|
Restriction Fragment Length Polymorphism
In molecular biology, restriction fragment length polymorphism (RFLP) is a technique that exploits variations in homologous DNA sequences, known as polymorphisms, populations, or species or to pinpoint the locations of genes within a sequence. The term may refer to a polymorphism itself, as detected through the differing locations of restriction enzyme sites, or to a related laboratory technique by which such differences can be illustrated. In RFLP analysis, a DNA sample is digested into fragments by one or more restriction enzymes, and the resulting ''restriction fragments'' are then separated by gel electrophoresis according to their size. RFLP analysis is now largely obsolete due to the emergence of inexpensive DNA sequencing technologies, but it was the first DNA profiling technique inexpensive enough to see widespread application. RFLP analysis was an important early tool in genome mapping, localization of genes for genetic disorders, determination of risk for disease, an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecular Biology
Molecular biology is a branch of biology that seeks to understand the molecule, molecular basis of biological activity in and between Cell (biology), cells, including biomolecule, biomolecular synthesis, modification, mechanisms, and interactions. Though cells and other microscopic structures had been observed in living organisms as early as the 18th century, a detailed understanding of the mechanisms and interactions governing their behavior did not emerge until the 20th century, when technologies used in physics and chemistry had advanced sufficiently to permit their application in the biological sciences. The term 'molecular biology' was first used in 1945 by the English physicist William Astbury, who described it as an approach focused on discerning the underpinnings of biological phenomena—i.e. uncovering the physical and chemical structures and properties of biological molecules, as well as their interactions with other molecules and how these interactions explain observ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Southern Blot
Southern blot is a method used for detection and quantification of a specific DNA sequence in DNA samples. This method is used in molecular biology. Briefly, purified DNA from a biological sample (such as blood or tissue) is digested with restriction enzymes, and the resulting DNA fragments are separated by electrophoresis using an electric current to move them through a sieve-like gel or matrix, which allows smaller fragments to move faster than larger fragments. The DNA fragments are transferred out of the gel or matrix onto a solid membrane, which is then exposed to a DNA probe labeled with a radioactive, fluorescent, or chemical tag. The tag allows any DNA fragments containing complementary sequences with the DNA probe sequence to be visualized within the Southern blot. The Southern blotting combines the transfer of electrophoresis-separated DNA fragments to a filter membrane in a process called blotting, and the subsequent fragment detection by probe hybridization. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
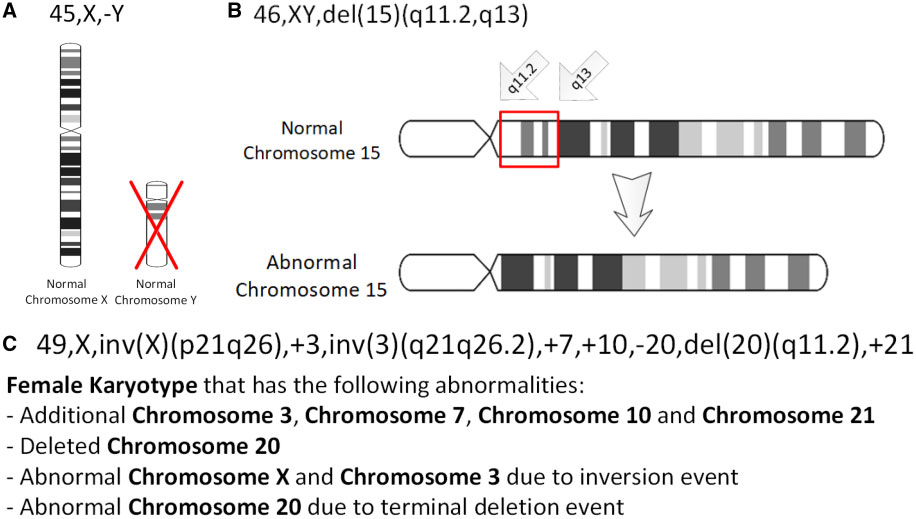
Genetic Deletion
In genetics, a deletion (also called gene deletion, deficiency, or deletion mutation) (sign: Δ) is a mutation (a genetic aberration) in which a part of a chromosome or a sequence of DNA is left out during DNA replication. Any number of nucleotides can be deleted, from a single base to an entire piece of chromosome. Some chromosomes have fragile spots where breaks occur, which result in the deletion of a part of the chromosome. The breaks can be induced by heat, viruses, radiation, or chemical reactions. When a chromosome breaks, if a part of it is deleted or lost, the missing piece of chromosome is referred to as a deletion or a deficiency. For synapsis to occur between a chromosome with a large intercalary deficiency and a normal complete homolog, the unpaired region of the normal homolog must loop out of the linear structure into a deletion or compensation loop. The smallest single base deletion mutations occur by a single base flipping in the template DNA, followed by temp ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genetic Insertion
In genetics, an insertion (also called an insertion mutation) is the addition of one or more nucleotide base pairs into a DNA sequence. This can often happen in microsatellite regions due to the DNA polymerase slipping. Insertions can be anywhere in size from one base pair incorrectly inserted into a DNA sequence to a section of one chromosome inserted into another. The mechanism of the smallest single base insertion mutations is believed to be through base-pair separation between the template and primer strands followed by non-neighbor base stacking, which can occur locally within the DNA polymerase active site. On a chromosome level, an ''insertion'' refers to the insertion of a larger sequence into a chromosome. This can happen due to unequal crossover during meiosis. N region addition is the addition of non-coded nucleotides during recombination by terminal deoxynucleotidyl transferase. P nucleotide insertion is the insertion of palindromic sequences encoded by the ends of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
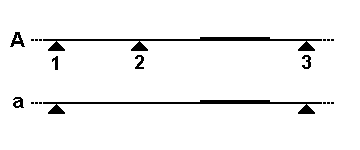
Variable Number Tandem Repeat
A variable number tandem repeat (or VNTR) is a location in a genome where a short nucleotide sequence is organized as a tandem repeat. These can be found on many chromosomes, and often show variations in length (number of repeats) among individuals. Each variant acts as an inherited allele, allowing them to be used for personal or parental identification. Their analysis is useful in genetics and biology research, forensics, and DNA fingerprinting. Structure and allelic variation In the schematic above, the rectangular blocks represent each of the repeated DNA sequences at a particular VNTR location. The repeats are in tandem – i.e. they are clustered together and oriented in the same direction. Individual repeats can be removed from (or added to) the VNTR via recombination or replication errors, leading to alleles with different numbers of repeats. Flanking regions are segments of repetitive sequence (shown here as thin lines), allowing the VNTR blocks to be extracte ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA or viral replication, mitosis, or meiosis or other types of damage to DNA (such as pyrimidine dimers caused by exposure to ultraviolet radiation), which then may undergo error-prone repair (especially microhomology-mediated end joining), cause an error during other forms of repair, or cause an error during replication ( translesion synthesis). Mutations may also result from substitution, insertion or deletion of segments of DNA due to mobile genetic elements. Mutations may or may not produce detectable changes in the observable characteristics ( phenotype) of an organism. Mutations play a part in both normal and abnormal biological processes including: evolution, cancer, and the development of the immune system, including junctional diversity. Mutati ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genome
A genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding genes, other functional regions of the genome such as regulatory sequences (see non-coding DNA), and often a substantial fraction of junk DNA with no evident function. Almost all eukaryotes have mitochondrial DNA, mitochondria and a small mitochondrial genome. Algae and plants also contain chloroplast DNA, chloroplasts with a chloroplast genome. The study of the genome is called genomics. The genomes of many organisms have been Whole-genome sequencing, sequenced and various regions have been annotated. The first genome to be sequenced was that of the virus φX174 in 1977; the first genome sequence of a prokaryote (''Haemophilus influenzae'') was published in 1995; the yeast (''Saccharomyces cerevisiae'') genome was the first eukaryotic genome to be sequenced in 1996. The Human Genome Project ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RFLP Genotyping
In molecular biology, restriction fragment length polymorphism (RFLP) is a technique that exploits variations in homologous DNA sequences, known as polymorphisms, populations, or species or to pinpoint the locations of genes within a sequence. The term may refer to a polymorphism itself, as detected through the differing locations of restriction enzyme sites, or to a related laboratory technique by which such differences can be illustrated. In RFLP analysis, a DNA sample is digested into fragments by one or more restriction enzymes, and the resulting ''restriction fragments'' are then separated by gel electrophoresis according to their size. RFLP analysis is now largely obsolete due to the emergence of inexpensive DNA sequencing technologies, but it was the first DNA profiling technique inexpensive enough to see widespread application. RFLP analysis was an important early tool in genome mapping, localization of genes for genetic disorders, determination of risk for disease, and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Coding DNA
The coding region of a gene, also known as the coding DNA sequence (CDS), is the portion of a gene's DNA or RNA that codes for a protein. Studying the length, composition, regulation, splicing, structures, and functions of coding regions compared to non-coding regions over different species and time periods can provide a significant amount of important information regarding gene organization and evolution of prokaryotes and eukaryotes. This can further assist in mapping the human genome and developing gene therapy. Definition Although this term is also sometimes used interchangeably with exon, it is not the exact same thing: the exon can be composed of the coding region as well as the 3' and 5' untranslated regions of the RNA, and so therefore, an exon would be partially made up of coding region. The 3' and 5' untranslated regions of the RNA, which do not code for protein, are termed non-coding regions and are not discussed on this page. There is often confusion between coding r ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Allele
An allele is a variant of the sequence of nucleotides at a particular location, or Locus (genetics), locus, on a DNA molecule. Alleles can differ at a single position through Single-nucleotide polymorphism, single nucleotide polymorphisms (SNP), but they can also have insertions and deletions of up to several thousand base pairs. Most alleles observed result in little or no change in the function or amount of the gene product(s) they code or regulate for. However, sometimes different alleles can result in different observable phenotypic traits, such as different pigmentation. A notable example of this is Gregor Mendel's discovery that the white and purple flower colors in pea plants were the result of a single gene with two alleles. Nearly all multicellular organisms have two sets of chromosomes at some point in their biological life cycle; that is, they are diploid. For a given locus, if the two chromosomes contain the same allele, they, and the organism, are homozygous with re ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |