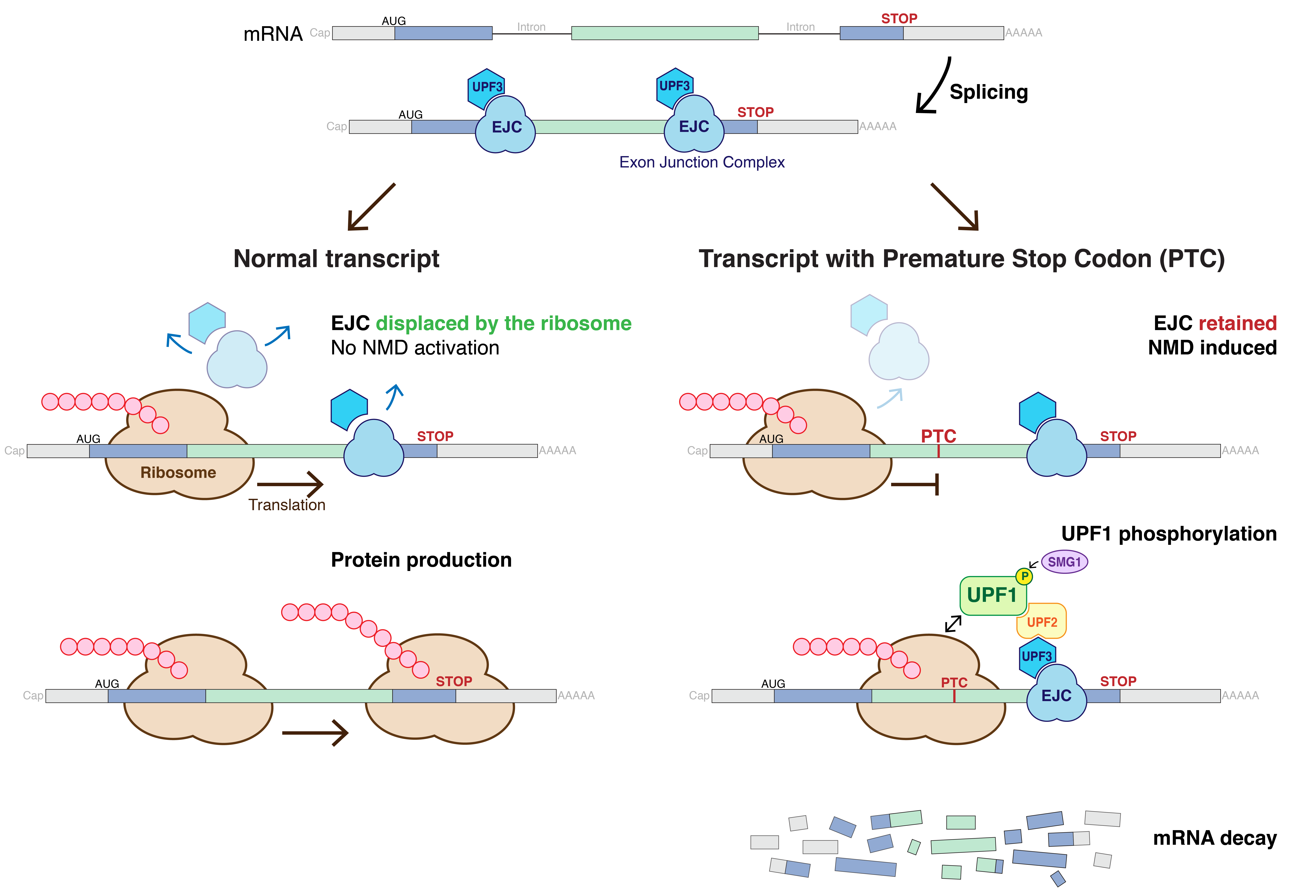
Nonsense-mediated mRNA decay (NMD) is a surveillance pathway that exists in all eukaryotes. Its main function is to reduce errors in gene expression by eliminating mRNA transcripts that contain premature stop codons. Translation of these aberrant mRNAs could, in some cases, lead to deleterious gain-of-function or dominant-negative activity of the resulting proteins. NMD was first described in human cells and in yeast almost simultaneously in 1979. This suggested broad phylogenetic conservation and an important biological role of this intriguing mechanism. NMD was discovered when it was realized that cells often contain unexpectedly low concentrations of mRNAs that are transcribed from alleles carrying nonsense mutations. Nonsense mutations code for a premature stop codon which causes the protein to be shortened. The truncated protein may or may not be functional, depending on the severity of what is not translated. In human genetics, NMD has the possibility to not only limit the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MicroRNA
MicroRNA (miRNA) are small, single-stranded, non-coding RNA molecules containing 21 to 23 nucleotides. Found in plants, animals and some viruses, miRNAs are involved in RNA silencing and post-transcriptional regulation of gene expression. miRNAs base-pair to complementary sequences in mRNA molecules, then gene silence said mRNA molecules by one or more of the following processes: (1) cleavage of mRNA strand into two pieces, (2) destabilization of mRNA by shortening its poly(A) tail, or (3) translation of mRNA into proteins. This last method of gene silencing is the least efficient of the three, and requires the aid of ribosomes. miRNAs resemble the small interfering RNAs (siRNAs) of the RNA interference (RNAi) pathway, except miRNAs derive from regions of RNA transcripts that fold back on themselves to form short hairpins, whereas siRNAs derive from longer regions of double-stranded RNA. The human genome may encode over 1900 miRNAs, although more recent analysis suggests that ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Messenger RNA Decapping
The process of messenger RNA decapping consists of hydrolysis of the 5' cap structure on the RNA exposing a 5' monophosphate. In eukaryotes, this 5' monophosphate is a substrate for the 5' exonuclease Xrn1 and the mRNA is quickly destroyed. There are many situations which may lead to the removal of the cap, some of which are discussed below. In prokaryotes, the initial mRNA transcript naturally possesses a 5'-triphosphate group after bacterial transcription; the enzyme RppH removes a pyrophosphate molecule from the 5' end, converting the 5'-triphosphate to a 5'-monophosphate, triggering mRNA degradation by ribonucleases. Translation and decay Inside cells, there is a balance between the processes of translation and mRNA decay. Messages which are being actively translated are bound by polysomes and the eukaryotic initiation factors eIF-4E and eIF-4G (in eukaryotes). This blocks access to the cap by the decapping enzyme DCP2 and protects the mRNA molecule. In nutrient-star ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA-induced Silencing Complex
The RNA-induced silencing complex, or RISC, is a multiprotein complex, specifically a ribonucleoprotein, which functions in gene silencing via a variety of pathways at the transcriptional and translational levels. Using single-stranded RNA (ssRNA) fragments, such as microRNA (miRNA), or double-stranded small interfering RNA (siRNA), the complex functions as a key tool in gene regulation. The single strand of RNA acts as a template for RISC to recognize complementary messenger RNA (mRNA) transcript. Once found, one of the proteins in RISC, Argonaute, activates and cleaves the mRNA. This process is called RNA interference (RNAi) and it is found in many eukaryotes; it is a key process in defense against viral infections, as it is triggered by the presence of double-stranded RNA (dsRNA). Discovery The biochemical identification of RISC was conducted by Gregory Hannon and his colleagues at the Cold Spring Harbor Laboratory. This was only a couple of years after the discovery of RNA ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Fluorescence-activated Cell Sorting
Flow cytometry (FC) is a technique used to detect and measure physical and chemical characteristics of a population of cells or particles. In this process, a sample containing cells or particles is suspended in a fluid and injected into the flow cytometer instrument. The sample is focused to ideally flow one cell at a time through a laser beam, where the light scattered is characteristic to the cells and their components. Cells are often labeled with fluorescent markers so light is absorbed and then emitted in a band of wavelengths. Tens of thousands of cells can be quickly examined and the data gathered are processed by a computer. Flow cytometry is routinely used in basic research, clinical practice, and clinical trials. Uses for flow cytometry include: * Cell counting * Cell sorting * Determining cell characteristics and function * Detecting microorganisms * Biomarker detection * Protein engineering detection * Diagnosis of health disorders such as blood cancers * Measuring ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Streptavidin
Streptavidin is a 66.0 (tetramer) kDa protein purified from the bacterium '' Streptomyces avidinii''. Streptavidin homo-tetramers have an extraordinarily high affinity for biotin (also known as vitamin B7 or vitamin H). With a dissociation constant (Kd) on the order of ≈10−14 mol/L, the binding of biotin to streptavidin is one of the strongest non-covalent interactions known in nature. Streptavidin is used extensively in molecular biology and bionanotechnology due to the streptavidin-biotin complex's resistance to organic solvents, denaturants (e.g. guanidinium chloride), detergents (e.g. SDS, Triton X-100), proteolytic enzymes, and extremes of temperature and pH. Structure The crystal structure of streptavidin with biotin bound was reported by two groups in 1989. The structure was solved using multi wavelength anomalous diffraction by Hendrickson et al. at Columbia University and using multiple isomorphous replacement by Weber et al. at E. I. DuPont Central Research ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Biotinylation
In biochemistry, biotinylation is the process of covalently attaching biotin to a protein, nucleic acid or other molecule. Biotinylation is rapid, specific and is unlikely to disturb the natural function of the molecule due to the small size of biotin (MW = 244.31 g/mol). Biotin binds to streptavidin and avidin with an extremely high affinity, fast on-rate, and high specificity, and these interactions are exploited in many areas of biotechnology to isolate biotinylated molecules of interest. Biotin-binding to streptavidin and avidin is resistant to extremes of heat, pH and proteolysis, making capture of biotinylated molecules possible in a wide variety of environments. Also, multiple biotin molecules can be conjugated to a protein of interest, which allows binding of multiple streptavidin, avidin or neutravidin protein molecules and increases the sensitivity of detection of the protein of interest. There is a large number of biotinylation reagents available that exploit t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Biotin
Biotin (or vitamin B7) is one of the B vitamins. It is involved in a wide range of metabolic processes, both in humans and in other organisms, primarily related to the utilization of fats, carbohydrates, and amino acids. The name ''biotin'', borrowed from the German , derives from the Ancient Greek word (; 'life') and the suffix "-in" (a suffix used in chemistry usually to indicate 'forming'). Chemical description Biotin is classified as a heterocyclic compound, with a sulfur-containing ring fused ureido and tetrahydrothiophene group. A C5-carboxylic acid side chain is appended to one of the rings. The ureido ring, containing the −N−CO−N− group, serves as the carbon dioxide carrier in carboxylation reactions. Biotin is a coenzyme for five carboxylase enzymes, which are involved in the catabolism of amino acids and fatty acids, synthesis of fatty acids, and gluconeogenesis. Biotinylation of histone proteins in nuclear chromatin plays a role in chromatin stability and g ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Proximity Labeling
Enzyme-catalyzed proximity labeling (PL), also known as proximity-based labeling, is a laboratory technique that labels biomolecules, usually proteins or RNA, proximal to a protein of interest. By creating a gene fusion in a living cell between the protein of interest and an engineered labeling enzyme, biomolecules spatially proximal to the protein of interest can then be selectively marked with biotin for pulldown and analysis. Proximity labeling has been used for identifying the components of novel cellular structures and for determining protein-protein interaction partners, among other applications. History Before the development of proximity labeling, determination of protein proximity in cells relied on studying protein-protein interactions through methods such as affinity purification-mass spectrometry and proximity ligation assays. DamID is a method developed in 2000 by Steven Henikoff for identifying parts of the genome proximal to a chromatin protein of interest. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |

.jpg)


