|
DXZ4
DXZ4 is a variable number tandemly repeated DNA sequence. In humans it is composed of 3kb monomers containing a highly conserved CTCF binding site. CTCF is a transcription factor protein and the main insulator responsible for partitioning of chromatin domains in the vertebrate genome. In addition to being enriched in CpG-islands, ''DXZ4'' transcribes long non-coding RNAs ( lncRNAs) and small RNAs of unknown function. Repeat copy number of ''DXZ4'' is highly polymorphic in human populations (varying between 50 and 100 copies). ''DXZ4'' is one of many large tandem repeat loci defined as macrosatellites. Several macrosatellites have been described in humans and share similar features, such as high GC content, large repeat monomers, and high variability for repeat copy number within populations. ''DXZ4'' plays an important role in the unique structural conformation of the inactive X chromosome (Xi) in female somatic cell A somatic cell (from Ancient Greek σῶμα ''sôma'', meani ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Macrosatellite
Macrosatellites are the largest of the tandem DNA repeats. Each macrosatellite repeat typically is several kilobases in length, and the entire repeat array often spans hundreds of kilobases. Reduced number of repeats on chromosome 4 (D4Z4 repeats) causes euchromatization of local DNA and is the predominant cause of facioscapulohumeral muscular dystrophy (FSHD). Other macrosatellites are RS447, NBL2 and DXZ4, although RS447 is also commonly referred to as a "megasatellite." See also * Microsatellite * Minisatellite * Satellite DNA Satellite DNA consists of very large arrays of tandemly repeating, non-coding DNA. Satellite DNA is the main component of functional centromeres, and form the main structural constituent of heterochromatin. The name "satellite DNA" refers to the ... References Genetics Repetitive DNA sequences {{Genetics-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
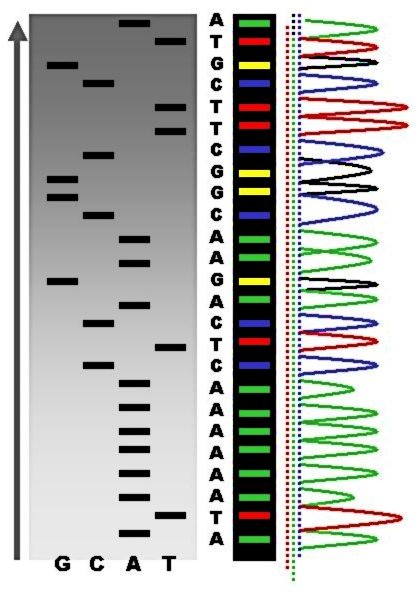
DNA Sequence
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery. Knowledge of DNA sequences has become indispensable for basic biological research, DNA Genographic Projects and in numerous applied fields such as medical diagnosis, biotechnology, forensic biology, virology and biological systematics. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers, characterize antibody repertoire, and can be used to guide patient treatment. Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged. The rapid speed of sequencing attained with modern D ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Monomers
In chemistry, a monomer ( ; ''mono-'', "one" + '' -mer'', "part") is a molecule that can react together with other monomer molecules to form a larger polymer chain or three-dimensional network in a process called polymerization. Classification Monomers can be classified in many ways. They can be subdivided into two broad classes, depending on the kind of the polymer that they form. Monomers that participate in condensation polymerization have a different stoichiometry than monomers that participate in addition polymerization: : Other classifications include: *natural vs synthetic monomers, e.g. glycine vs caprolactam, respectively *polar vs nonpolar monomers, e.g. vinyl acetate vs ethylene, respectively *cyclic vs linear, e.g. ethylene oxide vs ethylene glycol, respectively The polymerization of one kind of monomer gives a homopolymer. Many polymers are copolymers, meaning that they are derived from two different monomers. In the case of condensation polymerizations, the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
CTCF
Transcriptional repressor CTCF also known as 11-zinc finger protein or CCCTC-binding factor is a transcription factor that in humans is encoded by the ''CTCF'' gene. CTCF is involved in many cellular processes, including transcriptional regulation, insulator activity, V(D)J recombination and regulation of chromatin architecture. Discovery CCCTC-Binding factor or CTCF was initially discovered as a negative regulator of the chicken c-myc gene. This protein was found to be binding to three regularly spaced repeats of the core sequence CCCTC and thus was named CCCTC binding factor. Function The primary role of CTCF is thought to be in regulating the 3D structure of chromatin. CTCF binds together strands of DNA, thus forming chromatin loops, and anchors DNA to cellular structures like the nuclear lamina. It also defines the boundaries between active and heterochromatic DNA. Since the 3D structure of DNA influences the regulation of genes, CTCF's activity influences the expressi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transcription Factor
In molecular biology, a transcription factor (TF) (or sequence-specific DNA-binding factor) is a protein that controls the rate of transcription of genetic information from DNA to messenger RNA, by binding to a specific DNA sequence. The function of TFs is to regulate—turn on and off—genes in order to make sure that they are expressed in the desired cells at the right time and in the right amount throughout the life of the cell and the organism. Groups of TFs function in a coordinated fashion to direct cell division, cell growth, and cell death throughout life; cell migration and organization (body plan) during embryonic development; and intermittently in response to signals from outside the cell, such as a hormone. There are up to 1600 TFs in the human genome. Transcription factors are members of the proteome as well as regulome. TFs work alone or with other proteins in a complex, by promoting (as an activator), or blocking (as a repressor) the recruitment of RNA ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residue ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Insulator (genetics)
An insulator is a type of cis-regulatory element known as a long-range regulatory element. Found in multicellular eukaryotes and working over distances from the promoter element of the target gene, an insulator is typically 300 bp to 2000 bp in length. Insulators contain clustered binding sites for sequence specific DNA-binding proteins and mediate intra- and inter-chromosomal interactions. Insulators function either as an enhancer-blocker or a barrier, or both. The mechanisms by which an insulator performs these two functions include loop formation and nucleosome modifications. There are many examples of insulators, including the CTCF insulator, the ''gypsy'' insulator, and the β-globin locus. The CTCF insulator is especially important in vertebrates, while the ''gypsy'' insulator is implicated in ''Drosophila.'' The β-globin locus was first studied in chicken and then in humans for its insulator activity, both of which utilize CTCF. The genetic implications of insulators ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chromatin
Chromatin is a complex of DNA and protein found in eukaryotic cells. The primary function is to package long DNA molecules into more compact, denser structures. This prevents the strands from becoming tangled and also plays important roles in reinforcing the DNA during cell division, preventing DNA damage, and regulating gene expression and DNA replication. During mitosis and meiosis, chromatin facilitates proper segregation of the chromosomes in anaphase; the characteristic shapes of chromosomes visible during this stage are the result of DNA being coiled into highly condensed chromatin. The primary protein components of chromatin are histones. An octamer of two sets of four histone cores (Histone H2A, Histone H2B, Histone H3, and Histone H4) bind to DNA and function as "anchors" around which the strands are wound.Maeshima, K., Ide, S., & Babokhov, M. (2019). Dynamic chromatin organization without the 30-nm fiber. ''Current opinion in cell biology, 58,'' 95–104. https://doi.o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding genes, other functional regions of the genome such as regulatory sequences (see non-coding DNA), and often a substantial fraction of 'junk' DNA with no evident function. Almost all eukaryotes have mitochondria and a small mitochondrial genome. Algae and plants also contain chloroplasts with a chloroplast genome. The study of the genome is called genomics. The genomes of many organisms have been sequenced and various regions have been annotated. The International Human Genome Project reported the sequence of the genome for ''Homo sapiens'' in 200The Human Genome Project although the initial "finished" sequence was missing 8% of the genome consisting mostly of repetitive sequences. With advancements in technology that could handle sequenci ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
CpG-island
The CpG sites or CG sites are regions of DNA where a cytosine nucleotide is followed by a guanine nucleotide in the linear sequence of bases along its 5' → 3' direction. CpG sites occur with high frequency in genomic regions called CpG islands (or CG islands). Cytosines in CpG dinucleotides can be methylated to form 5-methylcytosines. Enzymes that add a methyl group are called DNA methyltransferases. In mammals, 70% to 80% of CpG cytosines are methylated. Methylating the cytosine within a gene can change its expression, a mechanism that is part of a larger field of science studying gene regulation that is called epigenetics. Methylated cytosines often mutate to thymines. In humans, about 70% of promoters located near the transcription start site of a gene (proximal promoters) contain a CpG island. CpG characteristics Definition ''CpG'' is shorthand for ''5'—C—phosphate—G—3' '', that is, cytosine and guanine separated by only one phosphate group; phosphate ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
LncRNA
Long non-coding RNAs (long ncRNAs, lncRNA) are a type of RNA, generally defined as transcripts more than 200 nucleotides that are not translated into protein. This arbitrary limit distinguishes long ncRNAs from small non-coding RNAs, such as microRNAs (miRNAs), small interfering RNAs (siRNAs), Piwi-interacting RNAs (piRNAs), small nucleolar RNAs (snoRNAs), and other short RNAs. Long intervening/intergenic noncoding RNAs (lincRNAs) are sequences of lncRNA which do not overlap protein-coding genes. Long non-coding RNAs include intergenic lincRNAs, intronic ncRNAs, and sense and antisense lncRNAs, each type showing different genomic positions in relation to genes and exons. Abundance In 2007 a study found only one-fifth of transcription across the human genome is associated with protein-coding genes, indicating at least four times more long non-coding than coding RNA sequences. Large-scale complementary DNA (cDNA) sequencing projects such as FANTOM reveal the complexity of this tr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Small RNA
Small RNA (sRNA) are polymeric RNA molecules that are less than 200 nucleotides in length, and are usually non-coding Non-coding DNA (ncDNA) sequences are components of an organism's DNA that do not encode protein sequences. Some non-coding DNA is transcribed into functional non-coding RNA molecules (e.g. transfer RNA, microRNA, piRNA, ribosomal RNA, and regula .... RNA silencing is often a function of these molecules, with the most common and well-studied example being RNA interference (RNAi), in which endogenously expressed microRNA (miRNA) or exogenously derived small interfering RNA (siRNA) induces the degradation of complementarity (molecular biology), complementary messenger RNA. Other classes of small RNA have been identified, including piwi-interacting RNA (piRNA) and its subspecies rasiRNA, repeat associated small interfering RNA (rasiRNA). Small RNA "is unable to induce RNAi alone, and to accomplish the task it must form the core of the RNA–protein complex termed the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |