|
Maturase K
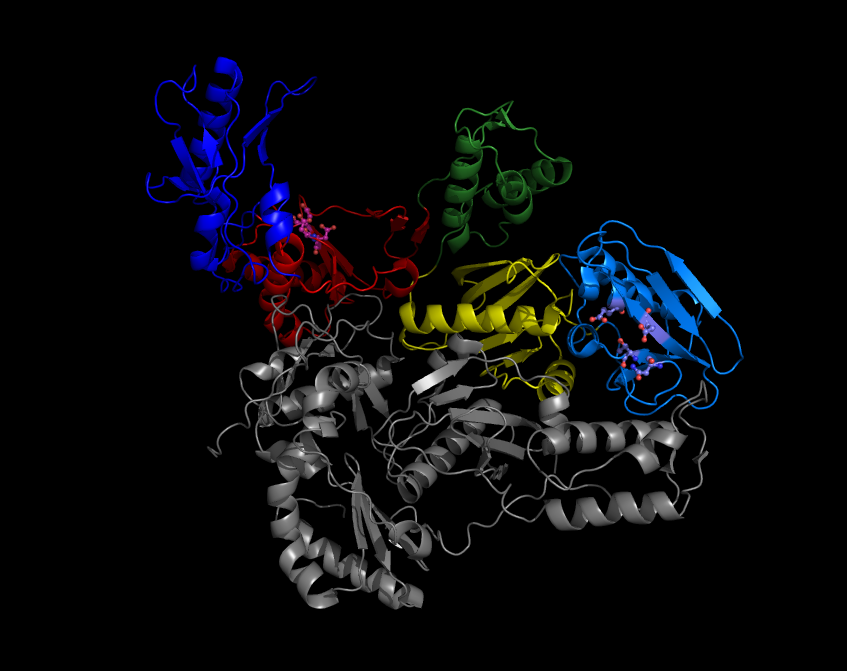
Maturase K (matK) is a plant plastidial gene. The protein it encodes is an organelle intron maturase, a protein that splices Group II introns. It is essential for ''in vivo'' splicing of Group II introns. Amongst other maturases, this protein retains only a well conserved domain X and remnants of a reverse transcriptase domain. Universal matK primers can be used for DNA barcoding DNA barcoding is a method of species identification using a short section of DNA from a specific gene or genes. The premise of DNA barcoding is that by comparison with a reference library of such DNA sections (also called " sequences"), an indi ... of angiosperms. See also * LtrA, an open reading frame found in the ''Lactococcus lactis'' group II introns LtrB. It is an intron-encoded protein, with three subdomains, one of which is a reverse-transcriptase/maturase. References Plant genes {{gene-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Arabidopsis Thaliana
''Arabidopsis thaliana'', the thale cress, mouse-ear cress or arabidopsis, is a small flowering plant native to Eurasia and Africa. ''A. thaliana'' is considered a weed; it is found along the shoulders of roads and in disturbed land. A winter annual with a relatively short lifecycle, ''A. thaliana'' is a popular model organism in plant biology and genetics. For a complex multicellular eukaryote, ''A. thaliana'' has a relatively small genome around 135 megabase pairs. It was the first plant to have its genome sequenced, and is a popular tool for understanding the molecular biology of many plant traits, including flower development and light sensing. Description ''Arabidopsis thaliana'' is an annual (rarely biennial) plant, usually growing to 20–25 cm tall. The leaves form a rosette at the base of the plant, with a few leaves also on the flowering stem. The basal leaves are green to slightly purplish in color, 1.5–5 cm long, and 2–10 mm broad, wit ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chloroplast DNA
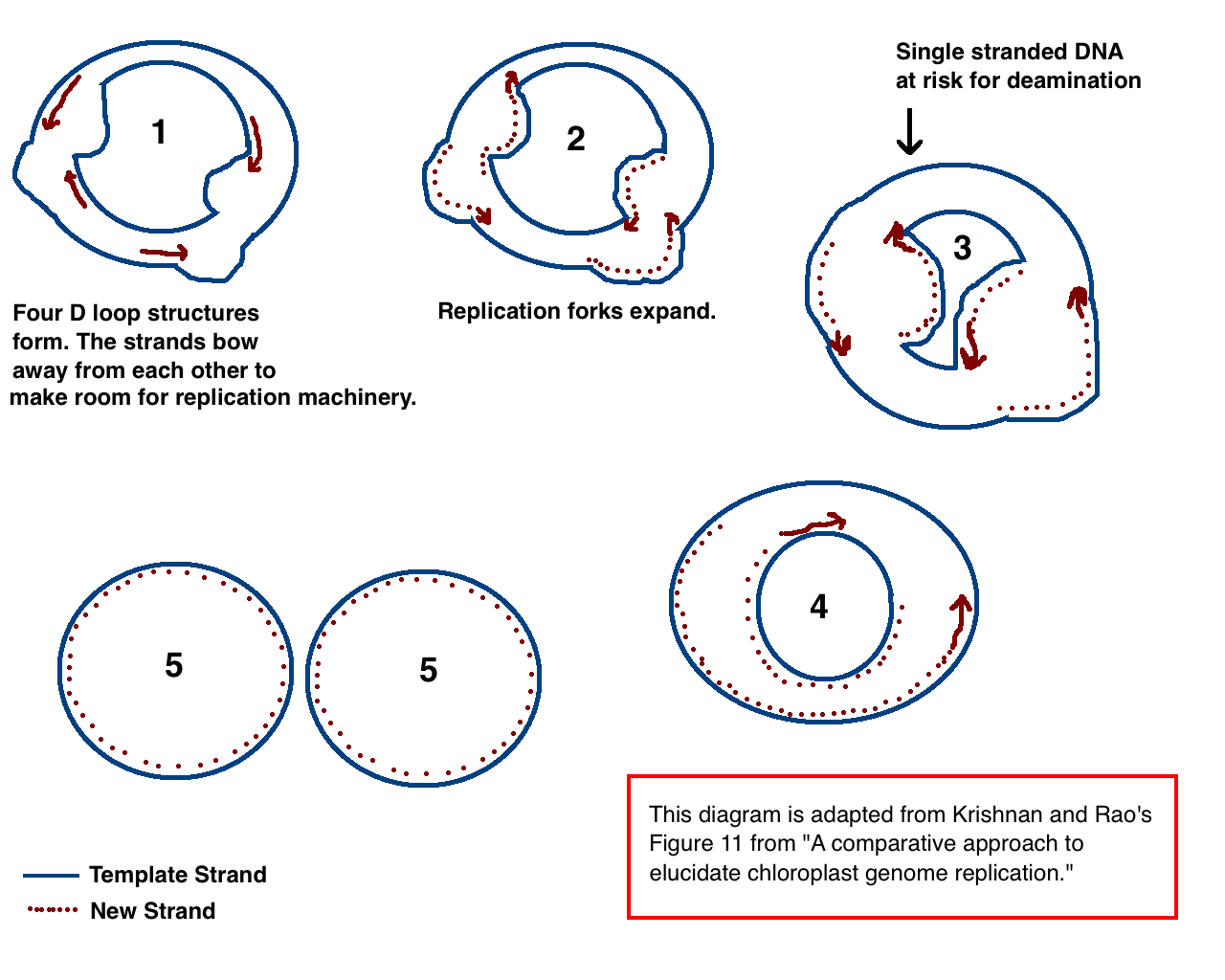
Chloroplast DNA (cpDNA) is the DNA located in chloroplasts, which are photosynthetic organelles located within the cells of some eukaryotic organisms. Chloroplasts, like other types of plastid, contain a genome separate from that in the cell nucleus. The existence of chloroplast DNA was identified biochemically in 1959, and confirmed by electron microscopy in 1962. The discoveries that the chloroplast contains ribosomes and performs protein synthesis revealed that the chloroplast is genetically semi-autonomous. The first complete chloroplast genome sequences were published in 1986, '' Nicotiana tabacum'' (tobacco) by Sugiura and colleagues and '' Marchantia polymorpha'' (liverwort) by Ozeki et al. Since then, a great number of chloroplast DNAs from various species have been sequenced. Molecular structure Chloroplast DNAs are circular, and are typically 120,000–170,000 base pairs long. They can have a contour length of around 30–60 micrometers, and have a mass of about 8 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA Splicing
RNA splicing is a process in molecular biology where a newly-made precursor messenger RNA (pre-mRNA) transcript is transformed into a mature messenger RNA (mRNA). It works by removing all the introns (non-coding regions of RNA) and ''splicing'' back together exons (coding regions). For nuclear-encoded genes, splicing occurs in the nucleus either during or immediately after transcription. For those eukaryotic genes that contain introns, splicing is usually needed to create an mRNA molecule that can be translated into protein. For many eukaryotic introns, splicing occurs in a series of reactions which are catalyzed by the spliceosome, a complex of small nuclear ribonucleoproteins (snRNPs). There exist self-splicing introns, that is, ribozymes that can catalyze their own excision from their parent RNA molecule. The process of transcription, splicing and translation is called gene expression, the central dogma of molecular biology. Splicing pathways Several methods of RNA spl ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Group II Intron
Group II introns are a large class of self-catalytic ribozymes and mobile genetic elements found within the genes of all three domains of life. Ribozyme activity (e.g., self- splicing) can occur under high-salt conditions ''in vitro''. However, assistance from proteins is required for ''in vivo'' splicing. In contrast to group I introns, intron excision occurs in the absence of GTP and involves the formation of a lariat, with an A-residue branchpoint strongly resembling that found in lariats formed during splicing of nuclear pre-mRNA. It is hypothesized that pre-mRNA splicing (see spliceosome) may have evolved from group II introns, due to the similar catalytic mechanism as well as the structural similarity of the Group II Domain V substructure to the U6/U2 extended snRNA. Finally, their ability to site-specifically insert into DNA sites has been exploited as a tool for biotechnology. For example, group II introns can be modified to make site-specific genome insertions and d ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Domain X
Group II introns are a large class of self-catalytic ribozymes and mobile genetic elements found within the genes of all three domains of life. Ribozyme activity (e.g., self- splicing) can occur under high-salt conditions ''in vitro''. However, assistance from proteins is required for ''in vivo'' splicing. In contrast to group I introns, intron excision occurs in the absence of GTP and involves the formation of a lariat, with an A-residue branchpoint strongly resembling that found in lariats formed during splicing of nuclear pre-mRNA. It is hypothesized that pre-mRNA splicing (see spliceosome) may have evolved from group II introns, due to the similar catalytic mechanism as well as the structural similarity of the Group II Domain V substructure to the U6/U2 extended snRNA. Finally, their ability to site-specifically insert into DNA sites has been exploited as a tool for biotechnology. For example, group II introns can be modified to make site-specific genome insertions and deli ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Reverse Transcriptase
A reverse transcriptase (RT) is an enzyme used to generate complementary DNA (cDNA) from an RNA template, a process termed reverse transcription. Reverse transcriptases are used by viruses such as HIV and hepatitis B to replicate their genomes, by retrotransposon mobile genetic elements to proliferate within the host genome, and by eukaryotic cells to extend the telomeres at the ends of their linear chromosomes. Contrary to a widely held belief, the process does not violate the flows of genetic information as described by the classical central dogma, as transfers of information from RNA to DNA are explicitly held possible. Retroviral RT has three sequential biochemical activities: RNA-dependent DNA polymerase activity, ribonuclease H (RNase H), and DNA-dependent DNA polymerase activity. Collectively, these activities enable the enzyme to convert single-stranded RNA into double-stranded cDNA. In retroviruses and retrotransposons, this cDNA can then integrate into the host geno ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Primer (molecular Biology)
Primer may refer to: Arts, entertainment, and media Films * ''Primer'' (film), a 2004 feature film written and directed by Shane Carruth * ''Primer'' (video), a documentary about the funk band Living Colour Literature * Primer (textbook), a textbook used in primary education to teach the alphabet and other basic subjects * Primer (prayer book), a common name for English prayer books used from the 13th to 16th centuries * '' The New England Primer'' (1688), a Puritan book from Colonial America with morality-themed rhymes Music * ''Primer'' (album), a 1995 music album by the musical group Rockapella * Primer 55, an American alternative metal band * "The Primer", a song from the 2005 album ''Alaska'' by Between the Buried and Me Firearms * Primer (firearms), a firearm powder charge-ignition mechanism ** Centerfire ammunition, Boxer or Berdan primers used in modern centerfire cartridges ** Detonator, a small explosive device also known as an explosive primer or blasting cap ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Barcoding
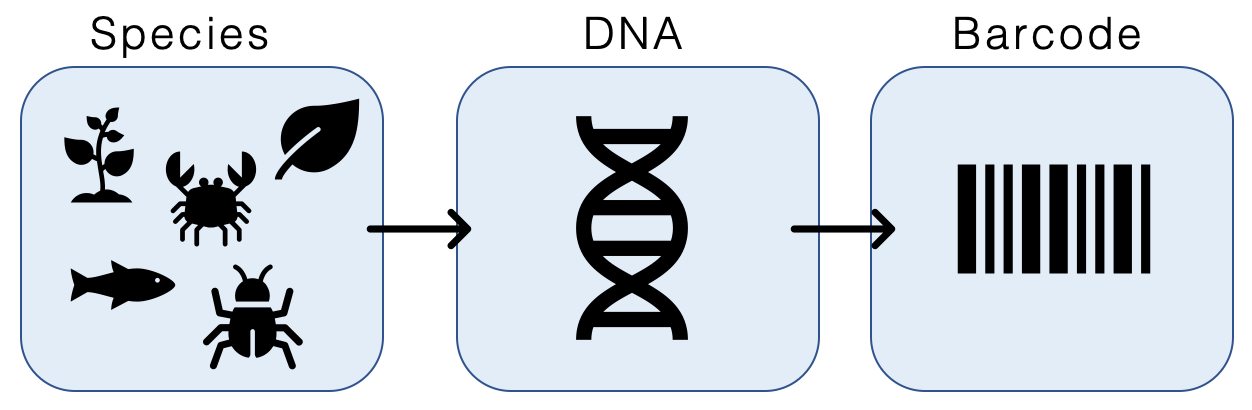
DNA barcoding is a method of species identification using a short section of DNA from a specific gene or genes. The premise of DNA barcoding is that by comparison with a reference library of such DNA sections (also called " sequences"), an individual sequence can be used to uniquely identify an organism to species, just as a supermarket scanner uses the familiar black stripes of the UPC barcode to identify an item in its stock against its reference database. These "barcodes" are sometimes used in an effort to identify unknown species or parts of an organism, simply to catalog as many taxa as possible, or to compare with traditional taxonomy in an effort to determine species boundaries. Different gene regions are used to identify the different organismal groups using barcoding. The most commonly used barcode region for animals and some protists is a portion of the cytochrome ''c'' oxidase I (COI or COX1) gene, found in mitochondrial DNA. Other genes suitable for DNA barcodin ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |