|
Sequence Alignment
In bioinformatics, a sequence alignment is a way of arranging the sequences of DNA, RNA, or protein to identify regions of similarity that may be a consequence of functional, structural, or evolutionary relationships between the sequences. Aligned sequences of nucleotide or amino acid residues are typically represented as rows within a matrix. Gaps are inserted between the residues so that identical or similar characters are aligned in successive columns. Sequence alignments are also used for non-biological sequences, such as calculating the distance cost between strings in a natural language or in financial data. Interpretation If two sequences in an alignment share a common ancestor, mismatches can be interpreted as point mutations and gaps as indels (that is, insertion or deletion mutations) introduced in one or both lineages in the time since they diverged from one another. In sequence alignments of proteins, the degree of similarity between amino acids occupying a p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bioinformatics
Bioinformatics () is an interdisciplinary field that develops methods and software tools for understanding biological data, in particular when the data sets are large and complex. As an interdisciplinary field of science, bioinformatics combines biology, chemistry, physics, computer science, information engineering, mathematics and statistics to analyze and interpret the biological data. Bioinformatics has been used for '' in silico'' analyses of biological queries using computational and statistical techniques. Bioinformatics includes biological studies that use computer programming as part of their methodology, as well as specific analysis "pipelines" that are repeatedly used, particularly in the field of genomics. Common uses of bioinformatics include the identification of candidates genes and single nucleotide polymorphisms ( SNPs). Often, such identification is made with the aim to better understand the genetic basis of disease, unique adaptations, desirable properties ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Heuristic Algorithm
In mathematical optimization and computer science, heuristic (from Greek εὑρίσκω "I find, discover") is a technique designed for solving a problem more quickly when classic methods are too slow for finding an approximate solution, or when classic methods fail to find any exact solution. This is achieved by trading optimality, completeness, accuracy, or precision for speed. In a way, it can be considered a shortcut. A heuristic function, also simply called a heuristic, is a function that ranks alternatives in search algorithms at each branching step based on available information to decide which branch to follow. For example, it may approximate the exact solution. Definition and motivation The objective of a heuristic is to produce a solution in a reasonable time frame that is good enough for solving the problem at hand. This solution may not be the best of all the solutions to this problem, or it may simply approximate the exact solution. But it is still valuable ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
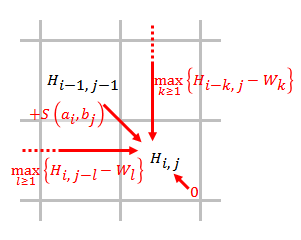
Smith–Waterman Algorithm
The Smith–Waterman algorithm performs local sequence alignment; that is, for determining similar regions between two strings of nucleic acid sequences or protein sequences. Instead of looking at the entire sequence, the Smith–Waterman algorithm compares segments of all possible lengths and optimizes the similarity measure. The algorithm was first proposed by Temple F. Smith and Michael S. Waterman in 1981. Like the Needleman–Wunsch algorithm, of which it is a variation, Smith–Waterman is a dynamic programming algorithm. As such, it has the desirable property that it is guaranteed to find the optimal local alignment with respect to the scoring system being used (which includes the substitution matrix and the gap-scoring scheme). The main difference to the Needleman–Wunsch algorithm is that negative scoring matrix cells are set to zero, which renders the (thus positively scoring) local alignments visible. Traceback procedure starts at the highest scoring matrix cell ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Needleman–Wunsch Algorithm
The Needleman–Wunsch algorithm is an algorithm used in bioinformatics to align protein or nucleotide sequences. It was one of the first applications of dynamic programming to compare biological sequences. The algorithm was developed by Saul B. Needleman and Christian D. Wunsch and published in 1970. The algorithm essentially divides a large problem (e.g. the full sequence) into a series of smaller problems, and it uses the solutions to the smaller problems to find an optimal solution to the larger problem. It is also sometimes referred to as the optimal matching algorithm and the global alignment technique. The Needleman–Wunsch algorithm is still widely used for optimal global alignment, particularly when the quality of the global alignment is of the utmost importance. The algorithm assigns a score to every possible alignment, and the purpose of the algorithm is to find all possible alignments having the highest score. Introduction This algorithm can be used for any two s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Read (biology)
In DNA sequencing, a read is an inferred sequence of base pairs (or base pair probabilities) corresponding to all or part of a single DNA fragment. A typical sequencing experiment involves fragmentation of the genome into millions of molecules, which are size-selected and ligated to adapters. The set of fragments is referred to as a sequencing library, which is sequenced to produce a set of reads. Read length Sequencing technologies vary in the length of reads produced. Reads of length 20-40 base pairs (bp) are referred to as ultra-short. Typical sequencers produce read lengths in the range of 100-500 bp. However, Pacific Biosciences platforms produce read lengths of approximately 1500 bp. Read length is a factor which can affect the results of biological studies. For example, longer read lengths improve the resolution of ''de novo'' genome assembly and detection of structural variants. It is estimated that read lengths greater than 100 kilobases (kb) will be required for routi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SAM (file Format)
Sequence Alignment Map (SAM) is a text-based format originally for storing biological sequences aligned to a reference sequence developed by Heng Li and Bob Handsaker ''et al''. It was developed when the 1000 Genomes Project wanted to move away from the MAQ mapper format and decided to design a new format. The overall TAB-delimited flavour of the format came from an earlier format inspired by BLAT’s PSL. The name of SAM came from Gabor Marth from University of Utah, who originally had a format under the same name but with a different syntax more similar to a BLAST output. It is widely used for storing data, such as nucleotide sequences, generated by next generation sequencing technologies, and the standard has been broadened to include unmapped sequences. The format supports short and long reads (up to 128 Mbp) produced by different sequencing platforms and is used to hold mapped data within the Genome Analysis Toolkit (GATK) and across the Broad Institute, the Wellcome ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
BioPerl
BioPerl is a collection of Perl modules that facilitate the development of Perl scripts for bioinformatics applications. It has played an integral role in the Human Genome Project. Background BioPerl is an active open source software project supported by the Open Bioinformatics Foundation. The first set of Perl codes of BioPerl was created by Tim Hubbard and Jong Bhak at MRC Centre Cambridge, where the first genome sequencing was carried out by Fred Sanger. MRC Centre was one of the hubs and birth places of modern bioinformatics as it had a large quantity of DNA sequences and 3D protein structures. Hubbard was using the th_lib.pl Perl library, which contained many useful Perl subroutines for bioinformatics. Bhak, Hubbard's first PhD student, created jong_lib.pl. Bhak merged the two Perl subroutine libraries into Bio.pl. The name BioPerl was coined jointly by Bhak and Steven Brenner at the Centre for Protein Engineering (CPE). In 1995, Brenner organized a BioPerl session at the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
BioRuby
BioRuby is a collection of open-source Ruby code, comprising classes for computational molecular biology and bioinformatics. It contains classes for DNA and protein sequence analysis, sequence alignment, biological database parsing, structural biology and other bioinformatics tasks. BioRuby is released under the GNU GPL version 2 or Ruby licence and is one of a number of Bio* projects, designed to reduce code duplication. In 2011, the BioRuby project introduced the Biogem software plugin system, with two or three new plugins added every month. BioRuby is managed via the BioRuby website and GitHub repository. History BioRuby The BioRuby project was first started in 2000 by Toshiaki Katayama as a Ruby implementation of similar bioinformatics packages such as BioPerl and BioPython. The initial release of version 0.1 was frequently updated by contributors both informally and at organised “hackathon” events; in June 2005, BioRuby was funded by IPA as an Exploratory Software Proj ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
EMBOSS
EMBOSS is a free open source software analysis package developed for the needs of the molecular biology and bioinformatics user community. The software automatically copes with data in a variety of formats and even allows transparent retrieval of sequence data from the web. Also, as extensive libraries are provided with the package, it is a platform to allow other scientists to develop and release software in true open source spirit. EMBOSS also integrates a range of currently available packages and tools for sequence analysis into a seamless whole. ''EMBOSS'' is an acronym for European Molecular Biology Open Software Suite. The ''European'' part of the name hints at the wider scope. The core EMBOSS groups are collaborating with many other groups to develop the new applications that the users need. This was done from the beginning with EMBnet, the European Molecular Biology Network. EMBnet has many nodes worldwide most of which are national bioinformatics services. EMBnet has the p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
GenBank
The GenBank sequence database is an open access, annotated collection of all publicly available nucleotide sequences and their protein translations. It is produced and maintained by the National Center for Biotechnology Information (NCBI; a part of the National Institutes of Health in the United States) as part of the International Nucleotide Sequence Database Collaboration (INSDC). GenBank and its collaborators receive sequences produced in laboratories throughout the world from more than 500,000 formally described species. The database started in 1982 by Walter Goad and Los Alamos National Laboratory. GenBank has become an important database for research in biological fields and has grown in recent years at an exponential rate by doubling roughly every 18 months. Release 250.0, published in June 2022, contained over 17 trillion nucleotide bases in more than 2,45 billion sequences. GenBank is built by direct submissions from individual laboratories, as well as from bulk submis ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
FASTA Format
In bioinformatics and biochemistry, the FASTA format is a text-based format for representing either nucleotide sequences or amino acid (protein) sequences, in which nucleotides or amino acids are represented using single-letter codes. The format also allows for sequence names and comments to precede the sequences. The format originates from the FASTA software package, but has now become a near universal standard in the field of bioinformatics. The simplicity of FASTA format makes it easy to manipulate and parse sequences using text-processing tools and scripting languages like the R programming language, Python, Ruby, Haskell, and Perl. Original format & overview The original FASTA/ Pearson format is described in the documentation for the FASTA suite of programs. It can be downloaded with any free distribution of FASTA (see fasta20.doc, fastaVN.doc or fastaVN.me—where VN is the Version Number). In the original format, a sequence was represented as a series of lines, each ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |