|
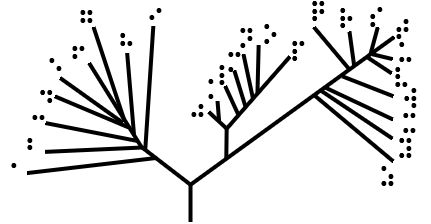
Cladogram
A cladogram (from Greek ''clados'' "branch" and ''gramma'' "character") is a diagram used in cladistics to show relations among organisms. A cladogram is not, however, an evolutionary tree because it does not show how ancestors are related to descendants, nor does it show how much they have changed, so many differing evolutionary trees can be consistent with the same cladogram. A cladogram uses lines that branch off in different directions ending at a clade, a group of organisms with a last common ancestor. There are many shapes of cladograms but they all have lines that branch off from other lines. The lines can be traced back to where they branch off. These branching off points represent a hypothetical ancestor (not an actual entity) which can be inferred to exhibit the traits shared among the terminal taxa above it. This hypothetical ancestor might then provide clues about the order of evolution of various features, adaptation, and other evolutionary narratives about an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cladograms
A cladogram (from Greek ''clados'' "branch" and ''gramma'' "character") is a diagram used in cladistics to show relations among organisms. A cladogram is not, however, an evolutionary tree because it does not show how ancestors are related to descendants, nor does it show how much they have changed, so many differing evolutionary trees can be consistent with the same cladogram. A cladogram uses lines that branch off in different directions ending at a clade, a group of organisms with a last common ancestor. There are many shapes of cladograms but they all have lines that branch off from other lines. The lines can be traced back to where they branch off. These branching off points represent a hypothetical ancestor (not an actual entity) which can be inferred to exhibit the traits shared among the terminal taxa above it. This hypothetical ancestor might then provide clues about the order of evolution of various features, adaptation, and other evolutionary narratives about ance ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cladistics
Cladistics (; ) is an approach to biological classification in which organisms are categorized in groups (" clades") based on hypotheses of most recent common ancestry. The evidence for hypothesized relationships is typically shared derived characteristics ( synapomorphies'')'' that are not present in more distant groups and ancestors. However, from an empirical perspective, common ancestors are inferences based on a cladistic hypothesis of relationships of taxa whose character states can be observed. Theoretically, a last common ancestor and all its descendants constitute a (minimal) clade. Importantly, all descendants stay in their overarching ancestral clade. For example, if the terms ''worms'' or ''fishes'' were used within a ''strict'' cladistic framework, these terms would include humans. Many of these terms are normally used paraphyletically, outside of cladistics, e.g. as a ' grade', which are fruitless to precisely delineate, especially when including extinct species. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Clade
A clade (), also known as a monophyletic group or natural group, is a group of organisms that are monophyletic – that is, composed of a common ancestor and all its lineal descendants – on a phylogenetic tree. Rather than the English term, the equivalent Latin term ''cladus'' (plural ''cladi'') is often used in taxonomical literature. The common ancestor may be an individual, a population, or a species (extinct or extant). Clades are nested, one in another, as each branch in turn splits into smaller branches. These splits reflect evolutionary history as populations diverged and evolved independently. Clades are termed monophyletic (Greek: "one clan") groups. Over the last few decades, the cladistic approach has revolutionized biological classification and revealed surprising evolutionary relationships among organisms. Increasingly, taxonomists try to avoid naming taxa that are not clades; that is, taxa that are not monophyletic. Some of the relationships between org ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phylogenetic Tree
A phylogenetic tree (also phylogeny or evolutionary tree Felsenstein J. (2004). ''Inferring Phylogenies'' Sinauer Associates: Sunderland, MA.) is a branching diagram or a tree showing the evolutionary relationships among various biological species or other entities based upon similarities and differences in their physical or genetic characteristics. All life on Earth is part of a single phylogenetic tree, indicating common ancestry. In a ''rooted'' phylogenetic tree, each node with descendants represents the inferred most recent common ancestor of those descendants, and the edge lengths in some trees may be interpreted as time estimates. Each node is called a taxonomic unit. Internal nodes are generally called hypothetical taxonomic units, as they cannot be directly observed. Trees are useful in fields of biology such as bioinformatics, systematics, and phylogenetics. ''Unrooted'' trees illustrate only the relatedness of the leaf nodes and do not require the ancestral root ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Plesiomorphy
In phylogenetics, a plesiomorphy ("near form") and symplesiomorphy are synonyms for an ancestral character shared by all members of a clade, which does not distinguish the clade from other clades. Plesiomorphy, symplesiomorphy, apomorphy, and synapomorphy, all mean a trait shared between species because they share an ancestral species. Apomorphic and synapomorphic characteristics convey much information about evolutionary clades and can be used to define taxa. However, plesiomorphic and symplesiomorphic characteristics cannot. The term ''symplesiomorphy'' was introduced in 1950 by German entomologist Willi Hennig. Examples A backbone is a plesiomorphic trait shared by birds and mammals, and does not help in placing an animal in one or the other of these two clades. Birds and mammals share this trait because both clades are descended from the same far distant ancestor. Other clades, e.g. snakes, lizards, turtles, fish, frogs, all have backbones and none are either birds no ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Homoplasy
Homoplasy, in biology and phylogenetics, is the term used to describe a feature that has been gained or lost independently in separate lineages over the course of evolution. This is different from homology, which is the term used to characterize the similarity of features that can be parsimoniously explained by common ancestry. Homoplasy can arise from both similar selection pressures acting on adapting species, and the effects of genetic drift. Most often, homoplasy is viewed as a similarity in morphological characters. However, homoplasy may also appear in other character types, such as similarity in the genetic sequence, life cycle types or even behavioral traits. Etymology The term homoplasy was first used by Ray Lankester in 1870. The corresponding adjective is either ''homoplasic'' or ''homoplastic''. It is derived from the two Ancient Greek words (), meaning "similar, alike, the same", and (), meaning "to shape, to mold". Parallelism and convergence Parallel a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Synapomorphy
In phylogenetics, an apomorphy (or derived trait) is a novel character or character state that has evolved from its ancestral form (or plesiomorphy). A synapomorphy is an apomorphy shared by two or more taxa and is therefore hypothesized to have evolved in their most recent common ancestor. ) In cladistics, synapomorphy implies homology. Examples of apomorphy are the presence of erect gait, fur, the evolution of three middle ear bones, and mammary glands in mammals but not in other vertebrate animals such as amphibians or reptiles, which have retained their ancestral traits of a sprawling gait and lack of fur. Thus, these derived traits are also synapomorphies of mammals in general as they are not shared by other vertebrate animals. Etymology The word —coined by German entomologist Willi Hennig—is derived from the Ancient Greek words (''sún''), meaning "with, together"; (''apó''), meaning "away from"; and (''morphḗ''), meaning "shape, form". Clade ana ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Maximum Parsimony (phylogenetics)
In phylogenetics, maximum parsimony is an optimality criterion under which the phylogenetic tree that minimizes the total number of character-state changes (or miminizes the cost of differentially weighted character-state changes) is preferred. Under the maximum-parsimony criterion, the optimal tree will minimize the amount of homoplasy (i.e., convergent evolution, parallel evolution, and evolutionary reversals). In other words, under this criterion, the shortest possible tree that explains the data is considered best. Some of the basic ideas behind maximum parsimony were presented by James S. Farris in 1970 and Walter M. Fitch in 1971. Maximum parsimony is an intuitive and simple criterion, and it is popular for this reason. However, although it is easy to ''score'' a phylogenetic tree (by counting the number of character-state changes), there is no algorithm to quickly ''generate'' the most-parsimonious tree. Instead, the most-parsimonious tree must be sought in "tree s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bayesian Inference
Bayesian inference is a method of statistical inference in which Bayes' theorem is used to update the probability for a hypothesis as more evidence or information becomes available. Bayesian inference is an important technique in statistics, and especially in mathematical statistics. Bayesian updating is particularly important in the Sequential analysis, dynamic analysis of a sequence of data. Bayesian inference has found application in a wide range of activities, including science, engineering, philosophy, medicine, sport, and law. In the philosophy of decision theory, Bayesian inference is closely related to subjective probability, often called "Bayesian probability". Introduction to Bayes' rule Formal explanation Bayesian inference derives the posterior probability as a consequence relation, consequence of two Antecedent (logic), antecedents: a prior probability and a "likelihood function" derived from a statistical model for the observed data. Bayesian inference computes ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Maximum Parsimony (phylogenetics)
In phylogenetics, maximum parsimony is an optimality criterion under which the phylogenetic tree that minimizes the total number of character-state changes (or miminizes the cost of differentially weighted character-state changes) is preferred. Under the maximum-parsimony criterion, the optimal tree will minimize the amount of homoplasy (i.e., convergent evolution, parallel evolution, and evolutionary reversals). In other words, under this criterion, the shortest possible tree that explains the data is considered best. Some of the basic ideas behind maximum parsimony were presented by James S. Farris in 1970 and Walter M. Fitch in 1971. Maximum parsimony is an intuitive and simple criterion, and it is popular for this reason. However, although it is easy to ''score'' a phylogenetic tree (by counting the number of character-state changes), there is no algorithm to quickly ''generate'' the most-parsimonious tree. Instead, the most-parsimonious tree must be sought in "tree s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Neighbor-joining
In bioinformatics, neighbor joining is a bottom-up (agglomerative) clustering method for the creation of phylogenetic trees, created by Naruya Saitou and Masatoshi Nei in 1987. Usually based on DNA or protein sequence data, the algorithm requires knowledge of the distance between each pair of taxa (e.g., species or sequences) to create the phylogenetic tree. The algorithm Neighbor joining takes a distance matrix, which specifies the distance between each pair of taxa, as input. The algorithm starts with a completely unresolved tree, whose topology corresponds to that of a star network, and iterates over the following steps, until the tree is completely resolved, and all branch lengths are known: # Based on the current distance matrix, calculate a matrix Q (defined below). # Find the pair of distinct taxa i and j (i.e. with i \neq j) for which Q(i,j) is smallest. Make a new node that joins the taxa i and j, and connect the new node to the central node. For example, in part (B ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |