|
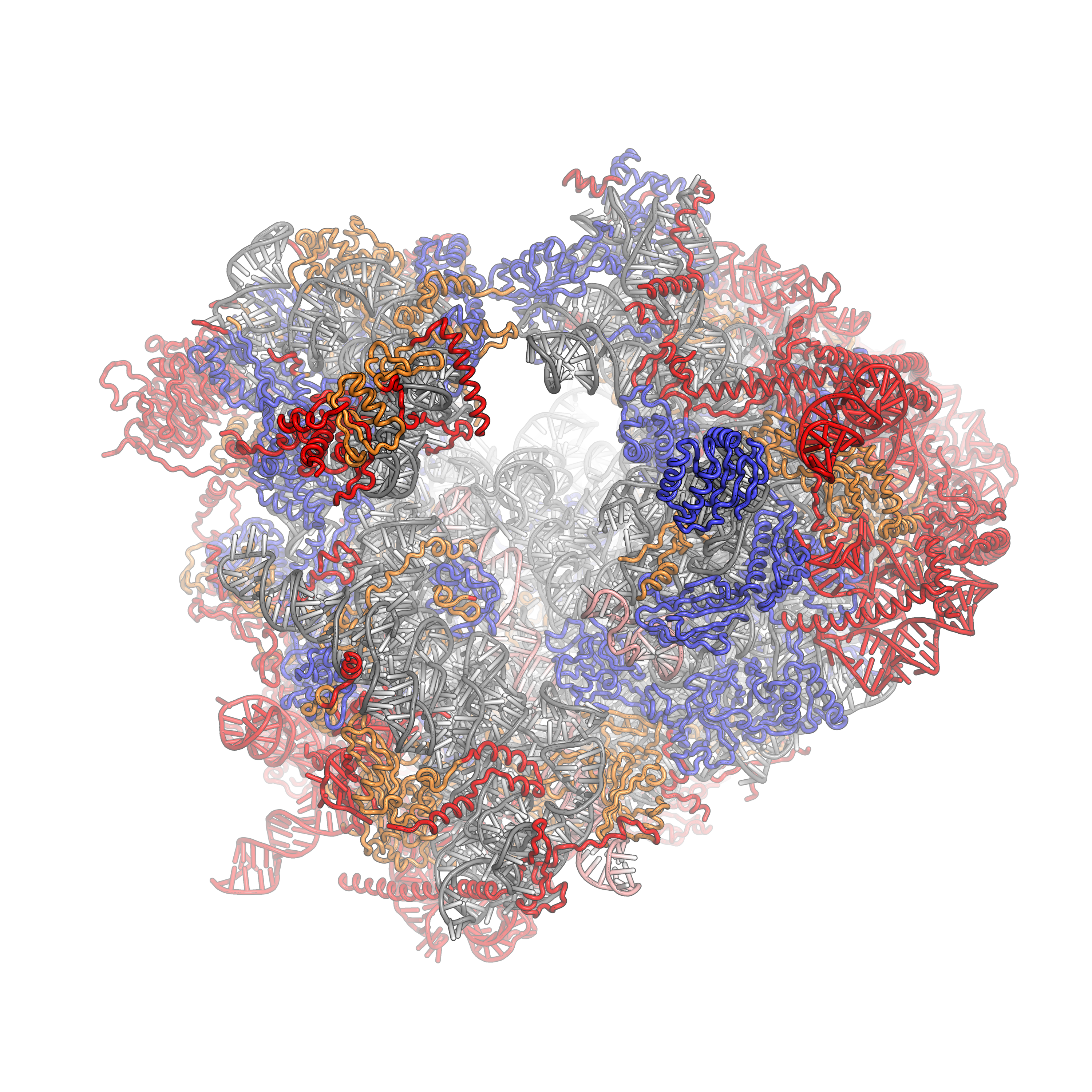
43S Preinitiation Complex
The 43S preinitiation complex (43S PIC) is a ribonucleoprotein complex that exists during an early step of eukaryotic translation initiation. The 43S PIC contains the small ribosomal subunit (40S) bound by the initiation factors eIF1, eIF1A, eIF3, and the eIF2-Met-tRNAiMet-GTP ternary complex (eIF2-TC). Function The 43S is an important intermediate complex during cap-dependent translation initiation. In the canonical model of translation initiation, the 43S PIC is pre-formed as a stable complex and recruited to the 5' cap of eukaryotic messenger RNAs (mRNAs) by the eIF4F complex. The 43S PIC then "scans" in the 5' --> 3' direction along the mRNA in an ATP-dependent fashion (via eIF4A and/or other RNA helicases such as Ded1/DDX3 and DHX29) to locate the start codon. Start codon recognition occurs through base-pairing between the Met-tRNAiMet and AUG in the ribosomal P-site and a number of associated changes, and is followed by joining of the large 60S ribosomal subunit to form ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribonucleoprotein
Nucleoproteins are proteins conjugated with nucleic acids (either DNA or RNA). Typical nucleoproteins include ribosomes, nucleosomes and viral nucleocapsid proteins. Structures Nucleoproteins tend to be positively charged, facilitating interaction with the negatively charged nucleic acid chains. The tertiary structures and biological functions of many nucleoproteins are understood.Graeme K. Hunter G. K. (2000): Vital Forces. The discovery of the molecular basis of life. Academic Press, London 2000, . Important techniques for determining the structures of nucleoproteins include X-ray diffraction, nuclear magnetic resonance and cryo-electron microscopy. Viruses Virus genomes (either DNA or RNA) are extremely tightly packed into the viral capsid. Many viruses are therefore little more than an organised collection of nucleoproteins with their binding sites pointing inwards. Structurally characterised viral nucleoproteins include influenza, rabies, Ebola, Bunyamwera, Schmall ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Helicase
Helicases are a class of enzymes thought to be vital to all organisms. Their main function is to unpack an organism's genetic material. Helicases are motor proteins that move directionally along a nucleic acid phosphodiester backbone, separating two hybridized nucleic acid strands (hence '' helic- + -ase''), using energy from ATP hydrolysis. There are many helicases, representing the great variety of processes in which strand separation must be catalyzed. Approximately 1% of eukaryotic genes code for helicases. The human genome codes for 95 non-redundant helicases: 64 RNA helicases and 31 DNA helicases. Many cellular processes, such as DNA replication, transcription, translation, recombination, DNA repair, and ribosome biogenesis involve the separation of nucleic acid strands that necessitates the use of helicases. Some specialized helicases are also involved in sensing of viral nucleic acids during infection and fulfill a immunological function. Function Helicases ar ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA-binding Proteins
RNA-binding proteins (often abbreviated as RBPs) are proteins that bind to the double or single stranded RNA in cells and participate in forming ribonucleoprotein complexes. RBPs contain various structural motifs, such as RNA recognition motif (RRM), dsRNA binding domain, zinc finger and others. They are cytoplasmic and nuclear proteins. However, since most mature RNA is exported from the nucleus relatively quickly, most RBPs in the nucleus exist as complexes of protein and pre-mRNA called heterogeneous ribonucleoprotein particles (hnRNPs). RBPs have crucial roles in various cellular processes such as: cellular function, transport and localization. They especially play a major role in post-transcriptional control of RNAs, such as: splicing, polyadenylation, mRNA stabilization, mRNA localization and translation. Eukaryotic cells express diverse RBPs with unique RNA-binding activity and protein–protein interaction. According to the Eukaryotic RBP Database (EuRBPDB), there are 2 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Expression
Gene expression is the process by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, protein or non-coding RNA, and ultimately affect a phenotype, as the final effect. These products are often proteins, but in non-protein-coding genes such as transfer RNA (tRNA) and small nuclear RNA (snRNA), the product is a functional non-coding RNA. Gene expression is summarized in the central dogma of molecular biology first formulated by Francis Crick in 1958, further developed in his 1970 article, and expanded by the subsequent discoveries of reverse transcription and RNA replication. The process of gene expression is used by all known life— eukaryotes (including multicellular organisms), prokaryotes (bacteria and archaea), and utilized by viruses—to generate the macromolecular machinery for life. In genetics, gene expression is the most fundamental level at which the genotype gives rise to the phen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecular Biology
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions. The study of chemical and physical structure of biological macromolecules is known as molecular biology. Molecular biology was first described as an approach focused on the underpinnings of biological phenomena - uncovering the structures of biological molecules as well as their interactions, and how these interactions explain observations of classical biology. In 1945 the term molecular biology was used by physicist William Astbury. In 1953 Francis Crick, James Watson, Rosalind Franklin, and colleagues, working at Medical Research Council unit, Cavendish laboratory, Cambridge (now the MRC Laboratory of Molecular Biology), made a double helix model of DNA which changed the entire research scenario. They proposed the DNA structure based on previous research done by ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cross-link
In chemistry and biology a cross-link is a bond or a short sequence of bonds that links one polymer chain to another. These links may take the form of covalent bonds or ionic bonds and the polymers can be either synthetic polymers or natural polymers (such as proteins). In polymer chemistry "cross-linking" usually refers to the use of cross-links to promote a change in the polymers' physical properties. When "crosslinking" is used in the biological field, it refers to the use of a probe to link proteins together to check for protein–protein interactions, as well as other creative cross-linking methodologies. Although the term is used to refer to the "linking of polymer chains" for both sciences, the extent of crosslinking and specificities of the crosslinking agents vary greatly. As with all science, there are overlaps, and the following delineations are a starting point to understanding the subtleties. Polymer chemistry Crosslinking is the general term for the process of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cryo-electron Microscopy
Cryogenic electron microscopy (cryo-EM) is a cryomicroscopy technique applied on samples cooled to cryogenic temperatures. For biological specimens, the structure is preserved by embedding in an environment of vitreous ice. An aqueous sample solution is applied to a grid-mesh and plunge-frozen in liquid ethane or a mixture of liquid ethane and propane. While development of the technique began in the 1970s, recent advances in detector technology and software algorithms have allowed for the determination of biomolecular structures at near-atomic resolution. This has attracted wide attention to the approach as an alternative to X-ray crystallography or NMR spectroscopy for macromolecular structure determination without the need for crystallization. In 2017, the Nobel Prize in Chemistry was awarded to Jacques Dubochet, Joachim Frank, and Richard Henderson "for developing cryo-electron microscopy for the high-resolution structure determination of biomolecules in solution." ''Nature ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Eukaryotic Ribosome (80S)
Ribosomes are a large and complex molecular machine that catalyzes the synthesis of proteins, referred to as translation. The ribosome selects aminoacylated transfer RNAs (tRNAs) based on the sequence of a protein-encoding messenger RNA (mRNA) and covalently links the amino acids into a polypeptide chain. Ribosomes from all organisms share a highly conserved catalytic center. However, the ribosomes of eukaryotes (animals, plants, fungi, and large number unicellular organisms all with a nucleus) are much larger than prokaryotic (bacterial and archaeal) ribosomes and subject to more complex regulation and biogenesis pathways. Eukaryotic ribosomes are also known as 80S ribosomes, referring to their sedimentation coefficients in Svedberg units, because they sediment faster than the prokaryotic ( 70S) ribosomes. Eukaryotic ribosomes have two unequal subunits, designated small subunit (40S) and large subunit (60S) according to their sedimentation coefficients. Both subunits contain ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Eukaryotic Large Ribosomal Subunit (60S)
Ribosomal particles are denoted according to their sedimentation coefficients in Svedberg units. The 60S subunit is the large subunit of eukaryotic 80S ribosomes. It is structurally and functionally related to the 50S subunit of 70S prokaryotic ribosomes. However, the 60S subunit is much larger than the prokaryotic 50S subunit and contains many additional protein segments, as well as ribosomal RNA expansion segments. Overall structure Characteristic features of the large subunit, shown below in the "Crown View", include the central protuberance (CP) and the two stalks, which are named according to their bacterial protein components (L1 stalk on the left as seen from the subunit interface and L7/L12 on the right). There are three binding sites for tRNA, the A-site, P-site and E-site (see article on protein translation for details). The core of the 60S subunit is formed by the 28S ribosomal RNA (abbreviated 28S rRNA), which is homologous to the prokaryotic 23S rRNA, which also c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
P-site
The P-site (for peptidyl) is the second binding site for tRNA in the ribosome. The other two sites are the A-site (aminoacyl), which is the first binding site in the ribosome, and the E-site (exit), the third. During protein translation, the P-site holds the tRNA which is linked to the growing polypeptide chain. When a stop codon is reached, the peptidyl-tRNA bond of the tRNA located in the P-site is cleaved releasing the newly synthesized protein. During the translocation step of the elongation phase, the mRNA is advanced by one codon, coupled to movement of the tRNAs from the ribosomal A to P and P to E sites, catalyzed by elongation factor EF-G. Overview The ribosomal P-site plays a vital role in all phases of translation. Initiation involves recognition of the start codon (AUG) by initiator tRNA in the P-site, elongation involves passage of many elongator tRNAs through the P site, termination involves hydrolysis of the mature polypeptide from tRNA bound to the P-site, and riboso ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DHX29
DExH-box helicase 29 (DHX29) is a 155 kDa protein that in humans is encoded by the DHX29 gene. Function This gene encodes a member of the DEAH (Asp-Glu-Ala-His) subfamily of proteins, part of the DEAD (Asp-Glu-Ala-Asp) box family of RNA helicases. The encoded protein functions in translation initiation, and is specifically required for ribosomal scanning across stable mRNA secondary structures during initiation codon selection. This protein may also play a role in sensing virally derived cytosolic nucleic acids. Knockdown of this gene results in reduced protein translation and impaired proliferation of cancer cells. Interactions DHX29 has been shown to interact with the eukaryotic small ribosomal subunit (40S) The eukaryotic small ribosomal subunit (40S) is the smaller subunit of the eukaryotic 80S ribosomes, with the other major component being the large ribosomal subunit (60S). The "40S" and "60S" names originate from the convention that ribosomal pa ... and eI ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DDX3X
ATP-dependent RNA helicase DDX3X is an enzyme that in humans is encoded by the ''DDX3X'' gene. Function DEAD box proteins, characterized by the conserved motif Asp-Glu-Ala-Asp (DEAD), are putative RNA helicases. They are implicated in a number of cellular processes involving alteration of RNA secondary structure such as translation initiation, nuclear and mitochondrial splicing, and ribosome and spliceosome assembly. Based on their distribution patterns, some members of this family are believed to be involved in embryogenesis, spermatogenesis, and cellular growth and division. This gene encodes a DEAD box protein, which interacts specifically with hepatitis C virus core protein resulting a change in intracellular location. This gene has a homolog located in the nonrecombining region of the Y chromosome. The protein sequence is 91% identical between this gene and the Y-linked homolog. Sub-cellular trafficking DDX3X performs its functions in the cell nucleus and cytoplasm, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |