|
MET (gene)
c-Met, also called tyrosine-protein kinase Met or hepatocyte growth factor receptor (HGFR), is a protein that in humans is encoded by the ''MET'' gene. The protein possesses tyrosine kinase activity. The primary single chain precursor protein is post-translationally cleaved to produce the alpha and beta subunits, which are disulfide linked to form the mature receptor. MET is a single pass tyrosine kinase receptor essential for embryonic development, organogenesis and wound healing. Hepatocyte growth factor/Scatter Factor (HGF/SF) and its splicing isoform (NK1, NK2) are the only known ligands of the MET receptor. MET is normally expressed by cells of epithelial origin, while expression of HGF/SF is restricted to cells of mesenchymal origin. When HGF/SF binds its cognate receptor MET it induces its dimerization through a not yet completely understood mechanism leading to its activation. Abnormal MET activation in cancer correlates with poor prognosis, where aberrantly active MET t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residue ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Disulfide Bridge
In biochemistry, a disulfide (or disulphide in British English) refers to a functional group with the structure . The linkage is also called an SS-bond or sometimes a disulfide bridge and is usually derived by the coupling of two thiol groups. In biology, disulfide bridges formed between thiol groups in two cysteine residues are an important component of the secondary and tertiary structure of proteins. ''Persulfide'' usually refers to compounds. In inorganic chemistry disulfide usually refers to the corresponding anion (−S−S−). Organic disulfides Symmetrical disulfides are compounds of the formula . Most disulfides encountered in organo sulfur chemistry are symmetrical disulfides. Unsymmetrical disulfides (also called heterodisulfides) are compounds of the formula . They are less common in organic chemistry, but most disulfides in nature are unsymmetrical. Properties The disulfide bonds are strong, with a typical bond dissociation energy of 60 kcal/mol (251&nbs ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Receptor Affinity
In chemistry, biochemistry, and pharmacology, a dissociation constant (K_D) is a specific type of equilibrium constant that measures the propensity of a larger object to separate (dissociate) reversibly into smaller components, as when a complex falls apart into its component molecules, or when a salt splits up into its component ions. The dissociation constant is the inverse of the association constant. In the special case of salts, the dissociation constant can also be called an ionization constant. For a general reaction: : A_\mathit B_\mathit \mathit A + \mathit B in which a complex \ce_x \ce_y breaks down into ''x'' A subunits and ''y'' B subunits, the dissociation constant is defined as : K_D = \frac where and ''x'' B''y''are the equilibrium concentrations of A, B, and the complex A''x'' B''y'', respectively. One reason for the popularity of the dissociation constant in biochemistry and pharmacology is that in the frequently encount ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Avidity
In biochemistry, avidity refers to the accumulated strength of ''multiple'' affinities of individual non-covalent binding interactions, such as between a protein receptor and its ligand, and is commonly referred to as functional affinity. Avidity differs from affinity, which describes the strength of a ''single'' interaction. However, because individual binding events increase the likelihood of occurrence of other interactions (i.e., increase the local concentration of each binding partner in proximity to the binding site), avidity should not be thought of as the mere sum of its constituent affinities but as the combined effect of all affinities participating in the biomolecular interaction. A particular important aspect relates to the phenomenon of 'avidity entropy'. Biomolecules often form heterogenous complexes or homogeneous oligomers and multimers or polymers. If clustered proteins form an organized matrix, such as the clathrin-coat, the interaction is described as a matricity. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
PI3K
Phosphoinositide 3-kinases (PI3Ks), also called phosphatidylinositol 3-kinases, are a family of enzymes involved in cellular functions such as cell growth, proliferation, differentiation, motility, survival and intracellular trafficking, which in turn are involved in cancer. PI3Ks are a family of related intracellular signal transducer enzymes capable of phosphorylating the 3 position hydroxyl group of the inositol ring of phosphatidylinositol (PtdIns). The pathway, with oncogene PIK3CA and tumor suppressor gene PTEN, is implicated in the sensitivity of cancer tumors to insulin and IGF1, and in calorie restriction. Discovery The discovery of PI3Ks by Lewis Cantley and colleagues began with their identification of a previously unknown phosphoinositide kinase associated with the polyoma middle T protein. They observed unique substrate specificity and chromatographic properties of the products of the lipid kinase, leading to the discovery that this phosphoinositide kinase had ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Src (gene)
Proto-oncogene tyrosine-protein kinase Src, also known as proto-oncogene c-Src, or simply c-Src (cellular Src; pronounced "sarc", as it is short for sarcoma), is a non-receptor tyrosine kinase protein that in humans is encoded by the ''SRC'' gene. It belongs to a family of Src family kinases and is similar to the v-Src (viral Src) gene of Rous sarcoma virus. It includes an SH2 domain, an SH3 domain and a tyrosine kinase domain. Two transcript variants encoding the same protein have been found for this gene. c-Src phosphorylates specific tyrosine residues in other tyrosine kinases. It plays a role in the regulation of embryonic development and cell growth. An elevated level of activity of c-Src is suggested to be linked to cancer progression by promoting other signals. Mutations in c-Src could be involved in the malignant progression of colon cancer. c-Src should not be confused with CSK (C-terminal Src kinase), an enzyme that phosphorylates c-Src at its C-terminus and provides ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Src Homology 2 Domain-containing
The SH2 (Src Homology 2) domain is a structurally conserved protein domain contained within the Src (gene), Src oncoprotein and in many other intracellular signal transduction, signal-transducing proteins. SH2 domains allow proteins containing those domains to dock to phosphorylated tyrosine residues on other proteins. SH2 domains are commonly found in Signal transducing adaptor protein, adaptor proteins that aid in the signal transduction of receptor tyrosine kinase pathways. Background SH2 is conserved by signalization of protein tyrosine kinase, which are binding on phosphotyrosine (pTyr). In the human proteome the class of pTyr-selective recognition domains is represented by SH2 domains. The N-terminal SH2 domains of cytoplasmic tyrosine kinase was at the beginning of evolution evolved with the occurrence of tyrosine phosphorylation. At the beginning it was supposed that, these domains serve as a substrate for their target kinase. Protein-protein interactions play a major role ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SH2 Domain
The SH2 (Src Homology 2) domain is a structurally conserved protein domain contained within the Src oncoprotein and in many other intracellular signal-transducing proteins. SH2 domains allow proteins containing those domains to dock to phosphorylated tyrosine residues on other proteins. SH2 domains are commonly found in adaptor proteins that aid in the signal transduction of receptor tyrosine kinase pathways. Background SH2 is conserved by signalization of protein tyrosine kinase, which are binding on phosphotyrosine (pTyr). In the human proteome the class of pTyr-selective recognition domains is represented by SH2 domains. The N-terminal SH2 domains of cytoplasmic tyrosine kinase was at the beginning of evolution evolved with the occurrence of tyrosine phosphorylation. At the beginning it was supposed that, these domains serve as a substrate for their target kinase. Protein-protein interactions play a major role in cellular growth and development. Modular domains, which are t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
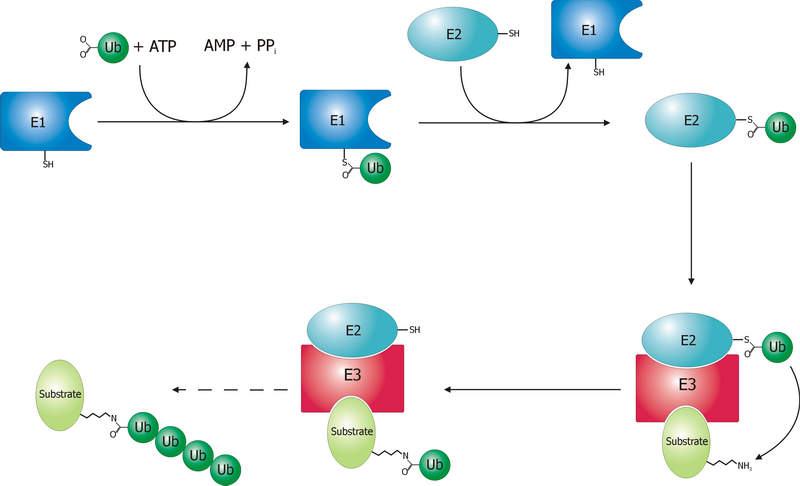
Ubiquitin Ligase
A ubiquitin ligase (also called an E3 ubiquitin ligase) is a protein that recruits an E2 ubiquitin-conjugating enzyme that has been loaded with ubiquitin, recognizes a protein substrate, and assists or directly catalyzes the transfer of ubiquitin from the E2 to the protein substrate. In simple and more general terms, the ligase enables movement of ubiquitin from a ubiquitin carrier to another thing (the substrate) by some mechanism. The ubiquitin, once it reaches its destination, ends up being attached by an isopeptide bond to a lysine residue, which is part of the target protein. E3 ligases interact with both the target protein and the E2 enzyme, and so impart substrate specificity to the E2. Commonly, E3s polyubiquitinate their substrate with Lys48-linked chains of ubiquitin, targeting the substrate for destruction by the proteasome. However, many other types of linkages are possible and alter a protein's activity, interactions, or localization. Ubiquitination by E3 ligases reg ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |