|
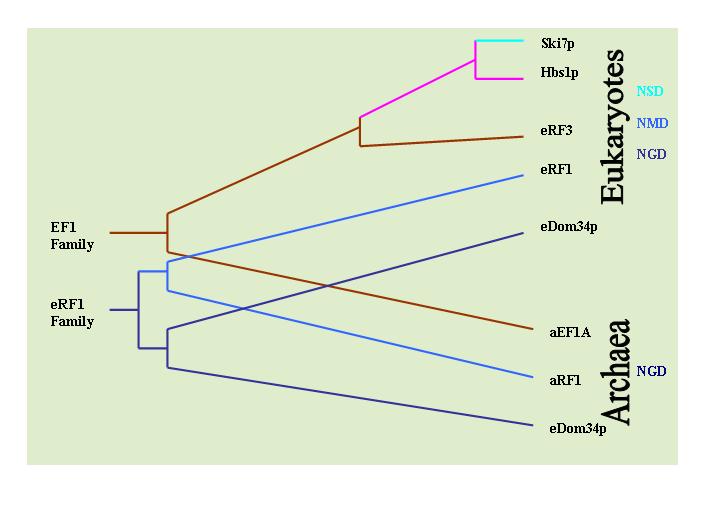
Non-stop Decay
Non-stop decay (NSD) is a cellular mechanism of mRNA surveillance to detect mRNA molecules lacking a stop codon and prevent these mRNAs from translation. The non-stop decay pathway releases ribosomes that have reached the far 3' end of an mRNA and guides the mRNA to the exosome complex, or to RNase R in bacteria for selective degradation. In contrast to Nonsense-mediated decay (NMD), polypeptides do not release from the ribosome, and thus, NSD seems to involve mRNA decay factors distinct from NMD. Non-Stop Decay (NSD) Non-stop decay (NSD) is a cellular pathway that identifies and degrades aberrant mRNA transcripts that do not contain a proper stop codon. Stop codons are signals in messenger RNA that signal for synthesis of proteins to end. Aberrant transcripts are identified during translation when the ribosome translates into the poly A tail at the 3' end of mRNA. A non-stop transcript can occur when point mutations damage the normal stop codon. Moreover, some transcriptional ev ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cell (biology)
The cell is the basic structural and functional unit of life forms. Every cell consists of a cytoplasm enclosed within a membrane, and contains many biomolecules such as proteins, DNA and RNA, as well as many small molecules of nutrients and metabolites.Cell Movements and the Shaping of the Vertebrate Body in Chapter 21 of Molecular Biology of the Cell '' fourth edition, edited by Bruce Alberts (2002) published by Garland Science. The Alberts text discusses how the "cellular building blocks" move to shape developing embryos. It is also common to describe small molecules such as ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein. mRNA is created during the process of Transcription (biology), transcription, where an enzyme (RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and, utilising amino acids carried by transfer RNA (tRNA), the ribosome creates the protein. This process is known as Translation (biology), translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genet ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Stop Codon
In molecular biology (specifically protein biosynthesis), a stop codon (or termination codon) is a codon (nucleotide triplet within messenger RNA) that signals the termination of the translation process of the current protein. Most codons in messenger RNA correspond to the addition of an amino acid to a growing polypeptide chain, which may ultimately become a protein; stop codons signal the termination of this process by binding release factors, which cause the ribosomal subunits to disassociate, releasing the amino acid chain. While start codons need nearby sequences or initiation factors to start translation, a stop codon alone is sufficient to initiate termination. Properties Standard codons In the standard genetic code, there are three different termination codons: Alternative stop codons There are variations on the standard genetic code, and alternative stop codons have been found in the mitochondrial genomes of vertebrates, ''Scenedesmus obliquus'', and '' Thra ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
3' End
Directionality, in molecular biology and biochemistry, is the end-to-end chemical orientation of a single strand of nucleic acid. In a single strand of DNA or RNA, the chemical convention of naming carbon atoms in the nucleotide pentose-sugar-ring means that there will be a 5′ end (usually pronounced "five-prime end"), which frequently contains a phosphate group attached to the 5′ carbon of the ribose ring, and a 3′ end (usually pronounced "three-prime end"), which typically is unmodified from the ribose -OH substituent. In a DNA double helix, the strands run in opposite directions to permit base pairing between them, which is essential for replication or transcription of the encoded information. Nucleic acids can only be synthesized in vivo in the 5′-to-3′ direction, as the polymerases that assemble various types of new strands generally rely on the energy produced by breaking nucleoside triphosphate bonds to attach new nucleoside monophosphates to the 3′- ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Exosome Complex
The exosome complex (or PM/Scl complex, often just called the exosome) is a multi-protein intracellular complex capable of degrading various types of RNA (ribonucleic acid) molecules. Exosome complexes are found in both eukaryotic cells and archaea, while in bacteria a simpler complex called the degradosome carries out similar functions. The core of the exosome contains a six-membered ring structure to which other proteins are attached. In eukaryotic cells, the exosome complex is present in the cytoplasm, nucleus, and especially the nucleolus, although different proteins interact with the exosome complex in these compartments regulating the RNA degradation activity of the complex to substrates specific to these cell compartments. Substrates of the exosome include messenger RNA, ribosomal RNA, and many species of small RNAs. The exosome has an exoribonucleolytic function, meaning it degrades RNA starting at one end (the 3′ end in this case), and in eukaryotes also an endori ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNase R
RNase R, or Ribonuclease R, is a 3'-->5' exoribonuclease, which belongs to the RNase II superfamily, a group of enzymes that hydrolyze RNA in the 3' - 5' direction. RNase R has been shown to be involved in selective mRNA degradation, particularly of non stop mRNAs in bacteria. RNase R has homologues in many other organisms. When a part of another larger protein has a domain that is very similar to RNase R, this is called an RNase R domain. Role in ''trans''-translation and ribosomal quality control RNase R ensures translation accuracy, correct rRNA maturation and elimination of abnormal rRNAs, and is employed by the ''trans''-translation system to break down damaged mRNAs. In ''Escherichia coli,'' RNase R is a 92 kD protein, with the characteristic capacity to degrade structured RNA substrates without displaying sequence specificity. Therefore, RNase R acts over a range of substrates, such as, ribosomal, transfer, messenger and small non-coding RNAs. RNase R is associated with ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
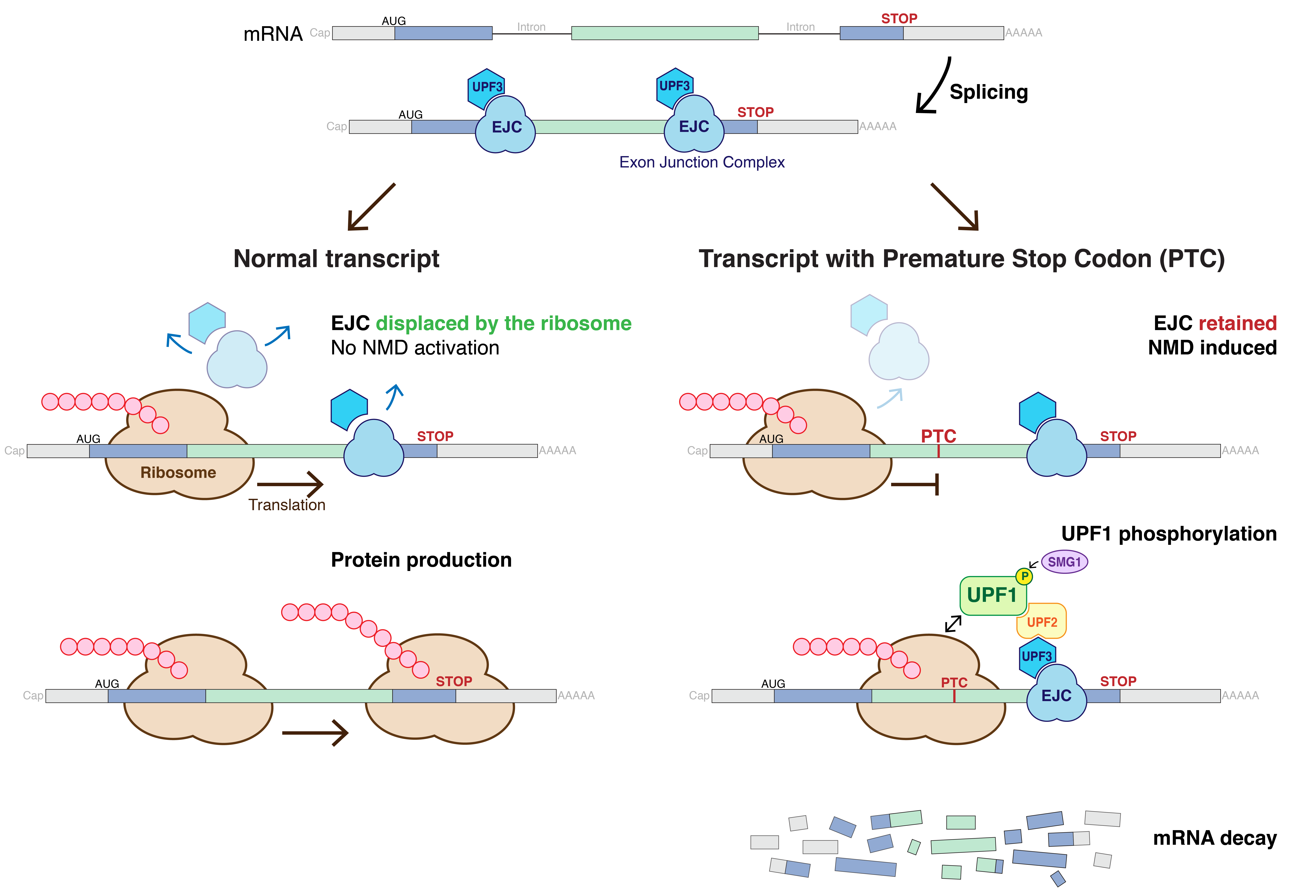
Nonsense-mediated Decay
Nonsense-mediated mRNA decay (NMD) is a surveillance pathway that exists in all eukaryotes. Its main function is to reduce errors in gene expression by eliminating mRNA transcripts that contain premature stop codons. Translation of these aberrant mRNAs could, in some cases, lead to deleterious gain-of-function or dominant-negative activity of the resulting proteins. NMD was first described in human cells and in yeast almost simultaneously in 1979. This suggested broad phylogenetic conservation and an important biological role of this intriguing mechanism. NMD was discovered when it was realized that cells often contain unexpectedly low concentrations of mRNAs that are transcribed from alleles carrying nonsense mutations. Nonsense mutations code for a premature stop codon which causes the protein to be shortened. The truncated protein may or may not be functional, depending on the severity of what is not translated. In human genetics, NMD has the possibility to not only limit the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MRNA Surveillance
mRNA surveillance mechanisms are pathways utilized by organisms to ensure fidelity and quality of messenger RNA (mRNA) molecules. There are a number of surveillance mechanisms present within cells. These mechanisms function at various steps of the mRNA biogenesis pathway to detect and degrade transcripts that have not properly been processed. Overview The translation of messenger RNA transcripts into proteins is a vital part of the central dogma of molecular biology. mRNA molecules are, however, prone to a host of fidelity errors which can cause errors in translation of mRNA into quality proteins. RNA surveillance mechanisms are methods cells use to assure the quality and fidelity of the mRNA molecules. This is generally achieved through marking aberrant mRNA molecule for degradation by various endogenous nucleases. mRNA surveillance has been documented in bacteria and yeast. In eukaryotes, these mechanisms are known to function in both the nucleus and cytoplasm. Fidelity c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Expression
Gene expression is the process by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, protein or non-coding RNA, and ultimately affect a phenotype, as the final effect. These products are often proteins, but in non-protein-coding genes such as transfer RNA (tRNA) and small nuclear RNA (snRNA), the product is a functional non-coding RNA. Gene expression is summarized in the central dogma of molecular biology first formulated by Francis Crick in 1958, further developed in his 1970 article, and expanded by the subsequent discoveries of reverse transcription and RNA replication. The process of gene expression is used by all known life—eukaryotes (including multicellular organisms), prokaryotes (bacteria and archaea), and utilized by viruses—to generate the macromolecular machinery for life. In genetics, gene expression is the most fundamental level at which the genotype gives rise to the phenotype, '' ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |