|
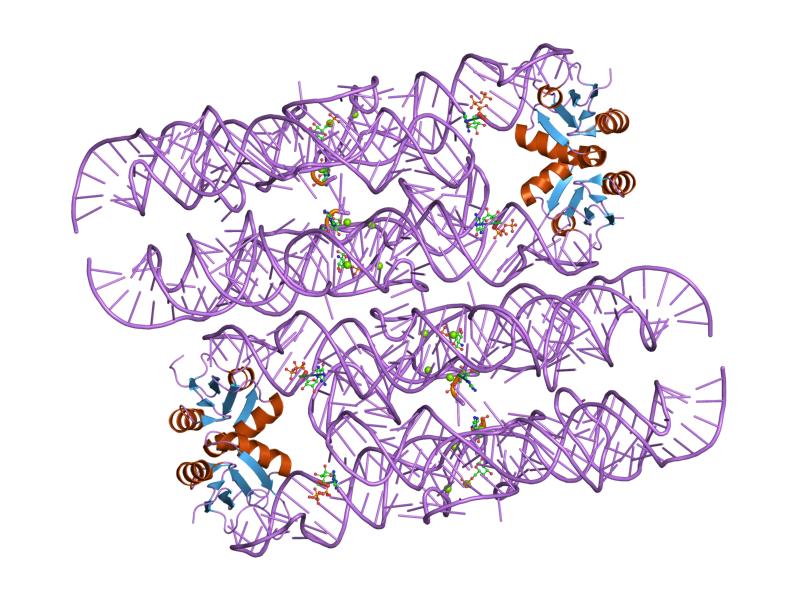
Hatchet Ribozyme
Background: The hatchet ribozyme is an RNA structure that catalyzes its own cleavage at a specific site. In other words, it is a self-cleaving ribozyme. Hatchet ribozymes were discovered by a bioinformatics strategy as RNAs Associated with Genes Associated with Twister and Hammerhead ribozymes, or RAGATH. Subsequent biochemical analysis supports the conclusion of a ribozyme function, and determined further characteristics of the chemical reaction catalyzed by the ribozyme. Nucleolytic ribozymes are small RNAs that adopt compact folds capable of site-specific cleavage/ligation reactions. 14 unique nucleolytic ribozymes have been identified to date, including recently discovered twister, pistol, twister-sister, and hatchet ribozymes that were identified based on application of comparative sequence and structural algorithms. The consensus sequence and secondary structure of this class includes 13 highly conserved and numerous other modestly conserved nucleotides inter-dispersed ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Secondary Structure
Protein secondary structure is the three dimensional conformational isomerism, form of ''local segments'' of proteins. The two most common Protein structure#Secondary structure, secondary structural elements are alpha helix, alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary structure elements typically spontaneously form as an intermediate before the protein protein folding, folds into its three dimensional protein tertiary structure, tertiary structure. Secondary structure is formally defined by the pattern of hydrogen bonds between the Amine, amino hydrogen and carboxyl oxygen atoms in the peptide backbone chain, backbone. Secondary structure may alternatively be defined based on the regular pattern of backbone Dihedral angle#Dihedral angles of proteins, dihedral angles in a particular region of the Ramachandran plot regardless of whether it has the correct hydrogen bonds. The concept of secondary structure was first introduced by Kaj Ulrik ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hairpin Ribozyme
The hairpin ribozyme is a small section of RNA that can act as a ribozyme. Like the hammerhead ribozyme it is found in RNA satellites of plant viruses. It was first identified in the minus strand of the tobacco ringspot virus (TRSV) satellite RNA where it catalyzes self-cleavage and joining (ligation) reactions to process the products of rolling circle virus replication into linear and circular satellite RNA molecules. The hairpin ribozyme is similar to the hammerhead ribozyme in that it does not require a metal ion for the reaction. Biological function The hairpin ribozyme is an RNA motif that catalyzes RNA processing reactions essential for replication of the satellite RNA molecules in which it is embedded. These reactions are self-processing, i.e. a molecule rearranging its own structure. Both cleavage and end joining reactions are mediated by the ribozyme motif, leading to a mixture of interconvertible linear and circular satellite RNA molecules. These reactions are imp ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Subgenomic MRNA
Subgenomic mRNAs are essentially smaller sections of the original transcribed template strand. 3' to 5' DNA or RNA During transcription, the original template strand is usually read from the 3' to the 5' end from beginning to end. Subgenomic mRNAs are created when transcription begins at the 3' end of the template strand (or 5' of the to-be-newly synthesized template) and begins to copy towards the 5' end of the template strand before "jumping" to the end of the template and copying the last nucleotides of the 5' end of the template, (finishing the 3' tail for the newly created strand). As a result, the translated strand will have a similar 5' end to varying degrees with the original template (depending on which part of the template the transcription jumped over) and a similar 3' end to the template. 5' to 3' (positive sense) viral RNA Positive-sense (5' to 3') viral RNA which may be directly translated into the desired viral proteins, undergoes a similar process as described ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Small Interfering RNA
Small interfering RNA (siRNA), sometimes known as short interfering RNA or silencing RNA, is a class of double-stranded RNA at first non-coding RNA molecules, typically 20-24 (normally 21) base pairs in length, similar to miRNA, and operating within the RNA interference (RNAi) pathway. It interferes with the expression of specific genes with complementary nucleotide sequences by degrading mRNA after transcription, preventing translation. Text was copied from this source, which is available under Creative Commons Attribution 4.0 International License Structure Naturally occurring siRNAs have a well-defined structure that is a short (usually 20 to 24- bp) double-stranded RNA (dsRNA) with phosphorylated 5' ends and hydroxylated 3' ends with two overhanging nucleotides. The Dicer enzyme catalyzes production of siRNAs from long dsRNAs and small hairpin RNAs. siRNAs can also be introduced into cells by transfection. Since in principle any gene can be knocked down by a syntheti ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Short Hairpin RNA
A short hairpin RNA or small hairpin RNA (shRNA/Hairpin Vector) is an artificial RNA molecule with a tight hairpin turn that can be used to silence target gene expression via RNA interference (RNAi). Expression of shRNA in cells is typically accomplished by delivery of plasmids or through viral or bacterial vectors. shRNA is an advantageous mediator of RNAi in that it has a relatively low rate of degradation and turnover. However, it requires use of an expression vector, which has the potential to cause side effects in medicinal applications. The promoter choice is essential to achieve robust shRNA expression. At first, polymerase III promoters such as U6 and H1 were used; however, these promoters lack spatial and temporal control. As such, there has been a shift to using polymerase II promoters, which are inducible, to regulate shRNA expression. Delivery Expression of shRNA in cells can be obtained by delivery of plasmids or through viral or bacterial vectors. Delivery of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Michaelis–Menten Kinetics
In biochemistry, Michaelis–Menten kinetics is one of the best-known models of enzyme kinetics. It is named after German biochemist Leonor Michaelis and Canadian physician Maud Menten. The model takes the form of an equation describing the rate of enzymatic reactions, by relating reaction rate v (rate of formation of product, ce P/math>) to ce S/math>, the concentration of a substrate ''S''. Its formula is given by : v = \frac = V_\max \frac This equation is called the Michaelis–Menten equation. Here, V_\max represents the maximum rate achieved by the system, happening at saturating substrate concentration for a given enzyme concentration. When the value of the Michaelis constant K_\mathrm is numerically equal to the substrate concentration, then the reaction rate is half of V_\max. Biochemical reactions involving a single substrate are often assumed to follow Michaelis–Menten kinetics, without regard to the model's underlying assumptions. Model In 1901, French ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Watson-Crick Base Pair
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA and RNA. Dictated by specific hydrogen bonding patterns, "Watson–Crick" (or "Watson–Crick–Franklin") base pairs (guanine–cytosine and adenine–thymine) allow the DNA helix to maintain a regular helical structure that is subtly dependent on its nucleotide sequence. The complementary nature of this based-paired structure provides a redundant copy of the genetic information encoded within each strand of DNA. The regular structure and data redundancy provided by the DNA double helix make DNA well suited to the storage of genetic information, while base-pairing between DNA and incoming nucleotides provides the mechanism through which DNA polymerase replicates DNA and RNA polymerase transcribes DNA into RNA. Many DNA-binding proteins ca ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hepatitis Delta Virus Ribozyme
The hepatitis delta virus (HDV) ribozyme is a non-coding RNA found in the hepatitis delta virus that is necessary for viral replication and is the only known human virus that utilizes ribozyme activity to infect its host. The ribozyme acts to process the RNA transcripts to unit lengths in a self-cleavage reaction during replication of the hepatitis delta virus, which is thought to propagate by a double rolling circle mechanism. The ribozyme is active ''in vivo'' in the absence of any protein factors and was the fastest known naturally occurring self-cleaving RNA at the time of its discovery. The crystal structure of this ribozyme has been solved using X-ray crystallography and shows five helical segments connected by a double pseudoknot. In addition to the sense (genomic version), all HDV viruses also have an antigenomic version of the HDV ribozyme. This version is not the exact complementary sequence but adopts the same structure as the sense (genomic) strand. The only "signi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
GlmS Glucosamine-6-phosphate Activated Ribozyme
The glucosamine-6-phosphate riboswitch ribozyme ( ''glmS'' ribozyme) is an RNA structure that resides in the 5' untranslated region (UTR) of the mRNA transcript of the ''glmS'' gene. This RNA regulates the ''glmS'' gene by responding to concentrations of a specific metabolite, glucosamine-6-phosphate (GlcN6P), in addition to catalyzing a self-cleaving chemical reaction upon activation. This cleavage leads to the degradation of the mRNA that contains the ribozyme, and lowers production of GlcN6P. The ''glmS'' gene encodes for an enzyme glutamine-fructose-6-phosphate amidotransferase, which catalyzes the formation of GlcN6P, a compound essential for cell wall biosynthesis, from fructose-6-phosphate and glutamine. Thus, when GlcN6P levels are high, the ''glmS'' ribozyme is activated and the mRNA transcript is degraded but in the absence of GlcN6P the gene continues to be translated into glutamine-fructose-6-phosphate amidotransferase and GlcN6P is produced. GlcN6P is a cofact ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hammerhead Ribozyme
The hammerhead ribozyme is an RNA motif that catalyzes reversible cleavage and ligation reactions at a specific site within an RNA molecule. It is one of several catalytic RNAs (ribozymes) known to occur in nature. It serves as a model system for research on the Nucleic acid structure, structure and properties of RNA, and is used for targeted RNA cleavage experiments, some with proposed therapeutic applications. Named for the resemblance of early secondary structure diagrams to a hammerhead shark, hammerhead ribozymes were originally discovered in two classes of plant virus-like RNAs: Satellite (biology), satellite RNAs and viroids. They are also known in some classes of Retrotransposon, retrotransposons, including the Retrozyme, retrozymes. The hammerhead ribozyme motif has been ubiquitously reported in lineages across the tree of life. The self-cleavage reactions, first reported in 1986, are part of a rolling circle replication mechanism. The hammerhead sequence is sufficient fo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequence Conservation
In evolutionary biology, conserved sequences are identical or similar sequences in nucleic acids ( DNA and RNA) or proteins across species ( orthologous sequences), or within a genome ( paralogous sequences), or between donor and receptor taxa ( xenologous sequences). Conservation indicates that a sequence has been maintained by natural selection. A highly conserved sequence is one that has remained relatively unchanged far back up the phylogenetic tree, and hence far back in geological time. Examples of highly conserved sequences include the RNA components of ribosomes present in all domains of life, the homeobox sequences widespread amongst Eukaryotes, and the tmRNA in Bacteria. The study of sequence conservation overlaps with the fields of genomics, proteomics, evolutionary biology, phylogenetics, bioinformatics and mathematics. History The discovery of the role of DNA in heredity, and observations by Frederick Sanger of variation between animal insulins in 1949, promp ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |