|
FASTQ Format
FASTQ format is a text-based format for storing both a biological sequence (usually nucleotide sequence) and its corresponding quality scores. Both the sequence letter and quality score are each encoded with a single ASCII character for brevity. It was originally developed at the Wellcome Trust Sanger Institute to bundle a FASTA formatted sequence and its quality data, but has recently become the ''de facto'' standard for storing the output of high-throughput sequencing instruments such as the Illumina Genome Analyzer. Format A FASTQ file has four line-separated fields per sequence: * Field 1 begins with a '@' character and is followed by a sequence identifier and an ''optional'' description (like a FASTA title line). * Field 2 is the raw sequence letters. * Field 3 begins with a '+' character and is ''optionally'' followed by the same sequence identifier (and any description) again. * Field 4 encodes the quality values for the sequence in Field 2, and must contain the same num ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Wellcome Trust Sanger Institute
The Wellcome Sanger Institute, previously known as The Sanger Centre and Wellcome Trust Sanger Institute, is a non-profit British genomics and genetics research institute, primarily funded by the Wellcome Trust. It is located on the Wellcome Genome Campus by the village of Hinxton, outside Cambridge. It shares this location with the European Bioinformatics Institute. It was established in 1992 and named after double Nobel Laureate Frederick Sanger. It was conceived as a large scale DNA sequencing centre to participate in the Human Genome Project, and went on to make the largest single contribution to the gold standard sequence of the human genome. From its inception the institute established and has maintained a policy of data sharing, and does much of its research in collaboration. Since 2000, the institute expanded its mission to understand "the role of genetics in health and disease". The institute now employs around 900 people and engages in five main areas of research ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
BioJava
BioJava is an open-source software project dedicated to provide Java tools to process biological data.VS Matha and P Kangueane, 2009, ''Bioinformatics: a concept-based introduction'', 2009. p26 BioJava is a set of library functions written in the programming language Java for manipulating sequences, protein structures, file parsers, Common Object Request Broker Architecture (CORBA) interoperability, Distributed Annotation System (DAS), access to AceDB, dynamic programming, and simple statistical routines. BioJava supports a huge range of data, starting from DNA and protein sequences to the level of 3D protein structures. The BioJava libraries are useful for automating many daily and mundane bioinformatics tasks such as to parsing a Protein Data Bank (PDB) file, interacting with Jmol and many more. This application programming interface (API) provides various file parsers, data models and algorithms to facilitate working with the standard data formats and enables rapid application d ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
BioRuby
BioRuby is a collection of open-source Ruby code, comprising classes for computational molecular biology and bioinformatics. It contains classes for DNA and protein sequence analysis, sequence alignment, biological database parsing, structural biology and other bioinformatics tasks. BioRuby is released under the GNU GPL version 2 or Ruby licence and is one of a number of Bio* projects, designed to reduce code duplication. In 2011, the BioRuby project introduced the Biogem software plugin system, with two or three new plugins added every month. BioRuby is managed via the BioRuby website and GitHub repository. History BioRuby The BioRuby project was first started in 2000 by Toshiaki Katayama as a Ruby implementation of similar bioinformatics packages such as BioPerl and BioPython. The initial release of version 0.1 was frequently updated by contributors both informally and at organised “hackathon” events; in June 2005, BioRuby was funded by IPA as an Exploratory Software Pro ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
BioPerl
BioPerl is a collection of Perl modules that facilitate the development of Perl scripts for bioinformatics applications. It has played an integral role in the Human Genome Project. Background BioPerl is an active open source software project supported by the Open Bioinformatics Foundation. The first set of Perl codes of BioPerl was created by Tim Hubbard and Jong Bhak at MRC Centre Cambridge, where the first genome sequencing was carried out by Fred Sanger. MRC Centre was one of the hubs and birth places of modern bioinformatics as it had a large quantity of DNA sequences and 3D protein structures. Hubbard was using the th_lib.pl Perl library, which contained many useful Perl subroutines for bioinformatics. Bhak, Hubbard's first PhD student, created jong_lib.pl. Bhak merged the two Perl subroutine libraries into Bio.pl. The name BioPerl was coined jointly by Bhak and Steven Brenner at the Centre for Protein Engineering (CPE). In 1995, Brenner organized a BioPerl session at the In ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
EMBOSS
EMBOSS is a free open source software analysis package developed for the needs of the molecular biology and bioinformatics user community. The software automatically copes with data in a variety of formats and even allows transparent retrieval of sequence data from the web. Also, as extensive libraries are provided with the package, it is a platform to allow other scientists to develop and release software in true open source spirit. EMBOSS also integrates a range of currently available packages and tools for sequence analysis into a seamless whole. ''EMBOSS'' is an acronym for European Molecular Biology Open Software Suite. The ''European'' part of the name hints at the wider scope. The core EMBOSS groups are collaborating with many other groups to develop the new applications that the users need. This was done from the beginning with EMBnet, the European Molecular Biology Network. EMBnet has many nodes worldwide most of which are national bioinformatics services. EMBnet has the p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
File Extension
A filename extension, file name extension or file extension is a suffix to the name of a computer file (e.g., .txt, .docx, .md). The extension indicates a characteristic of the file contents or its intended use. A filename extension is typically delimited from the rest of the filename with a full stop (period), but in some systems it is separated with spaces. Other extension formats include dashes and/or underscores on early versions of Linux and some versions of IBM AIX. Some file systems implement filename extensions as a feature of the file system itself and may limit the length and format of the extension, while others treat filename extensions as part of the filename without special distinction. Usage Filename extensions may be considered a type of metadata. They are commonly used to imply information about the way data might be stored in the file. The exact definition, giving the criteria for deciding what part of the file name is its extension, belongs to the rules of the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Burrows–Wheeler Transform
The Burrows–Wheeler transform (BWT, also called block-sorting compression) rearranges a character string into runs of similar characters. This is useful for compression, since it tends to be easy to compress a string that has runs of repeated characters by techniques such as move-to-front transform and run-length encoding. More importantly, the transformation is ''reversible'', without needing to store any additional data except the position of the first original character. The BWT is thus a "free" method of improving the efficiency of text compression algorithms, costing only some extra computation. The Burrows–Wheeler transform is an algorithm used to prepare data for use with data compression techniques such as bzip2. It was invented by Michael Burrows and David Wheeler in 1994 while Burrows was working at DEC Systems Research Center in Palo Alto, California. It is based on a previously unpublished transformation discovered by Wheeler in 1983. The algorithm can be imple ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
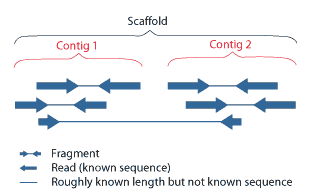
Contig
A contig (from ''contiguous'') is a set of overlapping DNA segments that together represent a consensus region of DNA.Gregory, S. ''Contig Assembly''. Encyclopedia of Life Sciences, 2005. In bottom-up sequencing projects, a contig refers to overlapping sequence data ( reads); in top-down sequencing projects, contig refers to the overlapping clones that form a physical map of the genome that is used to guide sequencing and assembly.Dear, P. H. ''Genome Mapping''. Encyclopedia of Life Sciences, 2005. . Contigs can thus refer both to overlapping DNA sequences and to overlapping physical segments (fragments) contained in clones depending on the context. Original definition of contig In 1980, Staden wrote: ''In order to make it easier to talk about our data gained by the shotgun method of sequencing we have invented the word "contig". A contig is a set of gel readings that are related to one another by overlap of their sequences. All gel readings belong to one and only one con ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
De Bruijn Graph
In graph theory, an -dimensional De Bruijn graph of symbols is a directed graph representing overlaps between sequences of symbols. It has vertices, consisting of all possible sequences of the given symbols; the same symbol may appear multiple times in a sequence. For a set of symbols the set of vertices is: :V=S^n=\. If one of the vertices can be expressed as another vertex by shifting all its symbols by one place to the left and adding a new symbol at the end of this vertex, then the latter has a directed edge to the former vertex. Thus the set of arcs (that is, directed edges) is :E=\. Although De Bruijn graphs are named after Nicolaas Govert de Bruijn, they were discovered independently by both De Bruijn and I. J. Good. Much earlier, Camille Flye Sainte-Marie implicitly used their properties. Properties * If , then the condition for any two vertices forming an edge holds vacuously, and hence all the vertices are connected, forming a total of edges. * Each vertex has ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SAM (file Format)
Sequence Alignment Map (SAM) is a text-based format originally for storing biological sequences aligned to a reference sequence developed by Heng Li and Bob Handsaker ''et al''. It was developed when the 1000 Genomes Project wanted to move away from the MAQ mapper format and decided to design a new format. The overall TAB-delimited flavour of the format came from an earlier format inspired by BLAT’s PSL. The name of SAM came from Gabor Marth from University of Utah, who originally had a format under the same name but with a different syntax more similar to a BLAST output. It is widely used for storing data, such as nucleotide sequences, generated by next generation sequencing technologies, and the standard has been broadened to include unmapped sequences. The format supports short and long reads (up to 128 Mbp) produced by different sequencing platforms and is used to hold mapped data within the Genome Analysis Toolkit (GATK) and across the Broad Institute, the Wellcome S ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |