|
Epistasis And Functional Genomics
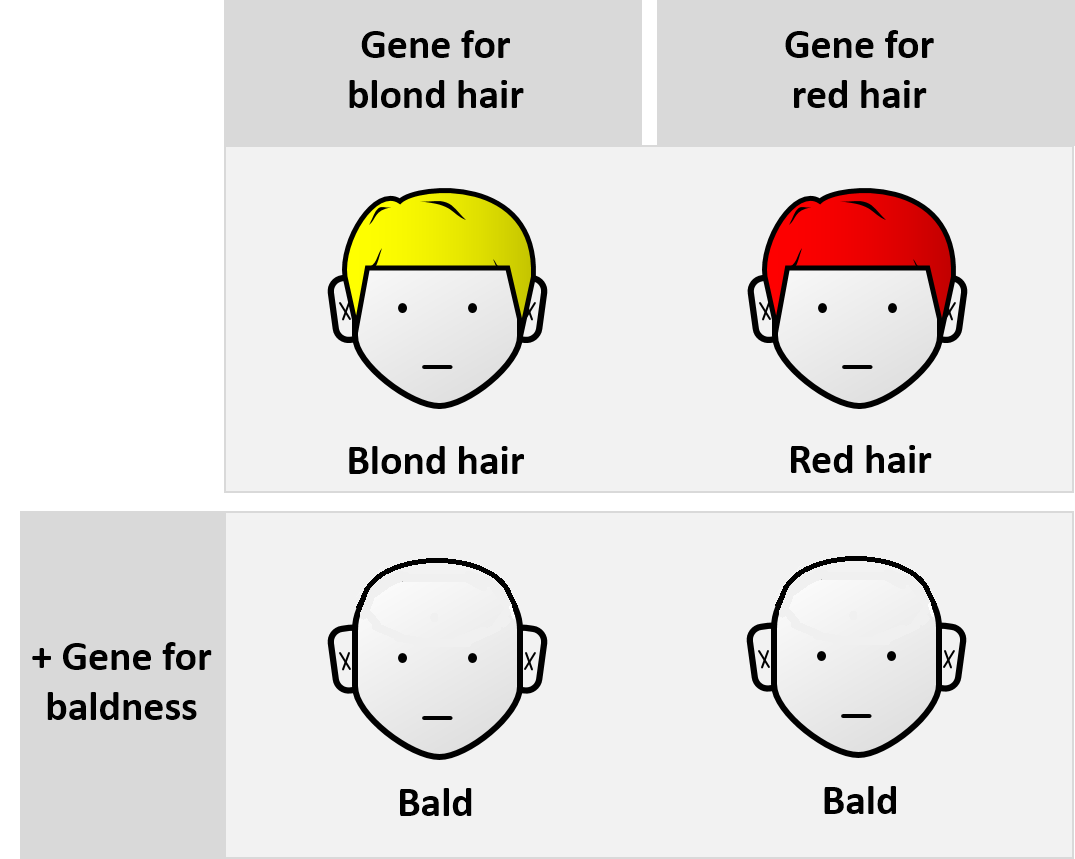
Epistasis refers to genetic interactions in which the mutation of one gene masks the phenotypic effects of a mutation at another genetic locus, locus. Systematic analysis of these epistatic interactions can provide insight into the structure and function of genetic pathways. Examining the phenotypes resulting from pairs of mutations helps in understanding how the function of these genes intersects. Genetic interactions are generally classified as either Positive/Alleviating or Negative/Aggravating. Fitness epistasis (an interaction between non-allelic genes) is positive (in other words, diminishing, antagonistic or buffering) when a loss of function mutation of two given genes results in exceeding the fitness (biology), fitness predicted from individual effects of deleterious mutations, and it is negative (that is, reinforcing, synergistic or aggravating) when it decreases fitness. Ryszard Korona and Lukas Jasnos showed that the epistatic effect is usually positive in Saccharomyces ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Epistasis
Epistasis is a phenomenon in genetics in which the effect of a gene mutation is dependent on the presence or absence of mutations in one or more other genes, respectively termed modifier genes. In other words, the effect of the mutation is dependent on the genetic background in which it appears. Epistatic mutations therefore have different effects on their own than when they occur together. Originally, the term ''epistasis'' specifically meant that the effect of a gene variant is masked by that of a different gene. The concept of ''epistasis'' originated in genetics in 1907 but is now used in biochemistry, computational biology and evolutionary biology. The phenomenon arises due to interactions, either between genes (such as mutations also being needed in regulators of gene expression) or within them (multiple mutations being needed before the gene loses function), leading to non-linear effects. Epistasis has a great influence on the shape of evolutionary landscapes, which l ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecularly Bar-coded
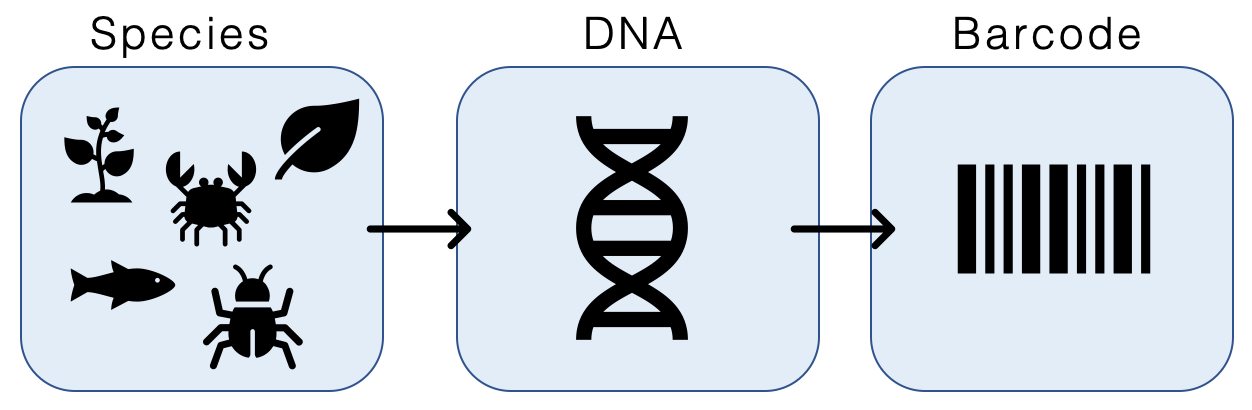
DNA barcoding is a method of species identification using a short section of DNA from a specific gene or genes. The premise of DNA barcoding is that by comparison with a reference library of such DNA sections (also called " sequences"), an individual sequence can be used to uniquely identify an organism to species, just as a supermarket scanner uses the familiar black stripes of the UPC barcode to identify an item in its stock against its reference database. These "barcodes" are sometimes used in an effort to identify unknown species or parts of an organism, simply to catalog as many taxa as possible, or to compare with traditional taxonomy in an effort to determine species boundaries. Different gene regions are used to identify the different organismal groups using barcoding. The most commonly used barcode region for animals and some protists is a portion of the cytochrome ''c'' oxidase I (COI or COX1) gene, found in mitochondrial DNA. Other genes suitable for DNA barcoding ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genetic Marker
A genetic marker is a gene or DNA sequence with a known location on a chromosome that can be used to identify individuals or species. It can be described as a variation (which may arise due to mutation or alteration in the genomic loci) that can be observed. A genetic marker may be a short DNA sequence, such as a sequence surrounding a single base-pair change ( single nucleotide polymorphism, SNP), or a long one, like minisatellites. Background For many years, gene mapping was limited to identifying organisms by traditional phenotypes markers. This included genes that encoded easily observable characteristics such as blood types or seed shapes. The insufficient number of these types of characteristics in several organisms limited the mapping efforts that could be done. This prompted the development of gene markers which could identify genetic characteristics that are not readily observable in organisms (such as protein variation). Types Some commonly used types of genetic markers ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Antibiotic
An antibiotic is a type of antimicrobial substance active against bacteria. It is the most important type of antibacterial agent for fighting bacterial infections, and antibiotic medications are widely used in the treatment and prevention of such infections. They may either kill or inhibit the growth of bacteria. A limited number of antibiotics also possess antiprotozoal activity. Antibiotics are not effective against viruses such as the common cold or influenza; drugs which inhibit viruses are termed antiviral drugs or antivirals rather than antibiotics. Sometimes, the term ''antibiotic''—literally "opposing life", from the Greek roots ἀντι ''anti'', "against" and βίος ''bios'', "life"—is broadly used to refer to any substance used against microbes, but in the usual medical usage, antibiotics (such as penicillin) are those produced naturally (by one microorganism fighting another), whereas non-antibiotic antibacterials (such as sulfonamides and antiseptics) ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein. mRNA is created during the process of Transcription (biology), transcription, where an enzyme (RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and, utilising amino acids carried by transfer RNA (tRNA), the ribosome creates the protein. This process is known as Translation (biology), translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genet ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Affinity Purification
Affinity chromatography is a method of separating a biomolecule from a mixture, based on a highly specific molecular binding, macromolecular binding interaction between the biomolecule and another substance. The specific type of binding interaction depends on the biomolecule of interest; antigen and antibody, enzyme and substrate (biochemistry), substrate, biochemistry receptor, receptor and ligand (biochemistry), ligand, or protein and nucleic acid binding interactions are frequently exploited for isolation of various biomolecules. Affinity chromatography is useful for its high selectivity (chromatography), selectivity and resolution (chromatography), resolution of separation, compared to other chromatographic methods. Principle Affinity chromatography has the advantage of specific binding interactions between the analyte of interest (normally dissolved in the mobile phase), and a binding partner or ligand (immobilized on the Stationary phase (chemistry), stationary phase). In ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cell Cycle
The cell cycle, or cell-division cycle, is the series of events that take place in a cell that cause it to divide into two daughter cells. These events include the duplication of its DNA (DNA replication) and some of its organelles, and subsequently the partitioning of its cytoplasm, chromosomes and other components into two daughter cells in a process called cell division. In cells with nuclei ( eukaryotes, i.e., animal, plant, fungal, and protist cells), the cell cycle is divided into two main stages: interphase and the mitotic (M) phase (including mitosis and cytokinesis). During interphase, the cell grows, accumulating nutrients needed for mitosis, and replicates its DNA and some of its organelles. During the mitotic phase, the replicated chromosomes, organelles, and cytoplasm separate into two new daughter cells. To ensure the proper replication of cellular components and division, there are control mechanisms known as cell cycle checkpoints after each of the key steps ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transcriptional Regulation
In molecular biology and genetics, transcriptional regulation is the means by which a cell regulates the conversion of DNA to RNA (transcription), thereby orchestrating gene activity. A single gene can be regulated in a range of ways, from altering the number of copies of RNA that are transcribed, to the temporal control of when the gene is transcribed. This control allows the cell or organism to respond to a variety of intra- and extracellular signals and thus mount a response. Some examples of this include producing the mRNA that encode enzymes to adapt to a change in a food source, producing the gene products involved in cell cycle specific activities, and producing the gene products responsible for cellular differentiation in multicellular eukaryotes, as studied in evolutionary developmental biology. The regulation of transcription is a vital process in all living organisms. It is orchestrated by transcription factors and other proteins working in concert to finely tune t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Elongation Factor
Elongation factors are a set of proteins that function at the ribosome, during protein synthesis, to facilitate translational elongation from the formation of the first to the last peptide bond of a growing polypeptide. Most common elongation factors in prokaryotes are EF-Tu, EF-Ts, EF-G. Bacteria and eukaryotes use elongation factors that are largely homologous to each other, but with distinct structures and different research nomenclatures. Elongation is the most rapid step in translation. In bacteria, it proceeds at a rate of 15 to 20 amino acids added per second (about 45-60 nucleotides per second). In eukaryotes the rate is about two amino acids per second (about 6 nucleotides read per second). Elongation factors play a role in orchestrating the events of this process, and in ensuring the high accuracy translation at these speeds. Nomenclature of homologous EFs In addition to their cytoplasmic machinery, eukaryotic mitochondria and plastids have their own translation ma ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hierarchical Clustering
In data mining and statistics, hierarchical clustering (also called hierarchical cluster analysis or HCA) is a method of cluster analysis that seeks to build a hierarchy of clusters. Strategies for hierarchical clustering generally fall into two categories: * Agglomerative: This is a " bottom-up" approach: Each observation starts in its own cluster, and pairs of clusters are merged as one moves up the hierarchy. * Divisive: This is a "top-down" approach: All observations start in one cluster, and splits are performed recursively as one moves down the hierarchy. In general, the merges and splits are determined in a greedy manner. The results of hierarchical clustering are usually presented in a dendrogram. The standard algorithm for hierarchical agglomerative clustering (HAC) has a time complexity of \mathcal(n^3) and requires \Omega(n^2) memory, which makes it too slow for even medium data sets. However, for some special cases, optimal efficient agglomerative methods (of c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Model Organism
A model organism (often shortened to model) is a non-human species that is extensively studied to understand particular biological phenomena, with the expectation that discoveries made in the model organism will provide insight into the workings of other organisms. Model organisms are widely used to research human disease when human experimentation would be unfeasible or unethical. This strategy is made possible by the common descent of all living organisms, and the conservation of metabolic and developmental pathways and genetic material over the course of evolution. Studying model organisms can be informative, but care must be taken when generalizing from one organism to another. In researching human disease, model organisms allow for better understanding the disease process without the added risk of harming an actual human. The species chosen will usually meet a determined taxonomic equivalency to humans, so as to react to disease or its treatment in a way that resembles ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |