|
ORF8a
ORF8 is a gene that encodes a viral accessory protein, Betacoronavirus NS8 protein, in coronaviruses of the subgenus ''Sarbecovirus''. It is one of the least well conserved and most variable parts of the genome. In some viruses, a deletion splits the region into two smaller open reading frames, called ORF8a and ORF8b - a feature present in many SARS-CoV viral isolates from later in the SARS epidemic, as well as in some bat coronaviruses. For this reason the full-length gene and its protein are sometimes called ORF8ab. The full-length gene, exemplified in SARS-CoV-2, encodes a protein with an immunoglobulin domain of unknown function, possibly involving interactions with the host immune system. It is similar in structure to the ORF7a protein, suggesting it may have originated through gene duplication. Structure ORF8 in SARS-CoV-2 encodes a protein of 121 amino acid residues with an N-terminal signal sequence. ORF8 forms a dimer that is covalently linked by disulfide bonds. It h ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sarbecovirus
''Severe acute respiratory syndrome–related coronavirus'' (SARSr-CoV or SARS-CoV)The terms ''SARSr-CoV'' and ''SARS-CoV'' are sometimes used interchangeably, especially prior to the discovery of SARS-CoV-2. This may cause confusion when some publications refer to SARS-CoV-1 as ''SARS-CoV''. is a species of virus consisting of many known strains phylogenetically related to severe acute respiratory syndrome coronavirus 1 (SARS-CoV-1) that have been shown to possess the capability to infect humans, bats, and certain other mammals. These enveloped, positive-sense single-stranded RNA viruses enter host cells by binding to the angiotensin-converting enzyme 2 (ACE2) receptor. The SARSr-CoV species is a member of the genus ''Betacoronavirus'' and of the subgenus ''Sarbecovirus'' (SARS Betacoronavirus). Two strains of the virus have caused outbreaks of severe respiratory diseases in humans: severe acute respiratory syndrome coronavirus 1 (SARS-CoV or SARS-CoV-1), which caused the 20 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SARS-CoV-2
Severe acute respiratory syndrome coronavirus 2 (SARS‑CoV‑2) is a strain of coronavirus that causes COVID-19 (coronavirus disease 2019), the respiratory illness responsible for the ongoing COVID-19 pandemic. The virus previously had a provisional name, 2019 novel coronavirus (2019-nCoV), and has also been called the human coronavirus 2019 (HCoV-19 or hCoV-19). First identified in the city of Wuhan, Hubei, China, the World Health Organization declared the outbreak a public health emergency of international concern on January 30, 2020, and a pandemic on March 11, 2020. SARS‑CoV‑2 is a positive-sense single-stranded RNA virus that is contagious in humans. SARS‑CoV‑2 is a virus of the species ''severe acute respiratory syndrome–related coronavirus'' (SARSr-CoV), related to the SARS-CoV-1 virus that caused the 2002–2004 SARS outbreak. Despite its close relation to SARS-CoV-1, its closest known relatives, with which it forms a sister group, are the derived SARS ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
ORF7a
ORF7a (also known by several other names, including SARS coronavirus X4, SARS-X4, ORF7a, or U122) is a gene found in coronaviruses of the ''Betacoronavirus'' genus. It gene expression, expresses the Betacoronavirus NS7A protein, a type I transmembrane protein with an immunoglobulin domain, immunoglobulin-like protein domain. It was first discovered in SARS-CoV, the virus that causes severe acute respiratory syndrome (SARS). The homology (biology), homolog in SARS-CoV-2, the virus that causes COVID-19, has about 85% sequence identity to the SARS-CoV protein. Function A number of possible functions for the ORF7a protein have been described. The primary function is thought to be immunomodulation and interferon antagonism. The protein is not essential gene, essential for viral replication. Viral protein interactions Studies in SARS-CoV suggest that the protein forms protein-protein interactions with coronavirus spike protein, spike protein and ORF3a, and is present in mature virions, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cysteine
Cysteine (symbol Cys or C; ) is a semiessential proteinogenic amino acid with the formula . The thiol side chain in cysteine often participates in enzymatic reactions as a nucleophile. When present as a deprotonated catalytic residue, sometimes the symbol Cyz is used. The deprotonated form can generally be described by the symbol Cym as well. The thiol is susceptible to oxidation to give the disulfide derivative cystine, which serves an important structural role in many proteins. In this case, the symbol Cyx is sometimes used. When used as a food additive, it has the E number E920. Cysteine is encoded by the codons UGU and UGC. The sulfur-containing amino acids cysteine and methionine are more easily oxidized than the other amino acids. Structure Like other amino acids (not as a residue of a protein), cysteine exists as a zwitterion. Cysteine has chirality in the older / notation based on homology to - and -glyceraldehyde. In the newer ''R''/''S'' system of designating chi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Complex
A protein complex or multiprotein complex is a group of two or more associated polypeptide chains. Protein complexes are distinct from multienzyme complexes, in which multiple catalytic domains are found in a single polypeptide chain. Protein complexes are a form of quaternary structure. Proteins in a protein complex are linked by non-covalent protein–protein interactions. These complexes are a cornerstone of many (if not most) biological processes. The cell is seen to be composed of modular supramolecular complexes, each of which performs an independent, discrete biological function. Through proximity, the speed and selectivity of binding interactions between enzymatic complex and substrates can be vastly improved, leading to higher cellular efficiency. Many of the techniques used to enter cells and isolate proteins are inherently disruptive to such large complexes, complicating the task of determining the components of a complex. Examples of protein complexes include the p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequence Identity
In bioinformatics, a sequence alignment is a way of arranging the sequences of DNA, RNA, or protein to identify regions of similarity that may be a consequence of functional, structural, or evolutionary relationships between the sequences. Aligned sequences of nucleotide or amino acid residues are typically represented as rows within a matrix. Gaps are inserted between the residues so that identical or similar characters are aligned in successive columns. Sequence alignments are also used for non-biological sequences, such as calculating the distance cost between strings in a natural language or in financial data. Interpretation If two sequences in an alignment share a common ancestor, mismatches can be interpreted as point mutations and gaps as indels (that is, insertion or deletion mutations) introduced in one or both lineages in the time since they diverged from one another. In sequence alignments of proteins, the degree of similarity between amino acids occupying a part ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transmembrane Protein
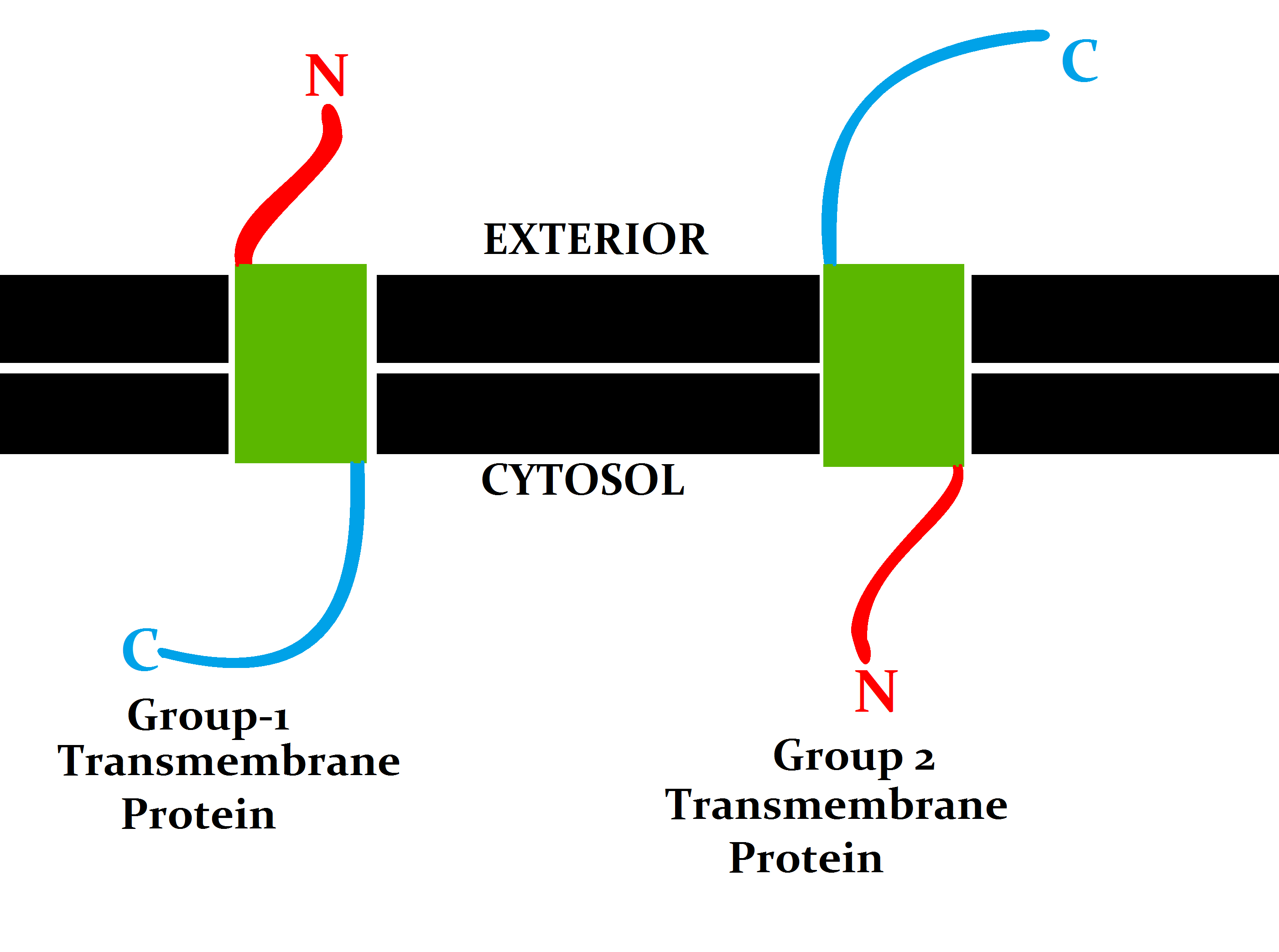
A transmembrane protein (TP) is a type of integral membrane protein that spans the entirety of the cell membrane. Many transmembrane proteins function as gateways to permit the transport of specific substances across the membrane. They frequently undergo significant conformational changes to move a substance through the membrane. They are usually highly hydrophobic and aggregate and precipitate in water. They require detergents or nonpolar solvents for extraction, although some of them (beta-barrels) can be also extracted using denaturing agents. The peptide sequence that spans the membrane, or the transmembrane segment, is largely hydrophobic and can be visualized using the hydropathy plot. Depending on the number of transmembrane segments, transmembrane proteins can be classified as single-span (or bitopic) or multi-span (polytopic). Some other integral membrane proteins are called monotopic, meaning that they are also permanently attached to the membrane, but do not pass ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transmembrane Helix
A transmembrane domain (TMD) is a membrane-spanning protein domain. TMDs generally adopt an alpha helix topological conformation, although some TMDs such as those in porins can adopt a different conformation. Because the interior of the lipid bilayer is hydrophobic, the amino acid residues in TMDs are often hydrophobic, although proteins such as membrane pumps and ion channels can contain polar residues. TMDs vary greatly in length, sequence, and hydrophobicity, adopting organelle-specific properties. Functions of transmembrane domains Transmembrane domains are known to perform a variety of functions. These include: * Anchoring transmembrane proteins to the membrane. *Facilitating molecular transport of molecules such as ions and proteins across biological membranes; usually hydrophilic residues and binding sites in the TMDs help in this process. * Signal transduction across the membrane; many transmembrane proteins, such as G protein-coupled receptors, receive extracellul ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
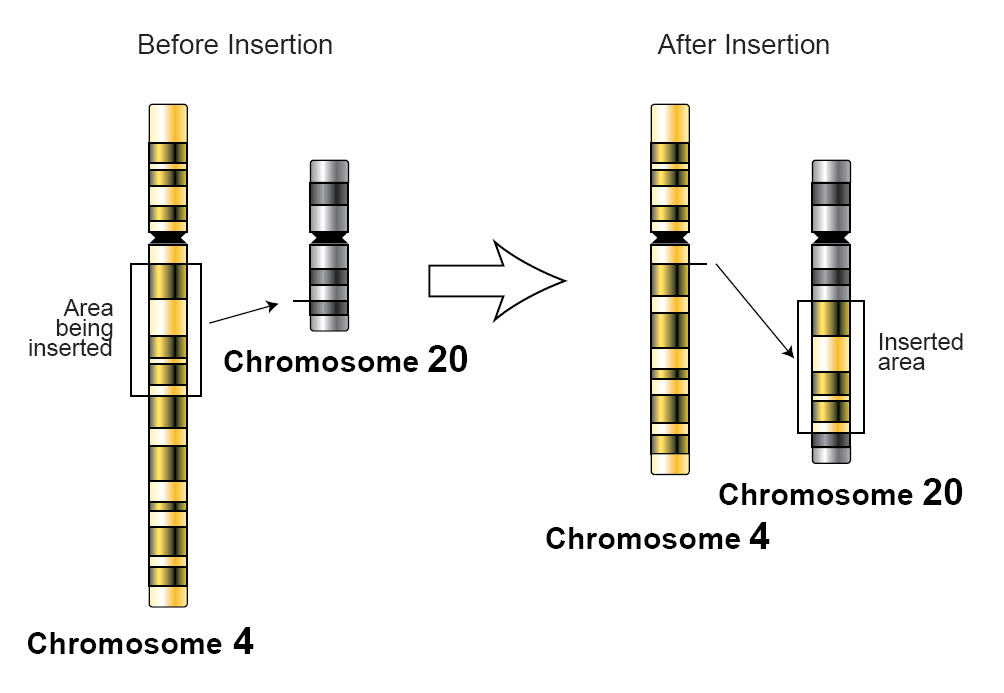
Insertion Mutation
In genetics, an insertion (also called an insertion mutation) is the addition of one or more nucleotide base pairs into a DNA sequence. This can often happen in microsatellite regions due to the DNA polymerase slipping. Insertions can be anywhere in size from one base pair incorrectly inserted into a DNA sequence to a section of one chromosome inserted into another. The mechanism of the smallest single base insertion mutations is believed to be through base-pair separation between the template and primer strands followed by non-neighbor base stacking, which can occur locally within the DNA polymerase active site. On a chromosome level, an ''insertion'' refers to the insertion of a larger sequence into a chromosome. This can happen due to unequal crossover during meiosis. N region addition is the addition of non-coded nucleotides during recombination by terminal deoxynucleotidyl transferase. P nucleotide insertion is the insertion of palindromic sequences encoded by the ends ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Domain
In molecular biology, a protein domain is a region of a protein's polypeptide chain that is self-stabilizing and that folds independently from the rest. Each domain forms a compact folded three-dimensional structure. Many proteins consist of several domains, and a domain may appear in a variety of different proteins. Molecular evolution uses domains as building blocks and these may be recombined in different arrangements to create proteins with different functions. In general, domains vary in length from between about 50 amino acids up to 250 amino acids in length. The shortest domains, such as zinc fingers, are stabilized by metal ions or disulfide bridges. Domains often form functional units, such as the calcium-binding EF hand domain of calmodulin. Because they are independently stable, domains can be "swapped" by genetic engineering between one protein and another to make chimeric proteins. Background The concept of the domain was first proposed in 1973 by Wetlaufer aft ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Covalent
A covalent bond is a chemical bond that involves the sharing of electrons to form electron pairs between atoms. These electron pairs are known as shared pairs or bonding pairs. The stable balance of attractive and repulsive forces between atoms, when they share electrons, is known as covalent bonding. For many molecules, the sharing of electrons allows each atom to attain the equivalent of a full valence shell, corresponding to a stable electronic configuration. In organic chemistry, covalent bonding is much more common than ionic bonding. Covalent bonding also includes many kinds of interactions, including σ-bonding, π-bonding, metal-to-metal bonding, agostic interactions, bent bonds, three-center two-electron bonds and three-center four-electron bonds. The term ''covalent bond'' dates from 1939. The prefix ''co-'' means ''jointly, associated in action, partnered to a lesser degree, '' etc.; thus a "co-valent bond", in essence, means that the atoms share " valence", such a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |





.png)