|
De Novo Transcriptome Assembly
''De novo'' transcriptome assembly is the de novo sequence assembly method of creating a transcriptome without the aid of a reference genome. Introduction As a result of the development of novel sequencing technologies, the years between 2008 and 2012 saw a large drop in the cost of sequencing. Per megabase and genome, the cost dropped to 1/100,000th and 1/10,000th of the price, respectively. Prior to this, only transcriptomes of organisms that were of broad interest and utility to scientific research were sequenced; however, these developed in 2010s high-throughput sequencing (also called next-generation sequencing) technologies are both cost- and labor- effective, and the range of organisms studied via these methods is expanding. Transcriptomes have subsequently been created for chickpea, planarians, '' Parhyale hawaiensis'', as well as the brains of the Nile crocodile, the corn snake, the bearded dragon, and the red-eared slider, to name just a few. Examining non-model organ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
De Novo Sequence Assemblers
De novo sequence assemblers are a type of program that assembles short nucleotide sequences into longer ones without the use of a reference genome. These are most commonly used in bioinformatic studies to assemble genomes or transcriptomes. Two common types of de novo assemblers are greedy algorithm assemblers and De Bruijn graph assemblers. Types of de novo assemblers There are two types of algorithms that are commonly utilized by these assemblers: greedy, which aim for local optima, and graph method algorithms, which aim for global optima. Different assemblers are tailored for particular needs, such as the assembly of (small) bacterial genomes, (large) eukaryotic genomes, or transcriptomes. Greedy algorithm assemblers are assemblers that find local optima in alignments of smaller reads. Greedy algorithm assemblers typically feature several steps: 1) pairwise distance calculation of reads, 2) clustering of reads with greatest overlap, 3) assembly of overlapping reads into larg ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Contig
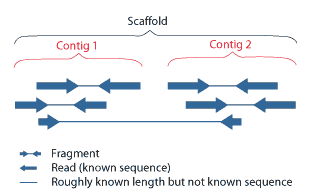
A contig (from ''contiguous'') is a set of overlapping DNA segments that together represent a consensus region of DNA.Gregory, S. ''Contig Assembly''. Encyclopedia of Life Sciences, 2005. In bottom-up sequencing projects, a contig refers to overlapping sequence data ( reads); in top-down sequencing projects, contig refers to the overlapping clones that form a physical map of the genome that is used to guide sequencing and assembly.Dear, P. H. ''Genome Mapping''. Encyclopedia of Life Sciences, 2005. . Contigs can thus refer both to overlapping DNA sequences and to overlapping physical segments (fragments) contained in clones depending on the context. Original definition of contig In 1980, Staden wrote: ''In order to make it easier to talk about our data gained by the shotgun method of sequencing we have invented the word "contig". A contig is a set of gel readings that are related to one another by overlap of their sequences. All gel readings belong to one and only one con ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Ontology
The Gene Ontology (GO) is a major bioinformatics initiative to unify the representation of gene and gene product attributes across all species. More specifically, the project aims to: 1) maintain and develop its controlled vocabulary of gene and gene product attributes; 2) annotate genes and gene products, and assimilate and disseminate annotation data; and 3) provide tools for easy access to all aspects of the data provided by the project, and to enable functional interpretation of experimental data using the GO, for example via enrichment analysis. GO is part of a larger classification effort, the Open Biomedical Ontologies, being one of the Initial Candidate Members of the OBO Foundry. Whereas gene nomenclature focuses on gene and gene products, the Gene Ontology focuses on the function of the genes and gene products. The GO also extends the effort by using markup language to make the data (not only of the genes and their products but also of curated attributes) machine read ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Blast2GO
Blast2GO, first published in 2005, is a bioinformatics software tool for the automatic, high-throughput functional annotation of novel sequence data (genes proteins). It makes use of the BLAST algorithm to identify similar sequences to then transfers existing functional annotation from yet characterised sequences to the novel one. The functional information is represented via the Gene Ontology (GO), a controlled vocabulary of functional attributes. The Gene Ontology, or GO, is a major bioinformatics initiative to unify the representation of gene and gene product attributes across all species. See also * Protein function prediction * Functional genomics * Bioinformatics Bioinformatics () is an interdisciplinary field that develops methods and software tools for understanding biological data, in particular when the data sets are large and complex. As an interdisciplinary field of science, bioinformatics combi ... References External links * Blast2GO - Tool for f ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
K-mer
In bioinformatics, ''k''-mers are substrings of length k contained within a biological sequence. Primarily used within the context of computational genomics and sequence analysis, in which ''k''-mers are composed of nucleotides (''i.e''. A, T, G, and C), ''k''-mers are capitalized upon to assemble DNA sequences, improve heterologous gene expression, identify species in metagenomic samples, and create attenuated vaccines. Usually, the term ''k''-mer refers to all of a sequence's subsequences of length k, such that the sequence AGAT would have four monomers (A, G, A, and T), three 2-mers (AG, GA, AT), two 3-mers (AGA and GAT) and one 4-mer (AGAT). More generally, a sequence of length L will have L - k + 1 ''k''-mers and n^ total possible ''k''-mers, where n is number of possible monomers (e.g. four in the case of DNA). Introduction ''k''-mers are simply length k subsequences. For example, all the possible ''k''-mers of a DNA sequence are shown below: A method of visualizi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
De Bruijn Graph
In graph theory, an -dimensional De Bruijn graph of symbols is a directed graph representing overlaps between sequences of symbols. It has vertices, consisting of all possible sequences of the given symbols; the same symbol may appear multiple times in a sequence. For a set of symbols the set of vertices is: :V=S^n=\. If one of the vertices can be expressed as another vertex by shifting all its symbols by one place to the left and adding a new symbol at the end of this vertex, then the latter has a directed edge to the former vertex. Thus the set of arcs (that is, directed edges) is :E=\. Although De Bruijn graphs are named after Nicolaas Govert de Bruijn, they were discovered independently by both De Bruijn and I. J. Good. Much earlier, Camille Flye Sainte-Marie implicitly used their properties. Properties * If , then the condition for any two vertices forming an edge holds vacuously, and hence all the vertices are connected, forming a total of edges. * Each vertex has ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequencing
In genetics and biochemistry, sequencing means to determine the primary structure (sometimes incorrectly called the primary sequence) of an unbranched biopolymer. Sequencing results in a symbolic linear depiction known as a sequence which succinctly summarizes much of the atomic-level structure of the sequenced molecule. DNA sequencing DNA sequencing is the process of determining the nucleotide order of a given DNA fragment. So far, most DNA sequencing has been performed using the chain termination method developed by Frederick Sanger. This technique uses sequence-specific termination of a DNA synthesis reaction using modified nucleotide substrates. However, new sequencing technologies such as pyrosequencing are gaining an increasing share of the sequencing market. More genome data are now being produced by pyrosequencing than Sanger DNA sequencing. Pyrosequencing has enabled rapid genome sequencing. Bacterial genomes can be sequenced in a single run with several times cover ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Overlap Graphs
Overlap may refer to: * In set theory, an overlap of elements shared between sets is called an intersection, as in a Venn diagram. * In music theory, overlap is a synonym for reinterpretation of a chord at the boundary of two musical phrases * Overlap (railway signalling), the length of track beyond a stop signal that is proved to be clear of obstructions as a safety margin * Overlap (road), a place where multiple road numbers overlap * Overlap (term rewriting), in mathematics, computer science, and logic, a property of the reduction rules in term rewriting systems * Overlap add, an efficient convolution method using FFT * Overlap coefficient, a similarity measure between sets * Orbital overlap, important concept in quantum mechanics describing a type of orbital interaction that affects bond strength Overlapping can refer to: * "Reaching over", term in Schenkerian theory, see Schenkerian analysis#Lines between voices, reaching over See also * Overlay (other) Overlay may ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sense (molecular Biology)
In molecular biology and genetics, the sense of a nucleic acid molecule, particularly of a strand of DNA or RNA, refers to the nature of the roles of the strand and its complement in specifying a sequence of amino acids. Depending on the context, sense may have slightly different meanings. For example, negative-sense strand of DNA is equivalent to the template strand, whereas the positive-sense strand is the non-template strand whose nucleotide sequence is equivalent to the sequence of the mRNA transcript. DNA sense Because of the complementary nature of base-pairing between nucleic acid polymers, a double-stranded DNA molecule will be composed of two strands with sequences that are reverse complements of each other. To help molecular biologists specifically identify each strand individually, the two strands are usually differentiated as the "sense" strand and the "antisense" strand. An individual strand of DNA is referred to as positive-sense (also positive (+) or simply sense) ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Assemblers
Assembler may refer to: Arts and media * Nobukazu Takemura, avant-garde electronic musician, stage name Assembler * Assemblers, a fictional race in the ''Star Wars'' universe * Assemblers, an alternative name of the superhero group Champions of Angor Biology *Assembler (bioinformatics), a program to perform genome assembly *Assembler (nanotechnology), a conjectured construction machine that would manipulate and build with individual atoms or molecules Computing *Assembler (computing), a computer program which translates assembly language to machine language **Assembly language, a more readable interpretation of a processor's machine code, allowing easier understanding and programming by humans, sometimes erroneously referenced as 'assembler' after the program which translates it Other uses * a worker on an assembly line See also * Assemble (other) * Assembly (other) * Constructor (other) Constructor may refer to: Science and technology * Constru ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
ABI Solid Sequencing
SOLiD (Sequencing by Oligonucleotide Ligation and Detection) is a next-generation DNA sequencing technology developed by Life Technologies and has been commercially available since 2006. This next generation technology generates 108 - 109 small sequence reads at one time. It uses 2 base encoding to decode the raw data generated by the sequencing platform into sequence data. This method should not be confused with "sequencing by synthesis," a principle used by Roche-454 pyrosequencing (introduced in 2005, generating millions of 200-400bp reads in 2009), and the Solexa system (now owned by Illumina) (introduced in 2006, generating hundreds of millions of 50-100bp reads in 2009) These methods have reduced the cost from $0.01/base in 2004 to nearly $0.0001/base in 2006 and increased the sequencing capacity from 1,000,000 bases/machine/day in 2004 to more than 5,000,000,000 bases/machine/day in 2009. Over 30 publications exist describing its use first for nucleosome positioning ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Illumina (company)
Illumina, Inc. is an American biotechnology company, headquartered in San Diego, California. Incorporated on April 1, 1998, Illumina develops, manufactures, and markets integrated systems for the analysis of genetic variation and biological function. The company provides a line of products and services that serves the sequencing, genotyping and gene expression, and proteomics markets. Illumina's technology had purportedly reduced the cost of sequencing a human genome to by 2014. Its customers include genomic research centers, pharmaceutical companies, academic institutions, clinical research organizations, and biotechnology companies. History Illumina was founded in April 1998 by David Walt, Larry Bock, John Stuelpnagel, Anthony Czarnik, and Mark Chee. While working with CW Group, a venture-capital firm, Bock and Stuelpnagel uncovered what would become Illumina's BeadArray technology at Tufts University and negotiated an exclusive license to that technology. In 1999, Illumin ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |

.jpg)