|
DNA Footprinting
DNA footprinting is a method of investigating the sequence specificity of DNA-binding proteins in vitro. This technique can be used to study protein-DNA interactions both outside and within cells. The regulation of transcription has been studied extensively, and yet there is still much that is unknown. Transcription factors and associated proteins that bind promoters, enhancers, or silencers to drive or repress transcription are fundamental to understanding the unique regulation of individual genes within the genome. Techniques like DNA footprinting help elucidate which proteins bind to these associated regions of DNA and unravel the complexities of transcriptional control. History In 1978, David Galas and Albert Schmitz developed the DNA footprinting technique to study the binding specificity of the lac repressor protein. It was originally a modification of the Maxam-Gilbert chemical sequencing technique. Method The simplest application of this technique is to assess whethe ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Proteins
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residues ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
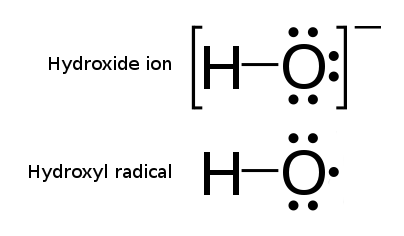
Hydroxyl Radicals
The hydroxyl radical is the diatomic molecule . The hydroxyl radical is very stable as a dilute gas, but it decays very rapidly in the condensed phase. It is pervasive in some situations. Most notably the hydroxyl radicals are produced from the decomposition of hydroperoxides (ROOH) or, in atmospheric chemistry, by the reaction of excited atomic oxygen with water. It is also important in the field of radiation chemistry, since it leads to the formation of hydrogen peroxide and oxygen, which can enhance corrosion and SCC in coolant systems subjected to radioactive environments. In organic synthesis, hydroxyl radicals are most commonly generated by photolysis of 1-hydroxy-2(1''H'')-pyridinethione. Notation The unpaired electron of the hydroxyl radical is officially represented by a middle dot, •, beside the O. Biology Hydroxyl radicals can occasionally be produced as a byproduct of immune action. Macrophages and microglia most frequently generate this compound when expose ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecular Biology
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions. The study of chemical and physical structure of biological macromolecules is known as molecular biology. Molecular biology was first described as an approach focused on the underpinnings of biological phenomena - uncovering the structures of biological molecules as well as their interactions, and how these interactions explain observations of classical biology. In 1945 the term molecular biology was used by physicist William Astbury. In 1953 Francis Crick, James Watson, Rosalind Franklin, and colleagues, working at Medical Research Council unit, Cavendish laboratory, Cambridge (now the MRC Laboratory of Molecular Biology), made a double helix model of DNA which changed the entire research scenario. They proposed the DNA structure based on previous research done by Ro ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Toeprinting Assay
The toeprinting assay, also known as the primer extension inhibition assay, is a method used in molecular biology that allows one to examine the interactions between messenger RNA and ribosomes or RNA-binding proteins. It is different from the more commonly used DNA footprinting assay. The toeprinting assay has been utilized to examine the formation of the translation initiation complex. To do a toeprint assay, one needs the mRNA of interest, ribosomes, a DNA primer, free nucleotides, and reverse transcriptase (RT), among other reagents. The assay involves letting the RT generate cDNA In genetics, complementary DNA (cDNA) is DNA synthesized from a single-stranded RNA (e.g., messenger RNA (mRNA) or microRNA (miRNA)) template in a reaction catalyzed by the enzyme reverse transcriptase. cDNA is often used to express a sp ... until it gets blocked by any bound ribosomes, resulting in shorter fragments called toeprints when the results are observed on a sequencing gel. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Footprinting
Protein footprinting is a term used to refer to a method of biochemical analysis that investigates protein structure, assembly, and interactions within a larger macromolecular assembly. It was originally coined in reference to the use of limited proteolysis to investigate contact sites within a monoclonal antibody - protein antigen complex and a year later to examine the protection from hydroxyl radical cleavage conferred by a protein bound to DNA within a DNA-protein complex. In DNA footprinting the protein is envisioned to make an imprint (or footprint) at a particular point of interaction. This latter method was adapted through the direct treatment of proteins and their complexes with hydroxyl radicals and can be generally denoted RP-MS (for Radical Probe - Mass Spectrometry) akin to the designation used for Hydrogen-deuterium exchange Mass Spectrometry (denoted HD-MS or HX-MS). Hydroxyl radical protein footprinting Time-resolved hydroxyl radical protein footprinting (HRPF) emplo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Position Weight Matrix
A position weight matrix (PWM), also known as a position-specific weight matrix (PSWM) or position-specific scoring matrix (PSSM), is a commonly used representation of motifs (patterns) in biological sequences. PWMs are often derived from a set of aligned sequences that are thought to be functionally related and have become an important part of many software tools for computational motif discovery. Background Creation Conversion of sequence to position probability matrix A PWM has one row for each symbol of the alphabet (4 rows for nucleotides in DNA sequences or 20 rows for amino acids in protein sequences) and one column for each position in the pattern. In the first step in constructing a PWM, a basic position frequency matrix (PFM) is created by counting the occurrences of each nucleotide at each position. From the PFM, a position probability matrix (PPM) can now be created by dividing that former nucleotide count at each position by the number of sequences, thereb ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hidden Markov Model
A hidden Markov model (HMM) is a statistical Markov model in which the system being modeled is assumed to be a Markov process — call it X — with unobservable ("''hidden''") states. As part of the definition, HMM requires that there be an observable process Y whose outcomes are "influenced" by the outcomes of X in a known way. Since X cannot be observed directly, the goal is to learn about X by observing Y. HMM has an additional requirement that the outcome of Y at time t=t_0 must be "influenced" exclusively by the outcome of X at t=t_0 and that the outcomes of X and Y at t handwriting recognition, handwriting, gesture recognition, part-of-speech tagging, musical score following, partial discharges and bioinformatics. Definition Let X_n and Y_n be discrete-time stochastic processes and n\geq 1. The pair (X_n,Y_n) is a ''hidden Markov model'' if * X_n is a Markov process whose behavior is not directly observable ("hidden"); * \operatorname\bigl(Y_n \i ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
FAIRE-Seq
FAIRE-Seq (Formaldehyde-Assisted Isolation of Regulatory Elements) is a method in molecular biology used for determining the sequences of DNA regions in the genome associated with regulatory activity. The technique was developed in the laboratory of Jason D. Lieb at the University of North Carolina, Chapel Hill. In contrast to DNase-Seq, the FAIRE-Seq protocol doesn't require the permeabilization of cells or isolation of nuclei, and can analyse any cell type. In a study of seven diverse human cell types, DNase-seq and FAIRE-seq produced strong cross-validation, with each cell type having 1-2% of the human genome as open chromatin. Workflow The protocol is based on the fact that the formaldehyde cross-linking is more efficient in nucleosome-bound DNA than it is in nucleosome-depleted regions of the genome. This method then segregates the non cross-linked DNA that is usually found in open chromatin, which is then sequenced. The protocol consists of cross linking, phenol extraction a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNase-Seq
DNase-seq (DNase I hypersensitive sites sequencing) is a method in molecular biology used to identify the location of regulatory regions, based on the genome-wide sequencing of regions sensitive to cleavage by DNase I. FAIRE-Seq is a successor of DNase-seq for the genome-wide identification of accessible DNA regions in the genome. Both the protocols for identifying open chromatin regions have biases depending on underlying nucleosome structure. For example, FAIRE-seq provides higher tag counts at non-promoter regions. On the other hand, DNase-seq signal is higher at promoter regions, and DNase-seq has been shown to have better sensitivity than FAIRE-seq even at non-promoter regions. DNase-seq Footprinting DNase-seq requires some downstream bioinformatics analyses in order to provide genome-wide DNA footprints. The computational tools proposed can be categorized in two classes: segmentation-based and site-centric approaches. Segmentation-based methods are based on the application ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Next-generation Sequencing
Massive parallel sequencing or massively parallel sequencing is any of several high-throughput approaches to DNA sequencing using the concept of massively parallel processing; it is also called next-generation sequencing (NGS) or second-generation sequencing. Some of these technologies emerged between 1994 and 1998 and have been commercially available since 2005. These technologies use miniaturized and parallelized platforms for sequencing of 1 million to 43 billion short reads (50 to 400 bases each) per instrument run. Many NGS platforms differ in engineering configurations and sequencing chemistry. They share the technical paradigm of massive parallel sequencing via spatially separated, clonally amplified DNA templates or single DNA molecules in a flow cytometry, flow cell. This design is very different from that of Sanger sequencing—also known as capillary sequencing or first-generation sequencing—which is based on electrophoretic separation of chain-termination products produ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Immunoprecipitation
Immunoprecipitation (IP) is the technique of precipitating a protein antigen out of solution using an antibody that specifically binds to that particular protein. This process can be used to isolate and concentrate a particular protein from a sample containing many thousands of different proteins. Immunoprecipitation requires that the antibody be coupled to a solid substrate at some point in the procedure. Types Individual protein immunoprecipitation (IP) Involves using an antibody that is specific for a known protein to isolate that particular protein out of a solution containing many different proteins. These solutions will often be in the form of a crude lysate of a plant or animal tissue. Other sample types could be body fluids or other samples of biological origin. Protein complex immunoprecipitation (Co-IP) Immunoprecipitation of intact protein complexes (i.e. antigen along with any proteins or ligands that are bound to it) is known as co-immunoprecipitation (Co-I ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |