|
2 Base Encoding
2 Base Encoding, also called SOLiD (sequencing by oligonucleotide ligation and detection), is a next-generation sequencing technology developed by Applied Biosystems and has been commercially available since 2008. These technologies generate hundreds of thousands of small sequence reads at one time. Well-known examples of such DNA sequencing methods include 454 pyrosequencing (introduced in 2005), the Solexa system (introduced in 2006) and the SOLiD system (introduced in 2007). These methods have reduced the cost from $0.01/base in 2004 to nearly $0.0001/base in 2006 and increased the sequencing capacity from 1,000,000 bases/machine/day in 2004 to more than 100,000,000 bases/machine/day in 2006. 2-base encoding is based on ligation sequencing rather than sequencing by synthesis. However, instead of using fluorescent labeled 9-mer probes that distinguish only 6 bases, 2-base encoding takes advantage of fluorescent labeled 8-mer probes that distinguish the two 3 prime most bases but ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Next-generation Sequencing
Massive parallel sequencing or massively parallel sequencing is any of several high-throughput approaches to DNA sequencing using the concept of massively parallel processing; it is also called next-generation sequencing (NGS) or second-generation sequencing. Some of these technologies emerged between 1994 and 1998 and have been commercially available since 2005. These technologies use miniaturized and parallelized platforms for sequencing of 1 million to 43 billion short reads (50 to 400 bases each) per instrument run. Many NGS platforms differ in engineering configurations and sequencing chemistry. They share the technical paradigm of massive parallel sequencing via spatially separated, clonally amplified DNA templates or single DNA molecules in a flow cell. This design is very different from that of Sanger sequencing—also known as capillary sequencing or first-generation sequencing—which is based on electrophoretic separation of chain-termination products produced in indi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Applied Biosystems
Applied Biosystems is one of various brands under the Life Technologies brand of Thermo Fisher Scientific corporation. The brand is focused on integrated systems for genetic analysis, which include computerized machines and the consumables used within them (such as reagents). In 2008, a merger between Applied Biosystems and Invitrogen was finalized, creating Life Technologies. The latter was acquired by Thermo Fisher Scientific in 2014. Prior to 2008, the Applied Biosystems brand was owned by various entities in a corporate group parented by PerkinElmer. The roots of Applied Biosystems trace back to GeneCo (Genetic Systems Company), a pioneer biotechnology company founded in 1981 in Foster City, California.Applied Biosystems Timeline , AppliedBiosystems.com Through the 1980s and early 1 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
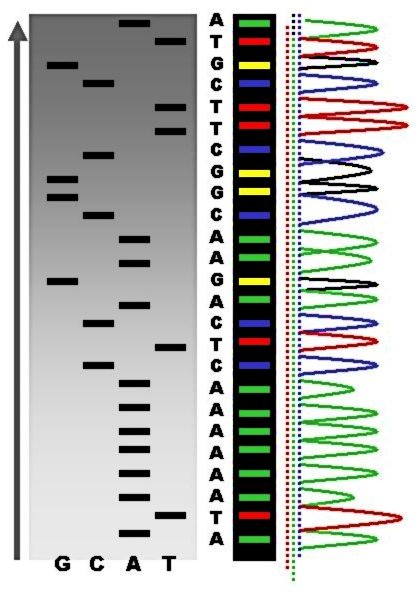
DNA Sequencing
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery. Knowledge of DNA sequences has become indispensable for basic biological research, DNA Genographic Projects and in numerous applied fields such as medical diagnosis, biotechnology, forensic biology, virology and biological systematics. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers, characterize antibody repertoire, and can be used to guide patient treatment. Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged. The rapid speed of sequencing attained with modern ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Pyrosequencing
Pyrosequencing is a method of DNA sequencing (determining the order of nucleotides in DNA) based on the "sequencing by synthesis" principle, in which the sequencing is performed by detecting the nucleotide incorporated by a DNA polymerase. Pyrosequencing relies on light detection based on a chain reaction when pyrophosphate is released. Hence, the name pyrosequencing. The principle of pyrosequencing was first described in 1993 by, Bertil Pettersson, Mathias Uhlen and Pål Nyren by combining the solid phase sequencing method using streptavidin coated magnetic beads with recombinant DNA polymerase lacking 3´to 5´exonuclease activity (proof-reading) and luminescence detection using the firefly luciferase enzyme. A mixture of three enzymes (DNA polymerase, ATP sulfurylase and firefly luciferase) and a nucleotide (dNTP) are added to single stranded DNA to be sequenced and the incorporation of nucleotide is followed by measuring the light emitted. The intensity of the light determ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polony Sequencing
Polony sequencing is an inexpensive but highly accurate multiplex sequencing technique that can be used to “read” millions of immobilized DNA sequences in parallel. This technique was first developed by Dr. George Church's group at Harvard Medical School. Unlike other sequencing techniques, Polony sequencing technology is an open platform with freely downloadable, open source software and protocols. Also, the hardware of this technique can be easily set up with a commonly available epifluorescence microscopy and a computer-controlled flowcell/fluidics system. Polony sequencing is generally performed on paired-end tags library that each molecule of DNA template is of 135 bp in length with two 17–18 bp paired genomic tags separated and flanked by common sequences. The current read length of this technique is 26 bases per amplicon and 13 bases per tag, leaving a gap of 4–5 bases in each tag. Workflow The protocol of Polony sequencing can be broken into three main parts, w ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequencing By Ligation
Sequencing by ligation is a DNA sequencing method that uses the enzyme DNA ligase to identify the nucleotide present at a given position in a DNA sequence. Unlike most currently popular DNA sequencing methods, this method does not use a DNA polymerase to create a second strand. Instead, the mismatch sensitivity of a DNA ligase enzyme is used to determine the underlying sequence of the target DNA molecule. Process DNA ligase is an enzyme that joins together ends of DNA molecules. Although commonly represented as joining two pairs of ends at once, as in the ligation of restriction enzyme fragments, ligase can also join the ends on only one of the two strands (for example, when the other strand is already continuous or lacks a terminal phosphate necessary for ligation). DNA ligase is sensitive to the structure of DNA and has very low efficiency when there are mismatches between the bases of the two strands. Sequencing by ligation relies upon the sensitivity of DNA ligase for base-p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Single Molecule Real Time Sequencing
Single-molecule real-time (SMRT) sequencing is a parallelized single molecule DNA sequencing method. Single-molecule real-time sequencing utilizes a zero-mode waveguide (ZMW). A single DNA polymerase enzyme is affixed at the bottom of a ZMW with a single molecule of DNA as a template. The ZMW is a structure that creates an illuminated observation volume that is small enough to observe only a single nucleotide of DNA being incorporated by DNA polymerase. Each of the four DNA bases is attached to one of four different fluorescent dyes. When a nucleotide is incorporated by the DNA polymerase, the fluorescent tag is cleaved off and diffuses out of the observation area of the ZMW where its fluorescence is no longer observable. A detector detects the fluorescent signal of the nucleotide incorporation, and the base call is made according to the corresponding fluorescence of the dye. Technology The DNA sequencing is done on a chip that contains many ZMWs. Inside each ZMW, a single active ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Biotechnology
Biotechnology is the integration of natural sciences and engineering sciences in order to achieve the application of organisms, cells, parts thereof and molecular analogues for products and services. The term ''biotechnology'' was first used by Károly Ereky in 1919, meaning the production of products from raw materials with the aid of living organisms. Definition The concept of biotechnology encompasses a wide range of procedures for modifying living organisms according to human purposes, going back to domestication of animals, cultivation of the plants, and "improvements" to these through breeding programs that employ artificial selection and hybridization. Modern usage also includes genetic engineering as well as cell and tissue culture technologies. The American Chemical Society defines biotechnology as the application of biological organisms, systems, or processes by various industries to learning about the science of life and the improvement of the value of materi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |