|
Restriction Digest
In molecular biology, a restriction digest is a procedure used to prepare DNA for analysis or other processing. It is sometimes termed ''DNA fragmentation'', though this term is used for other procedures as well. In a restriction digest, DNA molecules are cleaved at specific regions of 4-12 nucleotides in length (restriction sites) by use of restriction enzymes which recognize these sequences. Hartl, Daniel L., Jones, Elizabeth W. (2001), ''Genetics: Analysis of Genes and Genomes'', Fifth Edition. The resulting digested DNA is very often selectively amplified using polymerase chain reaction (PCR), making it more suitable for analytical techniques such as agarose gel electrophoresis, and chromatography. It is used in genetic fingerprinting, plasmid subcloning, and RFLP analysis. Restriction site A given restriction enzyme cuts DNA segments within a specific nucleotide sequence, at what is called a restriction site. These ''recognition sequences'' are typically four, six, eig ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Molecular Biology
Molecular biology is a branch of biology that seeks to understand the molecule, molecular basis of biological activity in and between Cell (biology), cells, including biomolecule, biomolecular synthesis, modification, mechanisms, and interactions. Though cells and other microscopic structures had been observed in living organisms as early as the 18th century, a detailed understanding of the mechanisms and interactions governing their behavior did not emerge until the 20th century, when technologies used in physics and chemistry had advanced sufficiently to permit their application in the biological sciences. The term 'molecular biology' was first used in 1945 by the English physicist William Astbury, who described it as an approach focused on discerning the underpinnings of biological phenomena—i.e. uncovering the physical and chemical structures and properties of biological molecules, as well as their interactions with other molecules and how these interactions explain observ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Vector (molecular Biology)
In molecular cloning, a vector is any particle (e.g., plasmids, cosmids, Lambda phages) used as a vehicle to artificially carry a foreign nucleic acid sequence, nucleic sequence – usually DNA – into another Cell (biology), cell, where it can DNA replication, be replicated and/or Gene expression, expressed. A vector containing foreign DNA is termed recombinant DNA. The four major types of vectors are plasmids, viral vectors, cosmids, and Bacterial artificial chromosome, artificial chromosomes. Of these, the most commonly used vectors are plasmids. Common to all engineered vectors are an origin of replication, a multiple cloning site, multicloning site, and a selectable marker. The vector itself generally carries a DNA sequence that consists of an Insert (molecular biology), insert (in this case the transgene) and a larger sequence that serves as the "backbone" of the vector. The purpose of a vector which transfers genetic information to another cell is typically to isolate, mu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Polymerase Chain Reaction
The polymerase chain reaction (PCR) is a method widely used to make millions to billions of copies of a specific DNA sample rapidly, allowing scientists to amplify a very small sample of DNA (or a part of it) sufficiently to enable detailed study. PCR was invented in 1983 by American biochemist Kary Mullis at Cetus Corporation. Mullis and biochemist Michael Smith (chemist), Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993. PCR is fundamental to many of the procedures used in genetic testing and research, including analysis of Ancient DNA, ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory research for a broad variety of applications including biomedical research and forensic ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Genetic Fingerprinting
DNA profiling (also called DNA fingerprinting and genetic fingerprinting) is the process of determining an individual's deoxyribonucleic acid (DNA) characteristics. DNA analysis intended to identify a species, rather than an individual, is called DNA barcoding. DNA profiling is a forensic technique in criminal investigations, comparing criminal suspects' profiles to DNA evidence so as to assess the likelihood of their involvement in the crime. It is also used in paternity testing, to establish immigration eligibility, and in genealogical and medical research. DNA profiling has also been used in the study of animal and plant populations in the fields of zoology, botany, and agriculture. Background Starting in the mid 1970s, scientific advances allowed the use of DNA as a material for the identification of an individual. The first patent covering the direct use of DNA variation for forensicsUS5593832A was filed by Jeffrey Glassberg in 1983, based upon work he had done while ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
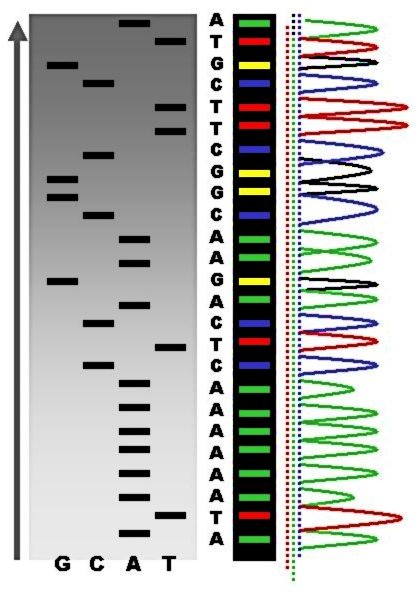
DNA Sequencing
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, thymine, cytosine, and guanine. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery. Knowledge of DNA sequences has become indispensable for basic biological research, Genographic Project, DNA Genographic Projects and in numerous applied fields such as medical diagnosis, biotechnology, forensic biology, virology and biological systematics. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers, characterize antibody repertoire, and can be used to guide patient treatment. Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged. The rapid advancements in DNA seque ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Agarose Gel Electrophoresis
Agarose gel electrophoresis is a method of gel electrophoresis used in biochemistry, molecular biology, genetics, and clinical chemistry to separate a mixed population of macromolecules such as DNA or proteins in a matrix of agarose, one of the two main components of agar. The proteins may be separated by charge and/or size (isoelectric focusing agarose electrophoresis is essentially size independent), and the DNA and RNA fragments by length. Biomolecules are separated by applying an electric field to move the charged molecules through an agarose matrix, and the biomolecules are separated by size in the agarose gel matrix. Agarose gel is easy to cast, has relatively fewer charged groups, and is particularly suitable for separating DNA of size range most often encountered in laboratories, which accounts for the popularity of its use. The separated DNA may be viewed with stain, most commonly under UV light, and the DNA fragments can be extracted from the gel with relative ease. Most ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Short Tandem Repeat Polymorphism
A microsatellite is a tract of repetitive DNA in which certain DNA motifs (ranging in length from one to six or more base pairs) are repeated, typically 5–50 times. Microsatellites occur at thousands of locations within an organism's genome. They have a higher mutation rate than other areas of DNA leading to high genetic diversity. Microsatellites are often referred to as short tandem repeats (STRs) by forensic geneticists and in genetic genealogy, or as simple sequence repeats (SSRs) by plant geneticists. Microsatellites and their longer cousins, the minisatellites, together are classified as VNTR (variable number of tandem repeats) DNA. The name "satellite" DNA refers to the early observation that centrifugation of genomic DNA in a test tube separates a prominent layer of bulk DNA from accompanying "satellite" layers of repetitive DNA. They are widely used for DNA profiling in cancer diagnosis, in kinship analysis (especially paternity testing) and in forensic identific ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Amplified Fragment Length Polymorphism
Amplified fragment length polymorphism (AFLP-PCR or AFLP) is a PCR-based tool used in genetics research, DNA fingerprinting, and in the practice of genetic engineering. Developed in the early 1990s by Pieter Vos, AFLP uses restriction enzymes to digest genomic DNA, followed by ligation of adaptors to the sticky ends of the restriction fragments. A subset of the restriction fragments is then selected to be amplified. This selection is achieved by using primers complementary to the adaptor sequence, the restriction site sequence and a few nucleotides inside the restriction site fragments (as described in detail below). The amplified fragments are separated and visualized on denaturing on agarose gel electrophoresis, either through autoradiography or fluorescence methodologies, or via automated capillary sequencing instruments. Although AFLP should not be used as an acronym, it is commonly referred to as "Amplified fragment length polymorphism". However, the resulting data are ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Restriction Fragment Length Polymorphism
In molecular biology, restriction fragment length polymorphism (RFLP) is a technique that exploits variations in homologous DNA sequences, known as polymorphisms, populations, or species or to pinpoint the locations of genes within a sequence. The term may refer to a polymorphism itself, as detected through the differing locations of restriction enzyme sites, or to a related laboratory technique by which such differences can be illustrated. In RFLP analysis, a DNA sample is digested into fragments by one or more restriction enzymes, and the resulting ''restriction fragments'' are then separated by gel electrophoresis according to their size. RFLP analysis is now largely obsolete due to the emergence of inexpensive DNA sequencing technologies, but it was the first DNA profiling technique inexpensive enough to see widespread application. RFLP analysis was an important early tool in genome mapping, localization of genes for genetic disorders, determination of risk for disease, an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Ligation (molecular Biology)
Ligation is the joining of two nucleotides, or two nucleic acid fragments, into a single polymeric chain through the action of an enzyme known as a ligase. The reaction involves the formation of a phosphodiester bond between the 3'-hydroxyl terminus of one nucleotide and the 5'-phosphoryl terminus of another nucleotide, which results in the two nucleotides being linked consecutively on a single strand. Ligation works in fundamentally the same way for both DNA and RNA. A cofactor is generally involved in the reaction, usually Adenosine triphosphate, ATP or Nicotinamide riboside, NAD+. Eukaryotic ligases belong to the ATP type, while the NAD+ type are found in bacteria (e.g. Escherichia coli, ''E. coli''). Ligation occurs naturally as part of numerous cellular processes, including DNA replication, transcription, splicing, and recombination, and is also an essential laboratory procedure in molecular cloning, whereby DNA fragments are joined to create recombinant DNA molecules (such a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Expression Vector
An expression vector, otherwise known as an expression construct, is usually a plasmid or virus designed for gene expression in cells. The vector (molecular biology), vector is used to introduce a specific gene into a target cell, and can commandeer the cell's mechanism for protein synthesis to produce the protein Genetic code, encoded by the gene. Expression vectors are the basic tools in biotechnology for the protein production, production of proteins. The Vector (molecular biology), vector is engineered to contain regulatory sequences that act as Enhancer (genetics), enhancer and Promoter (biology), promoter regions and lead to efficient transcription of the gene carried on the expression vector. The goal of a well-designed expression vector is the efficient production of protein, and this may be achieved by the production of significant amount of stable messenger RNA, which can then be Translation (biology), translated into protein. The expression of a protein may be tightly ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Cloning Vector
A cloning vector is a small piece of DNA that can be stably maintained in an organism, and into which a foreign DNA fragment can be inserted for cloning purposes. The cloning vector may be DNA taken from a virus, the cell of a higher organism, or it may be the plasmid of a bacterium. The vector contains features that allow for the convenient insertion of a DNA fragment into the vector or its removal from the vector, for example through the presence of restriction sites. The vector and the foreign DNA may be treated with a restriction enzyme that cuts the DNA, and DNA fragments thus generated contain either blunt ends or overhangs known as sticky ends, and vector DNA and foreign DNA with compatible ends can then be joined by molecular ligation. After a DNA fragment has been cloned into a cloning vector, it may be further subcloned into another vector designed for more specific use. There are many types of cloning vectors, but the most commonly used ones are genetically engineer ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |