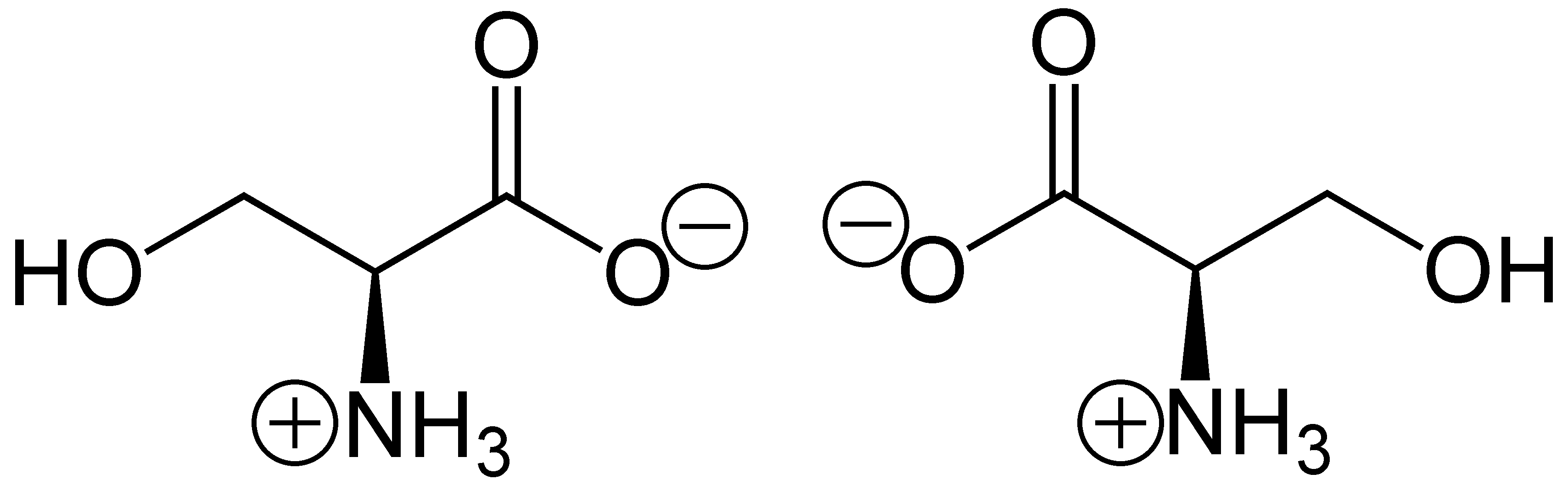|
C2orf16
C2orf16 is a protein that in humans is encoded by the C2orf16 gene. Isoform 2 of this protein (NCBI ID: CAH18189.1 henceforth referred to as C2orf16) is 1,984 amino acids long. The gene contains 1 exon and is located at 2p23.3. Aliases for C2orf16 include Open Reading Frame 16 on Chromosome 2 and P-S-E-R-S-H-H-S Repeats Containing Sequence. 68 orthologs are known for this gene, including in mice and sheep, but no paralogs have been found. Gene The C2orf16 isoform 2 is a 6.2 kb, 1 exon gene at locus 2p23.3, and contains P-S-E-R-S-H-H-S repeats on the C-terminal side of the gene from amino acid 1,559 to 1,903. These repeats appear to have arisen from a transposable element. Primates show more P-S-E-R-S-H-H-S repeats than other mammalian orthologs do. Expression C2orf16 is found to be highly expressed in the testes and a retinoic acid and mitogen-treated Embryonic stem cell, human embryonic stem cell line, but is not known to be expressed differently in age or disease phenotypes ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Orthologs
Sequence homology is the biological homology between DNA, RNA, or protein sequences, defined in terms of shared ancestry in the evolutionary history of life. Two segments of DNA can have shared ancestry because of three phenomena: either a speciation event (orthologs), or a duplication event (paralogs), or else a horizontal (or lateral) gene transfer event (xenologs). Homology among DNA, RNA, or proteins is typically inferred from their nucleotide or amino acid sequence similarity. Significant similarity is strong evidence that two sequences are related by evolutionary changes from a common ancestral sequence. Alignments of multiple sequences are used to indicate which regions of each sequence are homologous. Identity, similarity, and conservation The term "percent homology" is often used to mean "sequence similarity”, that is the percentage of identical residues (''percent identity''), or the percentage of residues conserved with similar physicochemical properties ('' ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
3'UTR
In molecular genetics, the three prime untranslated region (3′-UTR) is the section of messenger RNA (mRNA) that immediately follows the translation termination codon. The 3′-UTR often contains regulatory regions that post-transcriptionally influence gene expression. During gene expression, an mRNA molecule is transcribed from the DNA sequence and is later translated into a protein. Several regions of the mRNA molecule are not translated into a protein including the 5' cap, 5' untranslated region, 3′ untranslated region and poly(A) tail. Regulatory regions within the 3′-untranslated region can influence polyadenylation, translation efficiency, localization, and stability of the mRNA. The 3′-UTR contains both binding sites for regulatory proteins as well as microRNAs (miRNAs). By binding to specific sites within the 3′-UTR, miRNAs can decrease gene expression of various mRNAs by either inhibiting translation or directly causing degradation of the transcript. Th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transcription Factor
In molecular biology, a transcription factor (TF) (or sequence-specific DNA-binding factor) is a protein that controls the rate of transcription of genetic information from DNA to messenger RNA, by binding to a specific DNA sequence. The function of TFs is to regulate—turn on and off—genes in order to make sure that they are expressed in the desired cells at the right time and in the right amount throughout the life of the cell and the organism. Groups of TFs function in a coordinated fashion to direct cell division, cell growth, and cell death throughout life; cell migration and organization (body plan) during embryonic development; and intermittently in response to signals from outside the cell, such as a hormone. There are up to 1600 TFs in the human genome. Transcription factors are members of the proteome as well as regulome. TFs work alone or with other proteins in a complex, by promoting (as an activator), or blocking (as a repressor) the recruitment of RNA ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Secondary Structure
Protein secondary structure is the three dimensional conformational isomerism, form of ''local segments'' of proteins. The two most common Protein structure#Secondary structure, secondary structural elements are alpha helix, alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary structure elements typically spontaneously form as an intermediate before the protein protein folding, folds into its three dimensional protein tertiary structure, tertiary structure. Secondary structure is formally defined by the pattern of hydrogen bonds between the Amine, amino hydrogen and carboxyl oxygen atoms in the peptide backbone chain, backbone. Secondary structure may alternatively be defined based on the regular pattern of backbone Dihedral angle#Dihedral angles of proteins, dihedral angles in a particular region of the Ramachandran plot regardless of whether it has the correct hydrogen bonds. The concept of secondary structure was first introduced by Kaj Ulrik ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
C2orf16 Isoform 2 Predicted 3D Structure
C, or c, is the third letter in the Latin alphabet, used in the modern English alphabet, the alphabets of other western European languages and others worldwide. Its name in English is ''cee'' (pronounced ), plural ''cees''. History "C" comes from the same letter as "G". The Semites named it gimel. The sign is possibly adapted from an Egyptian hieroglyph for a staff sling, which may have been the meaning of the name ''gimel''. Another possibility is that it depicted a camel, the Semitic name for which was ''gamal''. Barry B. Powell, a specialist in the history of writing, states "It is hard to imagine how gimel = "camel" can be derived from the picture of a camel (it may show his hump, or his head and neck!)". In the Etruscan language, plosive consonants had no contrastive voicing, so the Greek ' Γ' (Gamma) was adopted into the Etruscan alphabet to represent . Already in the Western Greek alphabet, Gamma first took a '' form in Early Etruscan, then '' in Classical Etru ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nuclear Export Sequence
A nuclear export signal (NES) is a short target peptide containing 4 hydrophobic residues in a protein that targets it for export from the cell nucleus to the cytoplasm through the nuclear pore complex using nuclear transport. It has the opposite effect of a nuclear localization signal, which targets a protein located in the cytoplasm for import to the nucleus. The NES is recognized and bound by exportins. NESs serve several vital cellular functions. They assist in regulating the position of proteins within the cell. Through this NESs affect transcription and several other nuclear functions that are essential to proper cell function. The export of many types of RNA from the nucleus is required for proper cellular function. The NES determines what type of pathway the varying types of RNA may use to exit the nucleus and perform their function and the NESs may effect the directionality of molecules exiting the nucleus. Structure Computer analysis of known NESs found the most common ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nuclear Localization Sequence
A nuclear localization signal ''or'' sequence (NLS) is an amino acid sequence that 'tags' a protein for import into the cell nucleus by nuclear transport. Typically, this signal consists of one or more short sequences of positively charged lysines or arginines exposed on the protein surface. Different nuclear localized proteins may share the same NLS. An NLS has the opposite function of a nuclear export signal (NES), which targets proteins out of the nucleus. Types Classical These types of NLSs can be further classified as either monopartite or bipartite. The major structural differences between the two are that the two basic amino acid clusters in bipartite NLSs are separated by a relatively short spacer sequence (hence bipartite - 2 parts), while monopartite NLSs are not. The first NLS to be discovered was the sequence PKKKRKV in the SV40 Large T-antigen (a monopartite NLS). The NLS of nucleoplasmin, KR AATKKAGQAKKK, is the prototype of the ubiquitous bipartite signal: two cluster ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transmembrane Domains
A transmembrane domain (TMD) is a membrane-spanning protein domain. TMDs generally adopt an alpha helix topological conformation, although some TMDs such as those in porins can adopt a different conformation. Because the interior of the lipid bilayer is hydrophobic, the amino acid residues in TMDs are often hydrophobic, although proteins such as membrane pumps and ion channels can contain polar residues. TMDs vary greatly in length, sequence, and hydrophobicity, adopting organelle-specific properties. Functions of transmembrane domains Transmembrane domains are known to perform a variety of functions. These include: * Anchoring transmembrane proteins to the membrane. *Facilitating molecular transport of molecules such as ions and proteins across biological membranes; usually hydrophilic residues and binding sites in the TMDs help in this process. * Signal transduction across the membrane; many transmembrane proteins, such as G protein-coupled receptors, receive extracellula ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Histidine
Histidine (symbol His or H) is an essential amino acid that is used in the biosynthesis of proteins. It contains an α-amino group (which is in the protonated –NH3+ form under biological conditions), a carboxylic acid group (which is in the deprotonated –COO− form under biological conditions), and an imidazole side chain (which is partially protonated), classifying it as a positively charged amino acid at physiological pH. Initially thought essential only for infants, it has now been shown in longer-term studies to be essential for adults also. It is encoded by the codons CAU and CAC. Histidine was first isolated by Albrecht Kossel and Sven Gustaf Hedin in 1896. It is also a precursor to histamine, a vital inflammatory agent in immune responses. The acyl radical is histidyl. Properties of the imidazole side chain The conjugate acid (protonated form) of the imidazole side chain in histidine has a p''K''a of approximately 6.0. Thus, below a pH of 6, the imidazole ring ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Serine
Serine (symbol Ser or S) is an α-amino acid that is used in the biosynthesis of proteins. It contains an α-amino group (which is in the protonated − form under biological conditions), a carboxyl group (which is in the deprotonated − form under biological conditions), and a side chain consisting of a hydroxymethyl group, classifying it as a polar amino acid. It can be synthesized in the human body under normal physiological circumstances, making it a nonessential amino acid. It is encoded by the codons UCU, UCC, UCA, UCG, AGU and AGC. Occurrence This compound is one of the naturally occurring proteinogenic amino acids. Only the L-stereoisomer appears naturally in proteins. It is not essential to the human diet, since it is synthesized in the body from other metabolites, including glycine. Serine was first obtained from silk protein, a particularly rich source, in 1865 by Emil Cramer. Its name is derived from the Latin for silk, ''sericum''. Serine's structure was estab ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |

.png)