|
Chromosome 2 (human)
Chromosome 2 is one of the twenty-three pairs of chromosomes in humans. People normally have two copies of this chromosome. Chromosome 2 is the second-largest human chromosome, spanning more than 242 million base pairs and representing almost eight percent of the total DNA in human cells. Chromosome 2 contains the HOXD homeobox gene cluster. Chromosomes Humans have only twenty-three pairs of chromosomes, while all other extant members of Hominidae have twenty-four pairs. It is believed that Neanderthals and Denisovans had twenty-three pairs. Human chromosome 2 is a result of an end-to-end fusion of two ancestral chromosomes.It has been hypothesized that Human Chromosome 2 is a fusion of two ancestral chromosomes by Alec MacAndrew; accessed 18 May 2006. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
G Banding
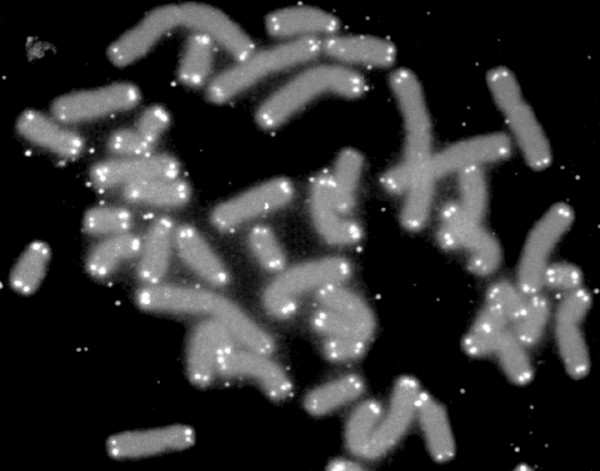
G-banding, G banding or Giemsa banding is a technique used in cytogenetics to produce a visible karyotype by staining condensed chromosomes. It is the most common chromosome banding method. It is useful for identifying genetic diseases through the photographic representation of the entire chromosome complement.Speicher, Michael R. and Nigel P. Carter. "The New Cytogenetics: Blurring the Boundaries with Molecular Biology." ''Nature'' Reviews Genetics, Vol 6. Oct 2005. The metaphase chromosomes are treated with trypsin (to partially digest the chromosome) and stained with Giemsa stain. Heterochromatic regions, which tend to be rich with adenine and thymine (AT-rich) DNA and relatively gene-poor, stain more darkly in G-banding. In contrast, less condensed chromatin ( Euchromatin)—which tends to be rich with guanine and cytosine ( GC-rich) and more transcriptionally active—incorporates less Giemsa stain, and these regions appear as light bands in G-banding. The pattern of ban ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gorilla
Gorillas are herbivorous, predominantly ground-dwelling great apes that inhabit the tropical forests of equatorial Africa. The genus ''Gorilla'' is divided into two species: the eastern gorilla and the western gorilla, and either four or five subspecies. The DNA of gorillas is highly similar to Human evolutionary genetics, that of humans, from 95 to 99% depending on what is included, and they are the next closest living relatives to humans after chimpanzees and bonobos. Gorillas are the largest Neontology#Extant taxa versus extinct taxa, living primates, reaching heights between 1.25 and 1.8 metres, weights between 100 and 270 kg, and arm spans up to 2.6 metres, depending on species and sex. They tend to live in troops, with the leader being called a silverback. The Eastern gorilla is distinguished from the Western by darker fur colour and some other minor morphological differences. Gorillas tend to live 35–40 years in the wild. The Oldest hominids, oldest gorilla kno ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
UniProt
UniProt is a freely accessible database of protein sequence and functional information, many entries being derived from genome sequencing projects. It contains a large amount of information about the biological function of proteins derived from the research literature. It is maintained by the UniProt consortium, which consists of several European bioinformatics organisations and a foundation from Washington, DC, United States. The UniProt consortium The UniProt consortium comprises the European Bioinformatics Institute (EBI), the Swiss Institute of Bioinformatics (SIB), and the Protein Information Resource (PIR). EBI, located at the Wellcome Trust Genome Campus in Hinxton, UK, hosts a large resource of bioinformatics databases and services. SIB, located in Geneva, Switzerland, maintains the ExPASy (Expert Protein Analysis System) servers that are a central resource for proteomics tools and databases. PIR, hosted by the National Biomedical Research Foundation (NBRF) at the Geo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ensembl Genome Database Project
Ensembl genome database project is a scientific project at the European Bioinformatics Institute, which provides a centralized resource for geneticists, molecular biologists and other researchers studying the genomes of our own species and other vertebrates and model organisms. Ensembl is one of several well known genome browsers for the retrieval of genomic information. Similar databases and browsers are found at NCBI and the University of California, Santa Cruz (UCSC). History The human genome consists of three billion base pairs, which code for approximately 20,000–25,000 genes. However the genome alone is of little use, unless the locations and relationships of individual genes can be identified. One option is manual annotation, whereby a team of scientists tries to locate genes using experimental data from scientific journals and public databases. However this is a slow, painstaking task. The alternative, known as automated annotation, is to use the power of computer ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
HUGO Gene Nomenclature Committee
The HUGO Gene Nomenclature Committee (HGNC) is a committee of the Human Genome Organisation (HUGO) that sets the standards for human gene nomenclature. The HGNC approves a ''unique'' and ''meaningful'' name for every known human gene, based on a query of experts. In addition to the name, which is usually 1 to 10 words long, the HGNC also assigns a symbol (a short group of characters) to every gene. As with an SI symbol, a gene symbol is like an abbreviation but is more than that, being a second unique name that can stand on its own just as much as substitute for the longer name. It may not necessarily "stand for" the initials of the name, although many gene symbols do reflect that origin. Purpose Especially gene abbreviations/symbols but also full gene names are often not specific for a single gene. A marked example is CAP which can refer to any of 6 different genes (BRD4'', CAP1'', HACD1'', LNPEP'', SERPINB6'', and SORBS1''). The HGNC short gene names, or gene symbols, unlike ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Pseudogene
Pseudogenes are nonfunctional segments of DNA that resemble functional genes. Most arise as superfluous copies of functional genes, either directly by DNA duplication or indirectly by reverse transcription of an mRNA transcript. Pseudogenes are usually identified when genome sequence analysis finds gene-like sequences that lack regulatory sequences needed for transcription or translation, or whose coding sequences are obviously defective due to frameshifts or premature stop codons. Most non-bacterial genomes contain many pseudogenes, often as many as functional genes. This is not surprising, since various biological processes are expected to accidentally create pseudogenes, and there are no specialized mechanisms to remove them from genomes. Eventually pseudogenes may be deleted from their genomes by chance DNA replication or DNA repair errors, or they may accumulate so many mutational changes that they are no longer recognizable as former genes. Analysis of these degenerati ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Non-coding RNA
A non-coding RNA (ncRNA) is a functional RNA molecule that is not Translation (genetics), translated into a protein. The DNA sequence from which a functional non-coding RNA is transcribed is often called an RNA gene. Abundant and functionally important list of RNAs, types of non-coding RNAs include transfer RNAs (tRNAs) and ribosomal RNAs (rRNAs), as well as small RNAs such as microRNAs, siRNAs, piRNAs, snoRNAs, snRNAs, Extracellular RNA, exRNAs, scaRNAs and the long noncoding RNA, long ncRNAs such as Xist and HOTAIR. The number of non-coding RNAs within the human genome is unknown; however, recent Transcriptomics, transcriptomic and Bioinformatics, bioinformatic studies suggest that there are thousands of non-coding transcripts. Many of the newly identified ncRNAs have not been validated for their function. There is no consensus in the literature on how much of non-coding transcription is functional. Some researchers have argued that many ncRNAs are non-functional (sometimes r ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein-coding Genes
The human genome is a complete set of nucleic acid sequences for humans, encoded as DNA within the 23 chromosome pairs in cell nuclei and in a small DNA molecule found within individual mitochondria. These are usually treated separately as the nuclear genome and the mitochondrial genome. Human genomes include both protein-coding DNA sequences and various types of DNA that does not encode proteins. The latter is a diverse category that includes DNA coding for non-translated RNA, such as that for ribosomal RNA, transfer RNA, ribozymes, small nuclear RNAs, and several types of regulatory RNAs. It also includes promoters and their associated gene-regulatory elements, DNA playing structural and replicatory roles, such as scaffolding regions, telomeres, centromeres, and origins of replication, plus large numbers of transposable elements, inserted viral DNA, non-functional pseudogenes and simple, highly-repetitive sequences. Introns make up a large percentage of non-coding DNA. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Number Of Genes
In biology, the word gene (from , ; "...Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a basic unit of heredity and the molecular gene is a sequence of nucleotides in DNA that is transcribed to produce a functional RNA. There are two types of molecular genes: protein-coding genes and noncoding genes. During gene expression, the DNA is first copied into RNA. The RNA can be directly functional or be the intermediate template for a protein that performs a function. The transmission of genes to an organism's offspring is the basis of the inheritance of phenotypic traits. These genes make up different DNA sequences called genotypes. Genotypes along with environmental and developmental factors determine what the phenotypes will be. Most biological traits are under the influence of polygenes (many different genes) as well as gene– ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Prediction
In computational biology, gene prediction or gene finding refers to the process of identifying the regions of genomic DNA that encode genes. This includes protein-coding genes as well as RNA genes, but may also include prediction of other functional elements such as regulatory regions. Gene finding is one of the first and most important steps in understanding the genome of a species once it has been Sequencing, sequenced. In its earliest days, "gene finding" was based on painstaking experimentation on living cells and organisms. Statistical analysis of the rates of homologous recombination of several different genes could determine their order on a certain chromosome, and information from many such experiments could be combined to create a genetic map specifying the rough location of known genes relative to each other. Today, with comprehensive genome sequence and powerful computational resources at the disposal of the research community, gene finding has been redefined as a largely ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genome Annotation
DNA annotation or genome annotation is the process of identifying the locations of genes and all of the coding regions in a genome and determining what those genes do. An annotation (irrespective of the context) is a note added by way of explanation or commentary. Once a genome is sequenced, it needs to be annotated to make sense of it. Genes in a eukaryotic genome can be annotated using various annotation tools such as FINDER. A modern annotation pipeline can support a user-friendly web interface and software containerization such as MOSGA. For DNA annotation, a previously unknown sequence representation of genetic material is enriched with information relating genomic position to intron-exon boundaries, regulatory sequences, repeats, gene names and protein products. This annotation is stored in genomic databases such as Mouse Genome Informatics, FlyBase, and WormBase. Educational materials on some aspects of biological annotation from the 2006 Gene Ontology annotation camp and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Telomere
A telomere (; ) is a region of repetitive nucleotide sequences associated with specialized proteins at the ends of linear chromosomes. Although there are different architectures, telomeres, in a broad sense, are a widespread genetic feature most commonly found in eukaryotes. In most, if not all species possessing them, they protect the terminal regions of chromosomal DNA from progressive degradation and ensure the integrity of linear chromosomes by preventing DNA repair systems from mistaking the very ends of the DNA strand for a double-strand break. Discovery In the early 1970s, Soviet theorist Alexei Olovnikov first recognized that chromosomes could not completely replicate their ends; this is known as the "end replication problem". Building on this, and accommodating Leonard Hayflick's idea of limited somatic cell division, Olovnikov suggested that DNA sequences are lost every time a cell replicates until the loss reaches a critical level, at which point cell division e ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |