|
CBL (gene)
''Cbl'' (named after Casitas B-lineage Lymphoma) is a mammalian gene encoding the protein CBL which is an E3 ubiquitin-protein ligase involved in cell signalling and protein ubiquitination. Mutations to this gene have been implicated in a number of human cancers, particularly acute myeloid leukaemia. Discovery In 1989 a virally encoded portion of the chromosomal mouse ''Cbl'' gene was the first member of the Cbl family to be discovered and was named ''v-Cbl'' to distinguish it from normal mouse ''c-Cbl''. The virus used in the experiment was a mouse-tropic strain of Murine leukemia virus isolated from the brain of a mouse captured at Lake Casitas, California known as ''Cas-Br-M'', and was found to have excised approximately a third of the original ''c-Cbl'' gene from a mouse into which it was injected. Sequencing revealed that the portion carried by the retrovirus encoded a ''tyrosine kinase binding domain'', and that this was the oncogenic form as retroviruses carrying full ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin Ligase
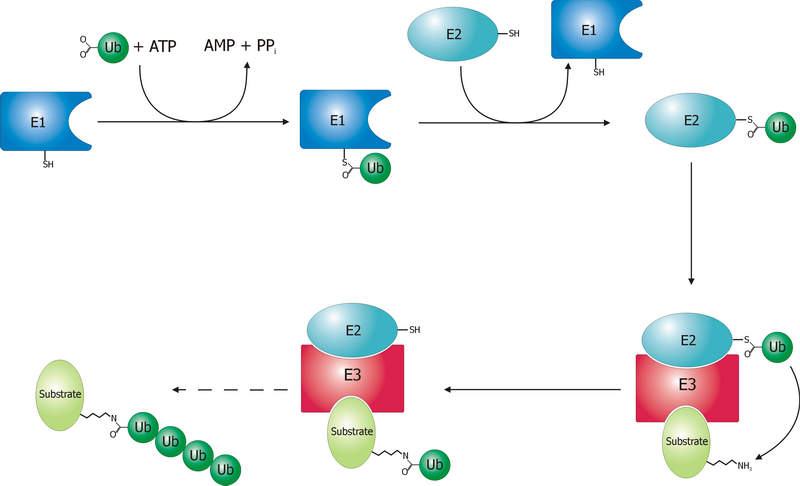
A ubiquitin ligase (also called an E3 ubiquitin ligase) is a protein that recruits an E2 ubiquitin-conjugating enzyme that has been loaded with ubiquitin, recognizes a protein substrate, and assists or directly catalyzes the transfer of ubiquitin from the E2 to the protein substrate. In simple and more general terms, the ligase enables movement of ubiquitin from a ubiquitin carrier to another thing (the substrate) by some mechanism. The ubiquitin, once it reaches its destination, ends up being attached by an isopeptide bond to a lysine residue, which is part of the target protein. E3 ligases interact with both the target protein and the E2 enzyme, and so impart substrate specificity to the E2. Commonly, E3s polyubiquitinate their substrate with Lys48-linked chains of ubiquitin, targeting the substrate for destruction by the proteasome. However, many other types of linkages are possible and alter a protein's activity, interactions, or localization. Ubiquitination by E3 ligases r ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Homology (biology)
In biology, homology is similarity due to shared ancestry between a pair of structures or genes in different taxa. A common example of homologous structures is the forelimbs of vertebrates, where the wings of bats and birds, the arms of primates, the front flippers of whales and the forelegs of four-legged vertebrates like dogs and crocodiles are all derived from the same ancestral tetrapod structure. Evolutionary biology explains homologous structures adapted to different purposes as the result of descent with modification from a common ancestor. The term was first applied to biology in a non-evolutionary context by the anatomist Richard Owen in 1843. Homology was later explained by Charles Darwin's theory of evolution in 1859, but had been observed before this, from Aristotle onwards, and it was explicitly analysed by Pierre Belon in 1555. In developmental biology, organs that developed in the embryo in the same manner and from similar origins, such as from matc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
C-Met
c-Met, also called tyrosine-protein kinase Met or hepatocyte growth factor receptor (HGFR), is a protein that in humans is encoded by the ''MET'' gene. The protein possesses tyrosine kinase activity. The primary single chain precursor protein is post-translationally cleaved to produce the alpha and beta subunits, which are disulfide linked to form the mature receptor. MET is a single pass tyrosine kinase receptor essential for embryonic development, organogenesis and wound healing. Hepatocyte growth factor/Scatter Factor (HGF/SF) and its splicing isoform (NK1, NK2) are the only known ligands of the MET receptor. MET is normally expressed by cells of epithelial origin, while expression of HGF/SF is restricted to cells of mesenchymal origin. When HGF/SF binds its cognate receptor MET it induces its dimerization through a not yet completely understood mechanism leading to its activation. Abnormal MET activation in cancer correlates with poor prognosis, where aberrantly active MET ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Abl Gene
Tyrosine-protein kinase ABL1 also known as ABL1 is a protein that, in humans, is encoded by the ''ABL1'' gene (previous symbol ''ABL'') located on chromosome 9. c-Abl is sometimes used to refer to the version of the gene found within the mammalian genome, while v-Abl refers to the viral gene, which was initially isolated from the Abelson murine leukemia virus. Function The ''ABL1'' proto-oncogene encodes a cytoplasmic and nuclear protein tyrosine kinase that has been implicated in processes of cell differentiation, cell division, cell adhesion, and stress response such as DNA repair. Activity of ABL1 protein is negatively regulated by its SH3 domain, and deletion of the SH3 domain turns ABL1 into an oncogene. The t(9;22) translocation results in the head-to-tail fusion of the '' BCR'' and ''ABL1'' genes, leading to a fusion gene present in many cases of chronic myelogenous leukemia. The DNA-binding activity of the ubiquitously expressed ABL1 tyrosine kinase is regulated b ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Lysosome
A lysosome () is a membrane-bound organelle found in many animal cells. They are spherical vesicles that contain hydrolytic enzymes that can break down many kinds of biomolecules. A lysosome has a specific composition, of both its membrane proteins, and its lumenal proteins. The lumen's pH (~4.5–5.0) is optimal for the enzymes involved in hydrolysis, analogous to the activity of the stomach. Besides degradation of polymers, the lysosome is involved in various cell processes, including secretion, plasma membrane repair, apoptosis, cell signaling, and energy metabolism. Lysosomes act as the waste disposal system of the cell by digesting used materials in the cytoplasm, from both inside and outside the cell. Material from outside the cell is taken up through endocytosis, while material from the inside of the cell is digested through autophagy. The sizes of the organelles vary greatly—the larger ones can be more than 10 times the size of the smaller ones. They we ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Receptor Tyrosine Kinase
Receptor tyrosine kinases (RTKs) are the high- affinity cell surface receptors for many polypeptide growth factors, cytokines, and hormones. Of the 90 unique tyrosine kinase genes identified in the human genome, 58 encode receptor tyrosine kinase proteins. Receptor tyrosine kinases have been shown not only to be key regulators of normal cellular processes but also to have a critical role in the development and progression of many types of cancer. Mutations in receptor tyrosine kinases lead to activation of a series of signalling cascades which have numerous effects on protein expression. Receptor tyrosine kinases are part of the larger family of protein tyrosine kinases, encompassing the receptor tyrosine kinase proteins which contain a transmembrane domain, as well as the non-receptor tyrosine kinases which do not possess transmembrane domains. History The first RTKs to be discovered were EGF and NGF in the 1960s, but the classification of receptor tyrosine kinases was ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ligase
In biochemistry, a ligase is an enzyme that can catalyze the joining ( ligation) of two large molecules by forming a new chemical bond. This is typically via hydrolysis of a small pendant chemical group on one of the larger molecules or the enzyme catalyzing the linking together of two compounds, e.g., enzymes that catalyze joining of C-O, C-S, C-N, etc. In general, a ligase catalyzes the following reaction: :Ab + C → A–C + b or sometimes :Ab + cD → A–D + b + c + d + e + f where the lowercase letters can signify the small, dependent groups. Ligase can join two complementary fragments of nucleic acid and repair single stranded breaks that arise in double stranded DNA during replication. Nomenclature The common names of ligases often include the word "ligase", such as DNA ligase, an enzyme commonly used in molecular biology laboratories to join together DNA fragments. Other common names for ligases include the word "synthetase", because they are used to synt ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin
Ubiquitin is a small (8.6 kDa) regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ''ubiquitously''. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 1980s. Four genes in the human genome code for ubiquitin: UBB, UBC, UBA52 and RPS27A. The addition of ubiquitin to a substrate protein is called ubiquitylation (or, alternatively, ubiquitination or ubiquitinylation). Ubiquitylation affects proteins in many ways: it can mark them for degradation via the proteasome, alter their cellular location, affect their activity, and promote or prevent protein interactions. Ubiquitylation involves three main steps: activation, conjugation, and ligation, performed by ubiquitin-activating enzymes (E1s), ubiquitin-conjugating enzymes (E2s), and ubiquitin ligases (E3s), respectively. The result of this sequential cascade is to bind ubiquitin to lysine residues on the protein substrate via an isopeptide ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Caenorhabditis Elegans
''Caenorhabditis elegans'' () is a free-living transparent nematode about 1 mm in length that lives in temperate soil environments. It is the type species of its genus. The name is a blend of the Greek ''caeno-'' (recent), ''rhabditis'' (rod-like) and Latin ''elegans'' (elegant). In 1900, Maupas initially named it ''Rhabditides elegans.'' Osche placed it in the subgenus ''Caenorhabditis'' in 1952, and in 1955, Dougherty raised ''Caenorhabditis'' to the status of genus. ''C. elegans'' is an unsegmented pseudocoelomate and lacks respiratory or circulatory systems. Most of these nematodes are hermaphrodites and a few are males. Males have specialised tails for mating that include spicules. In 1963, Sydney Brenner proposed research into ''C. elegans,'' primarily in the area of neuronal development. In 1974, he began research into the molecular and developmental biology of ''C. elegans'', which has since been extensively used as a model organism. It was the first multice ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Drosophila Melanogaster
''Drosophila melanogaster'' is a species of fly (the taxonomic order Diptera) in the family Drosophilidae. The species is often referred to as the fruit fly or lesser fruit fly, or less commonly the " vinegar fly" or " pomace fly". Starting with Charles W. Woodworth's 1901 proposal of the use of this species as a model organism, ''D. melanogaster'' continues to be widely used for biological research in genetics, physiology, microbial pathogenesis, and life history evolution. As of 2017, five Nobel Prizes have been awarded to drosophilists for their work using the insect. ''D. melanogaster'' is typically used in research owing to its rapid life cycle, relatively simple genetics with only four pairs of chromosomes, and large number of offspring per generation. It was originally an African species, with all non-African lineages having a common origin. Its geographic range includes all continents, including islands. ''D. melanogaster'' is a common pest in homes, restaurants, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |