|
CpG Dinucleotide
The CpG sites or CG sites are regions of DNA where a cytosine nucleotide is followed by a guanine nucleotide in the linear sequence of bases along its 5' → 3' direction. CpG sites occur with high frequency in genomic regions called CpG islands (or CG islands). Cytosines in CpG dinucleotides can be methylated to form 5-methylcytosines. Enzymes that add a methyl group are called DNA methyltransferases. In mammals, 70% to 80% of CpG cytosines are methylated. Methylating the cytosine within a gene can change its expression, a mechanism that is part of a larger field of science studying gene regulation that is called epigenetics. Methylated cytosines often mutate to thymines. In humans, about 70% of promoters located near the transcription start site of a gene (proximal promoters) contain a CpG island. CpG characteristics Definition ''CpG'' is shorthand for ''5'—C—phosphate—G—3' '', that is, cytosine and guanine separated by only one phosphate group; phosphate ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
CpG Vs C-G Bp
CpG can be: *CpG site - methylated sequences of DNA significant in gene regulation *CpG Oligodeoxynucleotide CpG oligodeoxynucleotides (or CpG ODN) are short single-stranded synthetic DNA molecules that contain a cytosine triphosphate deoxynucleotide ("C") followed by a guanine triphosphate deoxynucleotide ("G"). The "p" refers to the phosphodiester ... - unmethylated sequences of DNA that have immunostimulatory properties * CpG island - regions of DNA that contain several CpG sites {{disambig ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Base Pair
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA and RNA. Dictated by specific hydrogen bonding patterns, "Watson–Crick" (or "Watson–Crick–Franklin") base pairs (guanine–cytosine and adenine–thymine) allow the DNA helix to maintain a regular helical structure that is subtly dependent on its nucleotide sequence. The complementary nature of this based-paired structure provides a redundant copy of the genetic information encoded within each strand of DNA. The regular structure and data redundancy provided by the DNA double helix make DNA well suited to the storage of genetic information, while base-pairing between DNA and incoming nucleotides provides the mechanism through which DNA polymerase replicates DNA and RNA polymerase transcribes DNA into RNA. Many DNA-binding prot ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
CpG Island
The CpG sites or CG sites are regions of DNA where a cytosine nucleotide is followed by a guanine nucleotide in the linear sequence of bases along its 5' → 3' direction. CpG sites occur with high frequency in genomic regions called CpG islands (or CG islands). Cytosines in CpG dinucleotides can be methylated to form 5-methylcytosines. Enzymes that add a methyl group are called DNA methyltransferases. In mammals, 70% to 80% of CpG cytosines are methylated. Methylating the cytosine within a gene can change its expression, a mechanism that is part of a larger field of science studying gene regulation that is called epigenetics. Methylated cytosines often mutate to thymines. In humans, about 70% of promoters located near the transcription start site of a gene (proximal promoters) contain a CpG island. CpG characteristics Definition ''CpG'' is shorthand for ''5'—C—phosphate—G—3' '', that is, cytosine and guanine separated by only one phosphate group; phosphate ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Adenine Phosphoribosyltransferase
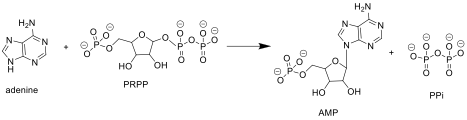
Adenine phosphoribosyltransferase (APRTase) is an enzyme encoded by the ''APRT'' gene, found in humans on chromosome 16. It is part of the Type I PRTase family and is involved in the nucleotide salvage pathway, which provides an alternative to nucleotide biosynthesis de novo in humans and most other animals. In parasitic protozoa such as giardia, APRTase provides the sole mechanism by which AMP can be produced. APRTase deficiency contributes to the formation of kidney stones ( urolithiasis) and to potential kidney failure. Function APRTase catalyzes the following reaction in the purine nucleotide salvage pathway: Adenine + Phosphoribosyl Pyrophosphate ( PRPP) → Adenylate (AMP) + Pyrophosphate ( PPi) In organisms that can synthesize purines de novo, the nucleotide salvage pathway provides an alternative that is energetically more efficient. It can salvage adenine from the polyamine biosynthetic pathway or from dietary sources of purines. Although APRTase is functional ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Norman Arnheim
Norman Arnheim is an American biologist specializing in aging and development biology, biochemistry, and molecular biology. He is currently a Distinguished Professor and the Ester Dornsife Chair at the University of Southern California, and an Elected Fellow A fellow is a concept whose exact meaning depends on context. In learned or professional societies, it refers to a privileged member who is specially elected in recognition of their work and achievements. Within the context of higher education ... of the American Association for the Advancement of Science. References Year of birth missing (living people) Living people University of Southern California faculty 21st-century American biologists {{US-biologist-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Philip Palmer Green
Philip Palmer Green is a theoretical and computational biologist noted for developing important algorithms and procedures used in Gene mapping and DNA sequencing. He earned his doctorate from Berkeley in mathematics in 1976 with a dissertation on C*-algebra under the direction of Marc Rieffel, but transitioned from pure mathematics into applied work in biology and bioinformatics. Green has obtained numerous important results, including in developing Phred, a widely used DNA trace analyzer, in mapping techniques, and in genetic analysis Genetic analysis is the overall process of studying and researching in fields of science that involve genetics and molecular biology. There are a number of applications that are developed from this research, and these are also considered parts of .... Green was elected to the National Academy of Sciences in 2001 and won the Gairdner Award in 2002. National Academy of Sciences (2004Biography of Phil Green PNAS 101(39), 13991–13993. See ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transition (genetics)
Transition, in genetics and molecular biology, refers to a point mutation that changes a purine nucleotide to another purine ( A ↔ G), or a pyrimidine nucleotide to another pyrimidine ( C ↔ T). Approximately two out of three single nucleotide polymorphisms (SNPs) are transitions. Transitions can be caused by oxidative deamination and tautomerization. Although there are twice as many possible transversions, transitions appear more often in genomes, possibly due to the molecular mechanisms that generate them. 5-Methylcytosine is more prone to transition than unmethylated cytosine, due to spontaneous deamination. This mechanism is important because it dictates the rarity of CpG islands The CpG sites or CG sites are regions of DNA where a cytosine nucleotide is followed by a guanine nucleotide in the linear sequence of bases along its 5' → 3' direction. CpG sites occur with high frequency in genomic regions called CpG i .... See also * Transversion References ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Base Excision Repair
Base excision repair (BER) is a cellular mechanism, studied in the fields of biochemistry and genetics, that repairs damaged DNA throughout the cell cycle. It is responsible primarily for removing small, non-helix-distorting base lesions from the genome. The related nucleotide excision repair pathway repairs bulky helix-distorting lesions. BER is important for removing damaged bases that could otherwise cause mutations by mispairing or lead to breaks in DNA during replication. BER is initiated by DNA glycosylases, which recognize and remove specific damaged or inappropriate bases, forming AP sites. These are then cleaved by an AP endonuclease. The resulting single-strand break can then be processed by either short-patch (where a single nucleotide is replaced) or long-patch BER (where 2–10 new nucleotides are synthesized). Lesions processed by BER Single bases in DNA can be chemically damaged by a variety of mechanisms, the most common ones being deamination, oxidation, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Uracil
Uracil () (symbol U or Ura) is one of the four nucleobases in the nucleic acid RNA. The others are adenine (A), cytosine (C), and guanine (G). In RNA, uracil binds to adenine via two hydrogen bonds. In DNA, the uracil nucleobase is replaced by thymine (T). Uracil is a demethylated form of thymine. Uracil is a common and naturally occurring pyrimidine derivative. The name "uracil" was coined in 1885 by the German chemist Robert Behrend, who was attempting to synthesize derivatives of uric acid. Originally discovered in 1900 by Alberto Ascoli, it was isolated by hydrolysis of yeast nuclein; it was also found in bovine thymus and spleen, herring sperm, and wheat germ. It is a planar, unsaturated compound that has the ability to absorb light. Based on 12C/13C isotopic ratios of organic compounds found in the Murchison meteorite, it is believed that uracil, xanthine, and related molecules can also be formed extraterrestrially. Data from the Cassini mission, orbiting in th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Thymine
Thymine () ( symbol T or Thy) is one of the four nucleobases in the nucleic acid of DNA that are represented by the letters G–C–A–T. The others are adenine, guanine, and cytosine. Thymine is also known as 5-methyluracil, a pyrimidine nucleobase. In RNA, thymine is replaced by the nucleobase uracil. Thymine was first isolated in 1893 by Albrecht Kossel and Albert Neumann from calf thymus glands, hence its name. Derivation As its alternate name (5-methyluracil) suggests, thymine may be derived by methylation of uracil at the 5th carbon. In RNA, thymine is replaced with uracil in most cases. In DNA, thymine (T) binds to adenine (A) via two hydrogen bonds, thereby stabilizing the nucleic acid structures. Thymine combined with deoxyribose creates the nucleoside deoxythymidine, which is synonymous with the term thymidine. Thymidine can be phosphorylated with up to three phosphoric acid groups, producing dTMP (deoxythymidine monophosphate), dTDP, or dTTP (for ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |