|
Glutamate-1-semialdehyde 2,1-aminomutase
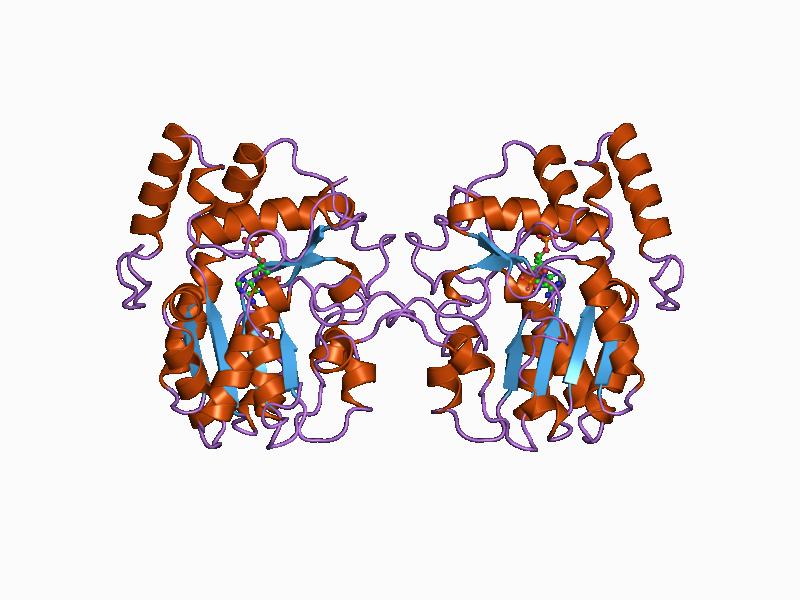
In enzymology, a glutamate-1-semialdehyde 2,1-aminomutase () is an enzyme that catalyzes the chemical reaction :L-glutamate 1-semialdehyde \rightleftharpoons 5-aminolevulinate Hence, this enzyme has one substrate, L-glutamate-1-semialdehyde, and one product, 5-aminolevulinate. This enzyme belongs to the family of isomerases, specifically those intramolecular transferases transferring amino groups. The systematic name of this enzyme class is (S)-4-amino-5-oxopentanoate 4,5-aminomutase. This enzyme is also called glutamate-1-semialdehyde aminotransferase. This enzyme participates in porphyrin and chlorophyll biosynthesis. It employs one cofactor, pyridoxal phosphate. Structural studies As of late 2007, 10 structures A structure is an arrangement and organization of interrelated elements in a material object or system, or the object or system so organized. Material structures include man-made objects such as buildings and machines and natural objects such as ... have ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Enzymology
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products. Almost all metabolic processes in the cell need enzyme catalysis in order to occur at rates fast enough to sustain life. Metabolic pathways depend upon enzymes to catalyze individual steps. The study of enzymes is called ''enzymology'' and the field of pseudoenzyme analysis recognizes that during evolution, some enzymes have lost the ability to carry out biological catalysis, which is often reflected in their amino acid sequences and unusual 'pseudocatalytic' properties. Enzymes are known to catalyze more than 5,000 biochemical reaction types. Other biocatalysts are catalytic RNA molecules, called ribozymes. Enzymes' specificity comes from their unique three-dimensional structures. Like all catalysts, enzymes increase the reaction ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
List Of Enzymes
This article lists enzymes by their classification in the International Union of Biochemistry and Molecular Biology's Enzyme Commission (EC) numbering system. * List of EC numbers (EC 5) * List of EC numbers (EC 6) :Oxidoreductases (EC 1) (Oxidoreductase) * Dehydrogenase * Luciferase * DMSO reductase :EC 1.1 (act on the CH-OH group of donors) * :EC 1.1.1 (with NAD+ or NADP+ as acceptor) ** Alcohol dehydrogenase (NAD) ** Alcohol dehydrogenase (NADP) ** Homoserine dehydrogenase ** Aminopropanol oxidoreductase ** Diacetyl reductase **Glycerol dehydrogenase ** Propanediol-phosphate dehydrogenase ** glycerol-3-phosphate dehydrogenase (NAD+) ** D-xylulose reductase **L-xylulose reductase ** Lactate dehydrogenase ** Malate dehydrogenase ** Isocitrate dehydrogenase **HMG-CoA reductase * :EC 1.1.2 (with a cytochrome as acceptor) * :EC 1.1.3 (with oxygen as acceptor) ** Glucose oxidase ** L-gulonolactone oxidase ** Thiamine oxidase ** Xanthine oxidase * :EC 1.1.4 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Data Bank
The Protein Data Bank (PDB) is a database for the three-dimensional structural data of large biological molecules, such as proteins and nucleic acids. The data, typically obtained by X-ray crystallography, NMR spectroscopy, or, increasingly, cryo-electron microscopy, and submitted by biologists and biochemists from around the world, are freely accessible on the Internet via the websites of its member organisations (PDBe, PDBj, RCSB, and BMRB). The PDB is overseen by an organization called the Worldwide Protein Data Bank, wwPDB. The PDB is a key in areas of structural biology, such as structural genomics. Most major scientific journals and some funding agencies now require scientists to submit their structure data to the PDB. Many other databases use protein structures deposited in the PDB. For example, SCOP and CATH classify protein structures, while PDBsum provides a graphic overview of PDB entries using information from other sources, such as Gene ontology. History Two ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tertiary Structure
Protein tertiary structure is the three dimensional shape of a protein. The tertiary structure will have a single polypeptide chain "backbone" with one or more protein secondary structures, the protein domains. Amino acid side chains may interact and bond in a number of ways. The interactions and bonds of side chains within a particular protein determine its tertiary structure. The protein tertiary structure is defined by its atomic coordinates. These coordinates may refer either to a protein domain or to the entire tertiary structure.Branden C. and Tooze J. "Introduction to Protein Structure" Garland Publishing, New York. 1990 and 1991. A number of tertiary structures may fold into a quaternary structure.Kyte, J. "Structure in Protein Chemistry." Garland Publishing, New York. 1995. History The science of the tertiary structure of proteins has progressed from one of hypothesis to one of detailed definition. Although Emil Fischer had suggested proteins were made of poly ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Pyridoxal Phosphate
Pyridoxal phosphate (PLP, pyridoxal 5'-phosphate, P5P), the active form of vitamin B6, is a coenzyme in a variety of enzymatic reactions. The International Union of Biochemistry and Molecular Biology has catalogued more than 140 PLP-dependent activities, corresponding to ~4% of all classified activities. The versatility of PLP arises from its ability to covalently bind the substrate, and then to act as an electrophilic catalyst, thereby stabilizing different types of carbanionic reaction intermediates. Role as a coenzyme PLP acts as a coenzyme in all transamination reactions, and in certain decarboxylation, deamination, and racemization reactions of amino acids. The aldehyde group of PLP forms a Schiff-base linkage (internal aldimine) with the ε-amino group of a specific lysine group of the aminotransferase enzyme. The α-amino group of the amino acid substrate displaces the ε-amino group of the active-site lysine residue in a process known as transaldimination. The resulti ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cofactor (biochemistry)
A cofactor is a non-protein chemical compound or metallic ion that is required for an enzyme's role as a catalyst (a catalyst is a substance that increases the rate of a chemical reaction). Cofactors can be considered "helper molecules" that assist in biochemical transformations. The rates at which these happen are characterized in an area of study called enzyme kinetics. Cofactors typically differ from ligands in that they often derive their function by remaining bound. Cofactors can be divided into two types: inorganic ions and complex organic molecules called coenzymes. Coenzymes are mostly derived from vitamins and other organic essential nutrients in small amounts. (Note that some scientists limit the use of the term "cofactor" for inorganic substances; both types are included here.) Coenzymes are further divided into two types. The first is called a "prosthetic group", which consists of a coenzyme that is tightly (or even covalently) and permanently bound to a protein ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chlorophyll
Chlorophyll (also chlorophyl) is any of several related green pigments found in cyanobacteria and in the chloroplasts of algae and plants. Its name is derived from the Greek words , ("pale green") and , ("leaf"). Chlorophyll allow plants to absorb energy from light. Chlorophylls absorb light most strongly in the blue portion of the electromagnetic spectrum as well as the red portion. Conversely, it is a poor absorber of green and near-green portions of the spectrum. Hence chlorophyll-containing tissues appear green because green light, diffusively reflected by structures like cell walls, is less absorbed. Two types of chlorophyll exist in the photosystems of green plants: chlorophyll ''a'' and ''b''. History Chlorophyll was first isolated and named by Joseph Bienaimé Caventou and Pierre Joseph Pelletier in 1817. The presence of magnesium in chlorophyll was discovered in 1906, and was that element's first detection in living tissue. After initial work done by German ch ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Porphyrin
Porphyrins ( ) are a group of heterocyclic macrocycle organic compounds, composed of four modified pyrrole subunits interconnected at their α carbon atoms via methine bridges (=CH−). The parent of porphyrin is porphine, a rare chemical compound of exclusively theoretical interest. Substituted porphines are called porphyrins. With a total of 26 π-electrons, of which 18 π-electrons form a planar, continuous cycle, the porphyrin ring structure is often described as aromatic. One result of the large conjugated system is that porphyrins typically absorb strongly in the visible region of the electromagnetic spectrum, i.e. they are deeply colored. The name "porphyrin" derives from the Greek word πορφύρα (''porphyra''), meaning ''purple''. Complexes of porphyrins Concomitant with the displacement of two N-''H'' protons, porphyrins bind metal ions in the N4 "pocket". The metal ion usually has a charge of 2+ or 3+. A schematic equation for these syntheses is shown: :H2porp ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transferase
A transferase is any one of a class of enzymes that catalyse the transfer of specific functional groups (e.g. a methyl or glycosyl group) from one molecule (called the donor) to another (called the acceptor). They are involved in hundreds of different biochemical pathways throughout biology, and are integral to some of life's most important processes. Transferases are involved in myriad reactions in the cell. Three examples of these reactions are the activity of coenzyme A (CoA) transferase, which transfers thiol esters, the action of N-acetyltransferase, which is part of the pathway that metabolizes tryptophan, and the regulation of pyruvate dehydrogenase (PDH), which converts pyruvate to acetyl CoA. Transferases are also utilized during translation. In this case, an amino acid chain is the functional group transferred by a peptidyl transferase. The transfer involves the removal of the growing amino acid chain from the tRNA molecule in the A-site of the ribosome an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products. Almost all metabolic processes in the cell need enzyme catalysis in order to occur at rates fast enough to sustain life. Metabolic pathways depend upon enzymes to catalyze individual steps. The study of enzymes is called ''enzymology'' and the field of pseudoenzyme analysis recognizes that during evolution, some enzymes have lost the ability to carry out biological catalysis, which is often reflected in their amino acid sequences and unusual 'pseudocatalytic' properties. Enzymes are known to catalyze more than 5,000 biochemical reaction types. Other biocatalysts are catalytic RNA molecules, called ribozymes. Enzymes' specificity comes from their unique three-dimensional structures. Like all catalysts, enzymes increase the react ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Isomerase
Isomerases are a general class of enzymes that convert a molecule from one isomer to another. Isomerases facilitate intramolecular rearrangements in which bonds are broken and formed. The general form of such a reaction is as follows: A–B → B–A There is only one substrate yielding one product. This product has the same molecular formula as the substrate but differs in bond connectivity or spatial arrangement. Isomerases catalyze reactions across many biological processes, such as in glycolysis and carbohydrate metabolism. Isomerization Isomerases catalyze changes within one molecule. They convert one isomer to another, meaning that the end product has the same molecular formula but a different physical structure. Isomers themselves exist in many varieties but can generally be classified as structural isomers or stereoisomers. Structural isomers have a different ordering of bonds and/or different bond connectivity from one another, as in the case of hexane and its ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |