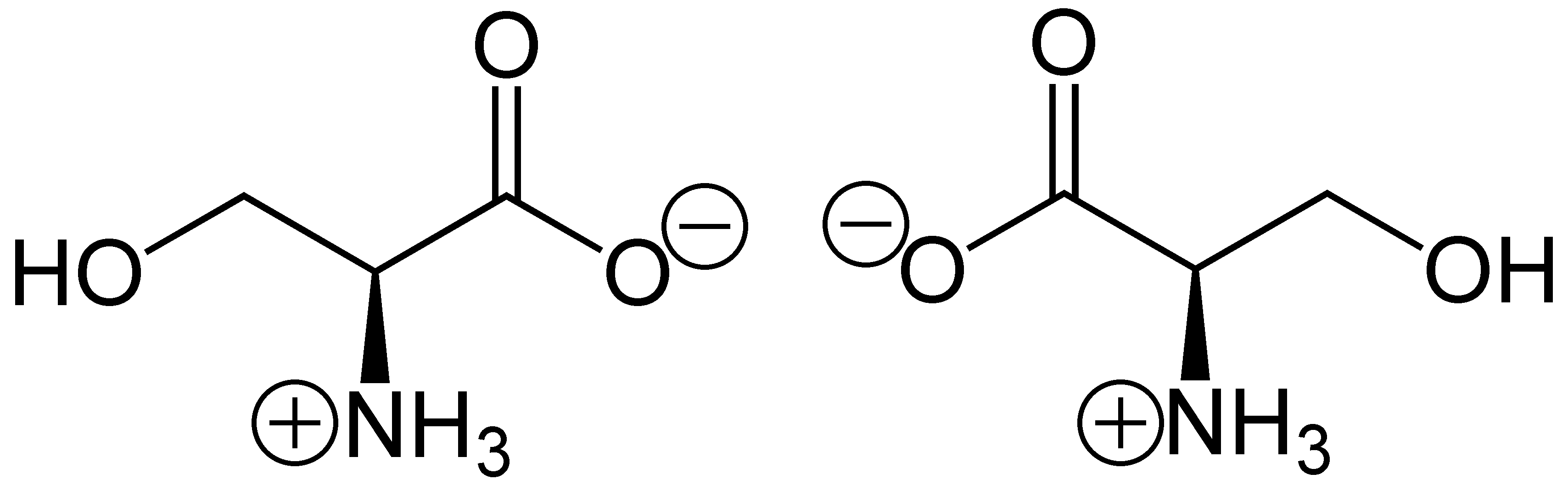|
DSP (gene)
Desmoplakin is a protein in humans that is encoded by the ''DSP'' gene. Desmoplakin is a critical component of desmosome structures in cardiac muscle and epidermal cells, which function to maintain the structural integrity at adjacent cell contacts. In cardiac muscle, desmoplakin is localized to intercalated discs which mechanically couple cardiac cells to function in a coordinated syncytial structure. Mutations in desmoplakin have been shown to play a role in dilated cardiomyopathy and arrhythmogenic right ventricular cardiomyopathy, where it may present with acute myocardial injury; striate palmoplantar keratoderma, Carvajal syndrome and paraneoplastic pemphigus. Structure Desmoplakin exists as two predominant isoforms; the first, known as "DPII", has molecular weight 260.0 kDa (2272 amino acids) and the second, known as "DPI", has molecular weight 332.0 kDa (2871 amino acids). These isoforms are identical except for the shorter rod domain in DPII. DPI is the predominant isoform ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Desmosome - 2
A desmosome (; "binding body"), also known as a macula adherens (plural: maculae adherentes) (Latin for ''adhering spot''), is a cell structure specialized for cell-to-cell adhesion. A type of junctional complex, they are localized spot-like adhesions randomly arranged on the lateral sides of plasma membranes. Desmosomes are one of the stronger cell-to-cell adhesion types and are found in tissue that experience intense mechanical stress, such as cardiac muscle tissue, bladder tissue, gastrointestinal mucosa, and epithelia. Structure Desmosomes are composed of desmosome-intermediate filament complexes (DIFC), which is a network of cadherin proteins, linker proteins and intermediate filaments. The DIFCs can be broken into three regions: the extracellular core region, or desmoglea, the outer dense plaque, or ODP, and the inner dense plaque, or IDP. The extracellular core region, approximately 34 nm in length, contains desmoglein and desmocollin, which are in the cadherin famil ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alpha Helix
The alpha helix (α-helix) is a common motif in the secondary structure of proteins and is a right hand-helix conformation in which every backbone N−H group hydrogen bonds to the backbone C=O group of the amino acid located four residues earlier along the protein sequence. The alpha helix is also called a classic Pauling–Corey–Branson α-helix. The name 3.613-helix is also used for this type of helix, denoting the average number of residues per helical turn, with 13 atoms being involved in the ring formed by the hydrogen bond. Among types of local structure in proteins, the α-helix is the most extreme and the most predictable from sequence, as well as the most prevalent. Discovery In the early 1930s, William Astbury showed that there were drastic changes in the X-ray fiber diffraction of moist wool or hair fibers upon significant stretching. The data suggested that the unstretched fibers had a coiled molecular structure with a characteristic repeat of ≈. Astb ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cardiomyocyte
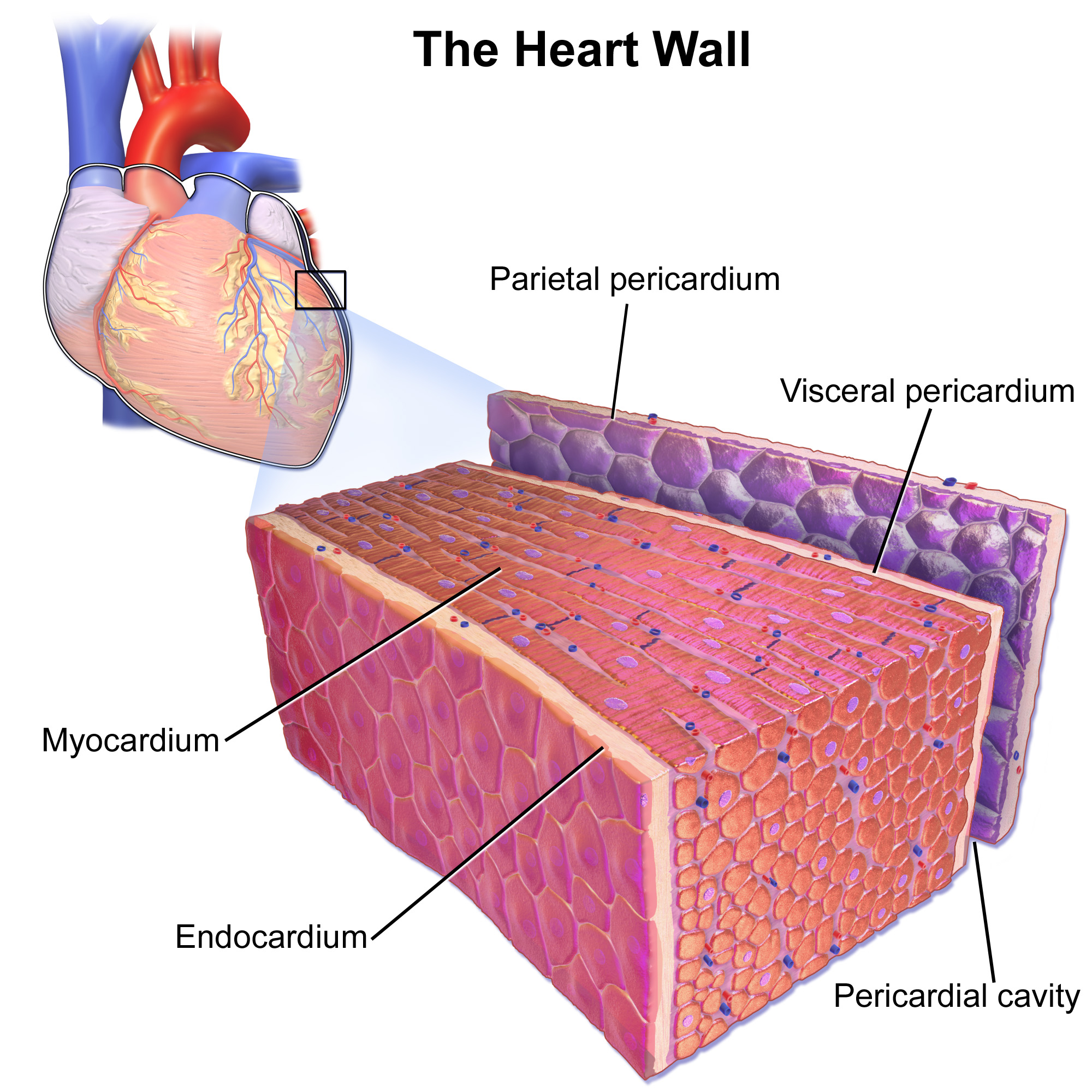
Cardiac muscle (also called heart muscle, myocardium, cardiomyocytes and cardiac myocytes) is one of three types of vertebrate muscle tissues, with the other two being skeletal muscle and smooth muscle. It is an involuntary, striated muscle that constitutes the main tissue of the wall of the heart. The cardiac muscle (myocardium) forms a thick middle layer between the outer layer of the heart wall (the pericardium) and the inner layer (the endocardium), with blood supplied via the coronary circulation. It is composed of individual cardiac muscle cells joined by intercalated discs, and encased by collagen fibers and other substances that form the extracellular matrix. Cardiac muscle contracts in a similar manner to skeletal muscle, although with some important differences. Electrical stimulation in the form of a cardiac action potential triggers the release of calcium from the cell's internal calcium store, the sarcoplasmic reticulum. The rise in calcium causes the cell's my ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Coiled-coil
A coiled coil is a structural motif in proteins in which 2–7 alpha helix, alpha-helices are coiled together like the strands of a rope. (Protein dimer, Dimers and Protein trimer, trimers are the most common types.) Many coiled coil-type proteins are involved in important biological functions, such as the regulation of gene expression — e.g., transcription factors. Notable examples are the oncoproteins c-Fos and c-Jun, as well as the muscle protein tropomyosin. Discovery The possibility of coiled coils for α-keratin was initially somewhat controversial. Linus Pauling and Francis Crick independently came to the conclusion that this was possible at about the same time. In the summer of 1952, Pauling visited the laboratory in England where Crick worked. Pauling and Crick met and spoke about various topics; at one point, Crick asked whether Pauling had considered "coiled coils" (Crick came up with the term), to which Pauling said he had. Upon returning to the United States, Paul ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phosphorylation
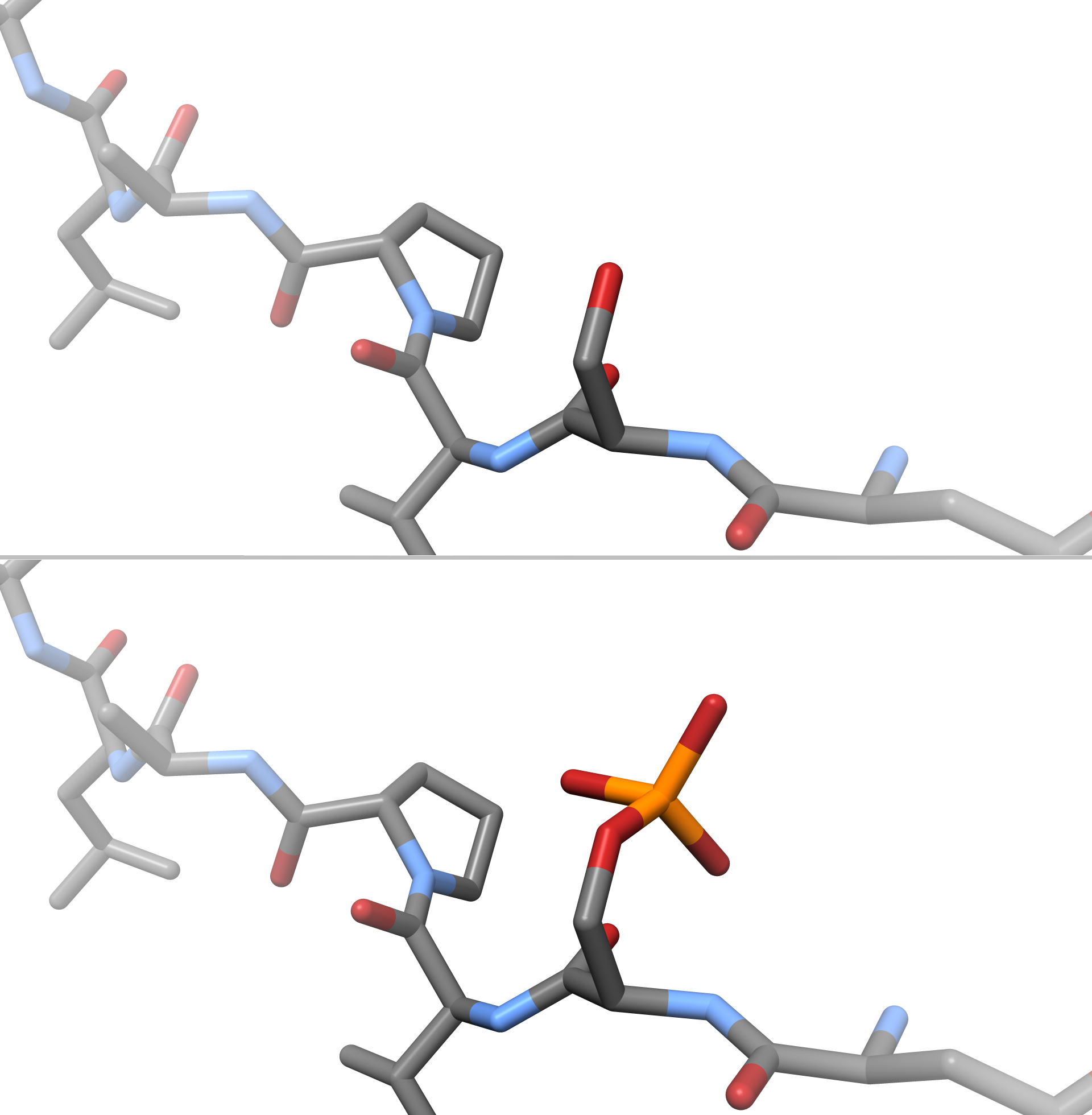
In chemistry, phosphorylation is the attachment of a phosphate group to a molecule or an ion. This process and its inverse, dephosphorylation, are common in biology and could be driven by natural selection. Text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. Protein phosphorylation often activates (or deactivates) many enzymes. Glucose Phosphorylation of sugars is often the first stage in their catabolism. Phosphorylation allows cells to accumulate sugars because the phosphate group prevents the molecules from diffusing back across their transporter. Phosphorylation of glucose is a key reaction in sugar metabolism. The chemical equation for the conversion of D-glucose to D-glucose-6-phosphate in the first step of glycolysis is given by :D-glucose + ATP → D-glucose-6-phosphate + ADP : ΔG° = −16.7 kJ/mol (° indicates measurement at standard condition) Hepatic cells are freely permeable to glucose, and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Arginine
Arginine is the amino acid with the formula (H2N)(HN)CN(H)(CH2)3CH(NH2)CO2H. The molecule features a guanidino group appended to a standard amino acid framework. At physiological pH, the carboxylic acid is deprotonated (−CO2−) and both the amino and guanidino groups are protonated, resulting in a cation. Only the -arginine (symbol Arg or R) enantiomer is found naturally. Arg residues are common components of proteins. It is encoded by the codons CGU, CGC, CGA, CGG, AGA, and AGG. The guanidine group in arginine is the precursor for the biosynthesis of nitric oxide. Like all amino acids, it is a white, water-soluble solid. History Arginine was first isolated in 1886 from yellow lupin seedlings by the German chemist Ernst Schulze and his assistant Ernst Steiger. He named it from the Greek ''árgyros'' (ἄργυρος) meaning "silver" due to the silver-white appearance of arginine nitrate crystals. In 1897, Schulze and Ernst Winterstein (1865–1949) determined the structure ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Serine
Serine (symbol Ser or S) is an α-amino acid that is used in the biosynthesis of proteins. It contains an α-amino group (which is in the protonated − form under biological conditions), a carboxyl group (which is in the deprotonated − form under biological conditions), and a side chain consisting of a hydroxymethyl group, classifying it as a polar amino acid. It can be synthesized in the human body under normal physiological circumstances, making it a nonessential amino acid. It is encoded by the codons UCU, UCC, UCA, UCG, AGU and AGC. Occurrence This compound is one of the naturally occurring proteinogenic amino acids. Only the L-stereoisomer appears naturally in proteins. It is not essential to the human diet, since it is synthesized in the body from other metabolites, including glycine. Serine was first obtained from silk protein, a particularly rich source, in 1865 by Emil Cramer. Its name is derived from the Latin for silk, ''sericum''. Serine's structure was estab ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Glycine
Glycine (symbol Gly or G; ) is an amino acid that has a single hydrogen atom as its side chain. It is the simplest stable amino acid (carbamic acid is unstable), with the chemical formula NH2‐ CH2‐ COOH. Glycine is one of the proteinogenic amino acids. It is encoded by all the codons starting with GG (GGU, GGC, GGA, GGG). Glycine is integral to the formation of alpha-helices in secondary protein structure due to its compact form. For the same reason, it is the most abundant amino acid in collagen triple-helices. Glycine is also an inhibitory neurotransmitter – interference with its release within the spinal cord (such as during a ''Clostridium tetani'' infection) can cause spastic paralysis due to uninhibited muscle contraction. It is the only achiral proteinogenic amino acid. It can fit into hydrophilic or hydrophobic environments, due to its minimal side chain of only one hydrogen atom. History and etymology Glycine was discovered in 1820 by the French chemist He ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Intermediate Filament
Intermediate filaments (IFs) are cytoskeletal structural components found in the cells of vertebrates, and many invertebrates. Homologues of the IF protein have been noted in an invertebrate, the cephalochordate ''Branchiostoma''. Intermediate filaments are composed of a family of related proteins sharing common structural and sequence features. Initially designated 'intermediate' because their average diameter (10 nm) is between those of narrower microfilaments (actin) and wider myosin filaments found in muscle cells, the diameter of intermediate filaments is now commonly compared to actin microfilaments (7 nm) and microtubules (25 nm). Animal intermediate filaments are subcategorized into six types based on similarities in amino acid sequence and protein structure. Most types are cytoplasmic, but one type, Type V is a nuclear lamin. Unlike microtubules, IF distribution in cells show no good correlation with the distribution of either mitochondria or endopla ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
C-terminus
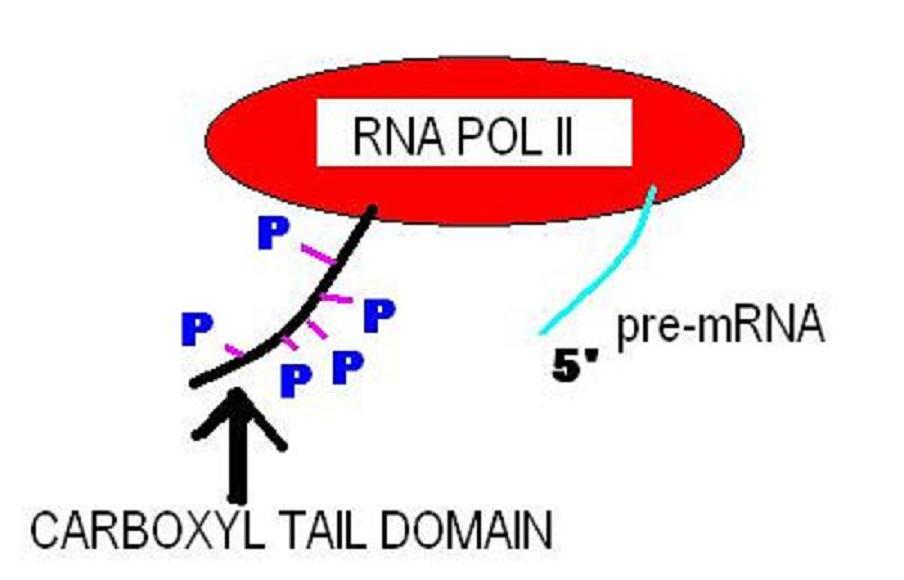
The C-terminus (also known as the carboxyl-terminus, carboxy-terminus, C-terminal tail, C-terminal end, or COOH-terminus) is the end of an amino acid chain (protein or polypeptide), terminated by a free carboxyl group (-COOH). When the protein is translated from messenger RNA, it is created from N-terminus to C-terminus. The convention for writing peptide sequences is to put the C-terminal end on the right and write the sequence from N- to C-terminus. Chemistry Each amino acid has a carboxyl group and an amine group. Amino acids link to one another to form a chain by a dehydration reaction which joins the amine group of one amino acid to the carboxyl group of the next. Thus polypeptide chains have an end with an unbound carboxyl group, the C-terminus, and an end with an unbound amine group, the N-terminus. Proteins are naturally synthesized starting from the N-terminus and ending at the C-terminus. Function C-terminal retention signals While the N-terminus of a protein often c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Small Angle X-ray Scattering
Small-angle X-ray scattering (SAXS) is a small-angle scattering technique by which nanoscale density differences in a sample can be quantified. This means that it can determine nanoparticle size distributions, resolve the size and shape of (monodisperse) macromolecules, determine pore sizes, characteristic distances of partially ordered materials, and much more. This is achieved by analyzing the elastic scattering behaviour of X-rays when travelling through the material, recording their scattering at small angles (typically 0.1 – 10°, hence the "Small-angle" in its name). It belongs to the family of small-angle scattering (SAS) techniques along with small-angle neutron scattering, and is typically done using hard X-rays with a wavelength of 0.07 – 0.2 nm.. Depending on the angular range in which a clear scattering signal can be recorded, SAXS is capable of delivering structural information of dimensions between 1 and 100 nm, and of repeat distances in partially ordered sys ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |