|
MiR-155
MiR-155 is a microRNA that in humans is encoded by the ''MIR155'' host gene or ''MIR155HG''. MiR-155 plays a role in various physiological and pathological processes. Exogenous molecular control ''in vivo'' of miR-155 expression may inhibit malignant growth, viral infections, and enhance the progression of cardiovascular diseases. Discovery The ''MIR155HG'' was initially identified as a gene that was transcriptionally activated by promoter insertion at a common retroviral integration site in B-cell lymphomas and was formerly called BIC (B-cell Integration Cluster). The ''MIR155HG'' is transcribed by RNA polymerase II and the resulting ~1,500 nucleotide RNA is capped and polyadenylated. The 23 nucleotide single-stranded miR-155, which is harbored in exon 3, is subsequently processed from the parent RNA molecule. Biogenesis The MIR155HG RNA transcript does not contain a long open reading frame (ORF), however, it does include an imperfectly base-paired stem loop that ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mir155 Gene
MiR-155 is a microRNA that in humans is encoded by the ''MIR155'' host gene or ''MIR155HG''. MiR-155 plays a role in various physiological and pathological processes. Exogenous molecular control ''in vivo'' of miR-155 expression may inhibit malignant growth, viral infections, and enhance the progression of cardiovascular diseases. Discovery The ''MIR155HG'' was initially identified as a gene that was transcriptionally activated by promoter insertion at a common retroviral integration site in B-cell lymphomas and was formerly called BIC (B-cell Integration Cluster). The ''MIR155HG'' is transcribed by RNA polymerase II and the resulting ~1,500 nucleotide RNA is capped and polyadenylated. The 23 nucleotide single-stranded miR-155, which is harbored in exon 3, is subsequently processed from the parent RNA molecule. Biogenesis The MIR155HG RNA transcript does not contain a long open reading frame (ORF), however, it does include an imperfectly base-paired stem loop that is ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MicroRNA
MicroRNA (miRNA) are small, single-stranded, non-coding RNA molecules containing 21 to 23 nucleotides. Found in plants, animals and some viruses, miRNAs are involved in RNA silencing and post-transcriptional regulation of gene expression. miRNAs base-pair to complementary sequences in mRNA molecules, then gene silence said mRNA molecules by one or more of the following processes: (1) cleavage of mRNA strand into two pieces, (2) destabilization of mRNA by shortening its poly(A) tail, or (3) translation of mRNA into proteins. This last method of gene silencing is the least efficient of the three, and requires the aid of ribosomes. miRNAs resemble the small interfering RNAs (siRNAs) of the RNA interference (RNAi) pathway, except miRNAs derive from regions of RNA transcripts that fold back on themselves to form short hairpins, whereas siRNAs derive from longer regions of double-stranded RNA. The human genome may encode over 1900 miRNAs, although more recent analysis sugges ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
NcRNA
A non-coding RNA (ncRNA) is a functional RNA molecule that is not translated into a protein. The DNA sequence from which a functional non-coding RNA is transcribed is often called an RNA gene. Abundant and functionally important types of non-coding RNAs include transfer RNAs (tRNAs) and ribosomal RNAs (rRNAs), as well as small RNAs such as microRNAs, siRNAs, piRNAs, snoRNAs, snRNAs, exRNAs, scaRNAs and the long ncRNAs such as Xist and HOTAIR. The number of non-coding RNAs within the human genome is unknown; however, recent transcriptomic and bioinformatic studies suggest that there are thousands of non-coding transcripts. Many of the newly identified ncRNAs have not been validated for their function. There is no consensus in the literature on how much of non-coding transcription is functional. Some researchers have argued that many ncRNAs are non-functional (sometimes referred to as "junk RNA"), spurious transcriptions. Others, however, disagree, arguing instead that ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Complementarity (molecular Biology)
In molecular biology, complementarity describes a relationship between two structures each following the lock-and-key principle. In nature complementarity is the base principle of DNA replication and transcription as it is a property shared between two DNA or RNA sequences, such that when they are aligned antiparallel to each other, the nucleotide bases at each position in the sequences will be complementary, much like looking in the mirror and seeing the reverse of things. This complementary base pairing allows cells to copy information from one generation to another and even find and repair damage to the information stored in the sequences. The degree of complementarity between two nucleic acid strands may vary, from complete complementarity (each nucleotide is across from its opposite) to no complementarity (each nucleotide is not across from its opposite) and determines the stability of the sequences to be together. Furthermore, various DNA repair functions as well as ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein. mRNA is created during the process of transcription, where an enzyme ( RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and, utilising amino acids carried by transfer RNA (tRNA), the ribosome creates the protein. This process is known as translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genetic information in a biological system. As in DNA, ge ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mature Mir-155-5p And -3p
Mature is the adjectival form of maturity, as immature is the adjectival form of immaturity, which have several meanings. Mature or immature may also refer to: * Mature, a character from ''The King of Fighters'' series *"Mature 17+", a rating in the Entertainment Software Rating Board video game rating system *Victor Mature Victor John Mature (January 29, 1913 – August 4, 1999) was an American stage, film, and television actor who was a leading man in Hollywood during the 1940s and 1950s. His best known film roles include ''One Million B.C.'' (1940), '' My Darlin ... (1913-1999), American actor * Immature (band), an American boy band See also * Adult (other) * Maturation (other) * Maturity (other) * Ripeness {{disambig, surname ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SiRNA
Small interfering RNA (siRNA), sometimes known as short interfering RNA or silencing RNA, is a class of double-stranded RNA at first non-coding RNA molecules, typically 20-24 (normally 21) base pairs in length, similar to miRNA, and operating within the RNA interference (RNAi) pathway. It interferes with the expression of specific genes with complementary nucleotide sequences by degrading mRNA after transcription, preventing translation. Text was copied from this source, which is available under Creative Commons Attribution 4.0 International License Structure Naturally occurring siRNAs have a well-defined structure that is a short (usually 20 to 24- bp) double-stranded RNA (dsRNA) with phosphorylated 5' ends and hydroxylated 3' ends with two overhanging nucleotides. The Dicer enzyme catalyzes production of siRNAs from long dsRNAs and small hairpin RNAs. siRNAs can also be introduced into cells by transfection. Since in principle any gene can be knocked down by a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA-induced Silencing Complex
The RNA-induced silencing complex, or RISC, is a multiprotein complex, specifically a ribonucleoprotein, which functions in gene silencing via a variety of pathways at the transcriptional and translational levels. Using single-stranded RNA (ssRNA) fragments, such as microRNA (miRNA), or double-stranded small interfering RNA (siRNA), the complex functions as a key tool in gene regulation. The single strand of RNA acts as a template for RISC to recognize complementary messenger RNA (mRNA) transcript. Once found, one of the proteins in RISC, Argonaute, activates and cleaves the mRNA. This process is called RNA interference (RNAi) and it is found in many eukaryotes; it is a key process in defense against viral infections, as it is triggered by the presence of double-stranded RNA (dsRNA). Discovery The biochemical identification of RISC was conducted by Gregory Hannon and his colleagues at the Cold Spring Harbor Laboratory. This was only a couple of years after the discovery of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Argonaute
The Argonaute protein family, first discovered for its evolutionarily conserved stem cell function, plays a central role in RNA silencing processes as essential components of the RNA-induced silencing complex (RISC). RISC is responsible for the gene silencing phenomenon known as RNA interference (RNAi). Argonaute proteins bind different classes of small non-coding RNAs, including microRNAs (miRNAs), small interfering RNAs (siRNAs) and Piwi-interacting RNAs (piRNAs). Small RNAs guide Argonaute proteins to their specific targets through sequence complementarity (base pairing), which then leads to mRNA cleavage, translation inhibition, and/or the initiation of mRNA decay. The name of this protein family is derived from a mutant phenotype resulting from mutation of AGO1 in ''Arabidopsis thaliana'', which was likened by Bohmert et al. to the appearance of the pelagic octopus ''Argonauta argo''. RNA interference RNA interference (RNAi) is a biological process in which the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Dicer
Dicer, also known as endoribonuclease Dicer or helicase with RNase motif, is an enzyme that in humans is encoded by the gene. Being part of the RNase III family, Dicer cleaves double-stranded RNA (dsRNA) and pre-microRNA (pre-miRNA) into short double-stranded RNA fragments called small interfering RNA and microRNA, respectively. These fragments are approximately 20–25 base pairs long with a two-base overhang on the 3′-end. Dicer facilitates the activation of the RNA-induced silencing complex (RISC), which is essential for RNA interference. RISC has a catalytic component Argonaute, which is an endonuclease capable of degrading messenger RNA (mRNA). Discovery Dicer was given its name in 2001 by Stony Brook PhD student Emily Bernstein while conducting research in Gregory Hannon's lab at Cold Spring Harbor Laboratory. Bernstein sought to discover the enzyme responsible for generating small RNA fragments from double-stranded RNA. Dicer's ability to generate around 22 n ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |

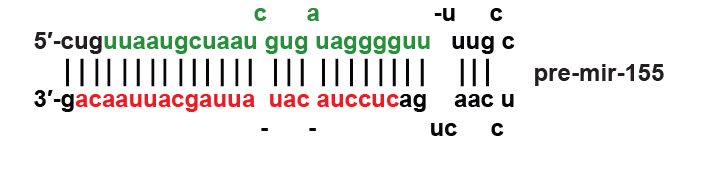

.jpg)

