|
ChIP-exo Process Diagram
ChIP-exo is a chromatin immunoprecipitation based method for mapping the locations at which a protein of interest (transcription factor) binds to the genome. It is a modification of the ChIP-seq protocol, improving the resolution of binding sites from hundreds of base pairs to almost one base pair. It employs the use of exonucleases to degrade strands of the protein-bound DNA in the 5'-3' direction to within a small number of nucleotides of the protein binding site. The nucleotides of the exonuclease-treated ends are determined using some combination of DNA sequencing, microarrays, and PCR. These sequences are then mapped to the genome to identify the locations on the genome at which the protein binds. __TOC__ Theory Chromatin immunoprecipitation (ChIP) techniques have been in use since 1984 to detect protein-DNA interactions. There have been many variations on ChIP to improve the quality of results. One such improvement, ChIP-on-chip (ChIP-chip), combines ChIP with microarray ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Exonuclease
Exonucleases are enzymes that work by cleaving nucleotides one at a time from the end (exo) of a polynucleotide chain. A hydrolyzing reaction that breaks phosphodiester bonds at either the 3′ or the 5′ end occurs. Its close relative is the endonuclease, which cleaves phosphodiester bonds in the middle (endo) of a polynucleotide chain. Eukaryotes and prokaryotes have three types of exonucleases involved in the normal turnover of mRNA: 5′ to 3′ exonuclease (Xrn1), which is a dependent decapping protein; 3′ to 5′ exonuclease, an independent protein; and poly(A)-specific 3′ to 5′ exonuclease. In both archaea and eukaryotes, one of the main routes of RNA degradation is performed by the multi-protein exosome complex, which consists largely of 3′ to 5′ exoribonucleases. Significance to polymerase RNA polymerase II is known to be in effect during transcriptional termination; it works with a 5' exonuclease (human gene Xrn2) to degrade the newly formed tra ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecular Biology Techniques
A molecule is a group of two or more atoms held together by attractive forces known as chemical bonds; depending on context, the term may or may not include ions which satisfy this criterion. In quantum physics, organic chemistry, and biochemistry, the distinction from ions is dropped and ''molecule'' is often used when referring to polyatomic ions. A molecule may be homonuclear, that is, it consists of atoms of one chemical element, e.g. two atoms in the oxygen molecule (O2); or it may be heteronuclear, a chemical compound composed of more than one element, e.g. water (molecule), water (two hydrogen atoms and one oxygen atom; H2O). In the kinetic theory of gases, the term ''molecule'' is often used for any gaseous particle regardless of its composition. This relaxes the requirement that a molecule contains two or more atoms, since the noble gases are individual atoms. Atoms and complexes connected by non-covalent interactions, such as hydrogen bonds or ionic bonds, are typic ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecular Biology
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions. The study of chemical and physical structure of biological macromolecules is known as molecular biology. Molecular biology was first described as an approach focused on the underpinnings of biological phenomena - uncovering the structures of biological molecules as well as their interactions, and how these interactions explain observations of classical biology. In 1945 the term molecular biology was used by physicist William Astbury. In 1953 Francis Crick, James Watson, Rosalind Franklin, and colleagues, working at Medical Research Council unit, Cavendish laboratory, Cambridge (now the MRC Laboratory of Molecular Biology), made a double helix model of DNA which changed the entire research scenario. They proposed the DNA structure based on previous research done by ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Methods
Protein methods are the techniques used to study proteins. There are experimental methods for studying proteins (e.g., for detecting proteins, for isolating and purifying proteins, and for characterizing the structure and function of proteins, often requiring that the protein first be purified). Computational methods typically use computer programs to analyze proteins. However, many experimental methods (e.g., mass spectrometry) require computational analysis of the raw data. Genetic methods Experimental analysis of proteins typically requires expression and purification of proteins. Expression is achieved by manipulating DNA that encodes the protein(s) of interest. Hence, protein analysis usually requires DNA methods, especially cloning. Some examples of genetic methods include conceptual translation, Site-directed mutagenesis, using a fusion protein, and matching allele with disease states. Some proteins have never been directly sequenced, however by translating codons from kn ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein-DNA Interaction
DNA-binding proteins are proteins that have DNA-binding domains and thus have a specific or general affinity for single- or double-stranded DNA. Sequence-specific DNA-binding proteins generally interact with the major groove of B-DNA, because it exposes more functional groups that identify a base pair. However, there are some known minor groove DNA-binding ligands such as netropsin, distamycin, Hoechst 33258, pentamidine, DAPI and others. Examples DNA-binding proteins include transcription factors which modulate the process of transcription, various polymerases, nucleases which cleave DNA molecules, and histones which are involved in chromosome packaging and transcription in the cell nucleus. DNA-binding proteins can incorporate such domains as the zinc finger, the helix-turn-helix, and the leucine zipper (among many others) that facilitate binding to nucleic acid. There are also more unusual examples such as transcription activator like effectors. Non-specific DNA-prote ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chip-sequencing
ChIP-sequencing, also known as ChIP-seq, is a method used to analyze protein interactions with DNA. ChIP-seq combines chromatin immunoprecipitation (ChIP) with massively parallel DNA sequencing to identify the binding sites of DNA-associated proteins. It can be used to map global binding sites precisely for any protein of interest. Previously, ChIP-on-chip was the most common technique utilized to study these protein–DNA relations. Uses ChIP-seq is primarily used to determine how transcription factors and other chromatin-associated proteins influence phenotype-affecting mechanisms. Determining how proteins interact with DNA to regulate gene expression is essential for fully understanding many biological processes and disease states. This epigenetic information is complementary to genotype and expression analysis. ChIP-seq technology is currently seen primarily as an alternative to ChIP-chip which requires a hybridization array. This introduces some bias, as an array is restr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chromatin Immunoprecipitation
Chromatin immunoprecipitation (ChIP) is a type of immunoprecipitation experimental technique used to investigate the interaction between proteins and DNA in the cell. It aims to determine whether specific proteins are associated with specific genomic regions, such as transcription factors on promoters or other DNA binding sites, and possibly define cistromes. ChIP also aims to determine the specific location in the genome that various histone modifications are associated with, indicating the target of the histone modifiers. ChIP is crucial for the advancements in the field of epigenomics and learning more about epigenetic phenomena. Briefly, the conventional method is as follows: # DNA and associated proteins on chromatin in living cells or tissues are crosslinked (this step is omitted in Native ChIP). # The DNA-protein complexes (chromatin-protein) are then sheared into ~500 bp DNA fragments by sonication or nuclease digestion. # Cross-linked DNA fragments associated with the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
High Throughput Sequencing
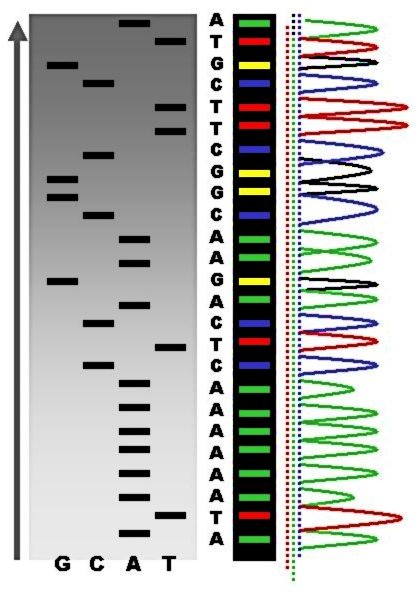
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery. Knowledge of DNA sequences has become indispensable for basic biological research, DNA Genographic Projects and in numerous applied fields such as medical diagnosis, biotechnology, forensic biology, virology and biological systematics. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers, characterize antibody repertoire, and can be used to guide patient treatment. Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged. The rapid speed of sequencing attained with mo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cross-link
In chemistry and biology a cross-link is a bond or a short sequence of bonds that links one polymer chain to another. These links may take the form of covalent bonds or ionic bonds and the polymers can be either synthetic polymers or natural polymers (such as proteins). In polymer chemistry "cross-linking" usually refers to the use of cross-links to promote a change in the polymers' physical properties. When "crosslinking" is used in the biological field, it refers to the use of a probe to link proteins together to check for protein–protein interactions, as well as other creative cross-linking methodologies. Although the term is used to refer to the "linking of polymer chains" for both sciences, the extent of crosslinking and specificities of the crosslinking agents vary greatly. As with all science, there are overlaps, and the following delineations are a starting point to understanding the subtleties. Polymer chemistry Crosslinking is the general term for the process of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sonication
A sonicator at the Weizmann Institute of Science during sonicationSonication is the act of applying sound energy to agitate particles in a sample, for various purposes such as the extraction of multiple compounds from plants, microalgae and seaweeds. Ultrasonic frequencies (> 20 kHz) are usually used, leading to the process also being known as ultrasonication or ultra-sonication. In the laboratory, it is usually applied using an ''ultrasonic bath'' or an '' ultrasonic probe'', colloquially known as a ''sonicator''. In a paper machine, an ultrasonic foil can distribute cellulose fibres more uniformly and strengthen the paper. Effects Sonication has numerous effects, both chemical and physical. The chemical effects of ultrasound are concerned with understanding the effect of sonic waves on chemical systems, this is called sonochemistry. The chemical effects of ultrasound do not come from a direct interaction with molecular species. Studies have shown that no direct c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nuclease Protection Assay
Nuclease protection assay is a laboratory technique used in biochemistry and genetics to identify individual RNA molecules in a heterogeneous RNA sample extracted from cells. The technique can identify one or more RNA molecules of known sequence even at low total concentration. The extracted RNA is first mixed with antisense RNA or DNA probes that are complementary to the sequence or sequences of interest and the complementary strands are hybridized to form double-stranded RNA (or a DNA-RNA hybrid). The mixture is then exposed to ribonucleases that specifically cleave only ''single''-stranded RNA but have no activity against double-stranded RNA. When the reaction runs to completion, susceptible RNA regions are degraded to very short oligomers or to individual nucleotides; the surviving RNA fragments are those that were complementary to the added antisense strand and thus contained the sequence of interest. Probe The probes are prepared by cloning part of the gene of interest ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |