Southern blot on:
[Wikipedia]
[Google]
[Amazon]


 A Southern blot is a method used in
A Southern blot is a method used in
OpenWetWare
{{Molecular probes Molecular biology techniques



 A Southern blot is a method used in
A Southern blot is a method used in molecular biology
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions. The study of chemical and physi ...
for detection of a specific DNA sequence
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. Th ...
in DNA samples. Southern blotting combines transfer of electrophoresis
Electrophoresis, from Ancient Greek ἤλεκτρον (ḗlektron, "amber") and φόρησις (phórēsis, "the act of bearing"), is the motion of dispersed particles relative to a fluid under the influence of a spatially uniform electric fie ...
-separated DNA fragments to a filter membrane and subsequent fragment detection by probe hybridization.
The method is named after the British
British may refer to:
Peoples, culture, and language
* British people, nationals or natives of the United Kingdom, British Overseas Territories, and Crown Dependencies.
** Britishness, the British identity and common culture
* British English, ...
biologist
A biologist is a scientist who conducts research in biology. Biologists are interested in studying life on Earth, whether it is an individual Cell (biology), cell, a multicellular organism, or a Community (ecology), community of Biological inter ...
Edwin Southern
Sir Edwin Mellor Southern (born 7 June 1938) is an English Lasker Award-winning molecular biologist, Emeritus Professor of Biochemistry at the University of Oxford and a fellow of Trinity College, Oxford. He is most widely known for the inventio ...
, who first published it in 1975. Other blot
Blot may refer to:
Surname
* Guillaume Blot (born 1985), French racing cyclist
* Harold W. Blot (born 1938), served as United States Deputy Chief of Staff for Aviation
* Jean-François Joseph Blot (1781–1857), French soldier and politician
*Yv ...
ting methods (i.e., western blot
The western blot (sometimes called the protein immunoblot), or western blotting, is a widely used analytical technique in molecular biology and immunogenetics to detect specific proteins in a sample of tissue homogenate or extract. Besides detect ...
, northern blot
The northern blot, or RNA blot,Gilbert, S. F. (2000) Developmental Biology, 6th Ed. Sunderland MA, Sinauer Associates. is a technique used in molecular biology research to study gene expression by detection of RNA (or isolated mRNA) in a sample.K ...
, eastern blot
The eastern blot, or eastern blotting, is a biochemical technique used to analyze protein post-translational modifications including the addition of lipids, phosphates, and glycoconjugates. It is most often used to detect carbohydrate epitopes. Thu ...
, southwestern blot
The southwestern blot, is a lab technique that involves identifying as well as characterizing DNA-binding proteins by their ability to bind to specific oligonucleotide probes. Determination of molecular weight of proteins binding to DNA is also ...
) that employ similar principles, but using RNA or protein, have later been named in reference to Edwin Southern's name. As the label is eponymous, Southern is capitalised, as is conventional of proper noun
A proper noun is a noun that identifies a single entity and is used to refer to that entity (''Africa'', ''Jupiter'', ''Sarah'', ''Microsoft)'' as distinguished from a common noun, which is a noun that refers to a class of entities (''continent, ...
s. The names for other blotting methods may follow this convention, by analogy.
Method
#Restrictionendonuclease
Endonucleases are enzymes that cleave the phosphodiester bond within a polynucleotide chain. Some, such as deoxyribonuclease I, cut DNA relatively nonspecifically (without regard to sequence), while many, typically called restriction endonucleases ...
s are used to cut high-molecular-weight DNA strands into smaller fragments.
#The DNA fragments are then electrophoresed on an agarose gel
Agarose gel electrophoresis is a method of gel electrophoresis used in biochemistry, molecular biology, genetics, and clinical chemistry to separate a mixed population of macromolecules such as DNA or proteins in a matrix of agarose, one of the ...
to separate them by size.
# If some of the DNA fragments are larger than 15 kb, then prior to blotting, the gel may be treated with an acid, such as dilute HCl HCL may refer to:
Science and medicine
* Hairy cell leukemia, an uncommon and slowly progressing B cell leukemia
* Harvard Cyclotron Laboratory, from 1961 to 2002, a proton accelerator used for research and development
* Hollow-cathode lamp, a spe ...
. This depurinates the DNA fragments, breaking the DNA into smaller pieces, thereby allowing more efficient transfer from the gel to membrane.
# If alkaline transfer methods are used, the DNA gel is placed into an alkaline solution (typically containing sodium hydroxide
Sodium hydroxide, also known as lye and caustic soda, is an inorganic compound with the formula NaOH. It is a white solid ionic compound consisting of sodium cations and hydroxide anions .
Sodium hydroxide is a highly caustic base and alkali ...
) to denature the double-stranded DNA. The denaturation in an alkaline environment may improve binding of the negatively charged thymine residues of DNA to a positively charged amino groups of membrane, separating it into single DNA strands for later hybridization to the probe (see below), and destroys any residual RNA that may still be present in the DNA. The choice of alkaline over neutral transfer methods, however, is often empirical and may result in equivalent results.
# A sheet of nitrocellulose
Nitrocellulose (also known as cellulose nitrate, flash paper, flash cotton, guncotton, pyroxylin and flash string, depending on form) is a highly flammable compound formed by nitrating cellulose through exposure to a mixture of nitric acid and ...
(or, alternatively, nylon
Nylon is a generic designation for a family of synthetic polymers composed of polyamides ( repeating units linked by amide links).The polyamides may be aliphatic or semi-aromatic.
Nylon is a silk-like thermoplastic, generally made from petro ...
) membrane
A membrane is a selective barrier; it allows some things to pass through but stops others. Such things may be molecules, ions, or other small particles. Membranes can be generally classified into synthetic membranes and biological membranes. B ...
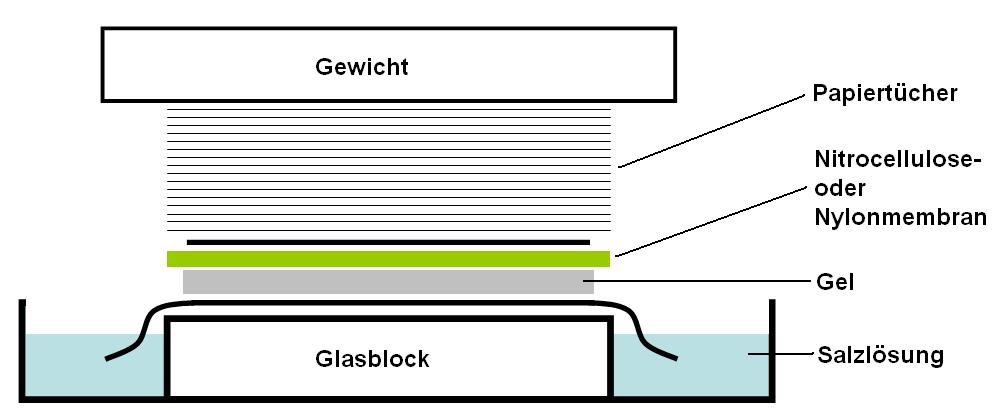
is placed on top of (or below, depending on the direction of the transfer) the gel. Pressure is applied evenly to the gel (either using suction, or by placing a stack of paper towels and a weight on top of the membrane and gel), to ensure good and even contact between gel and membrane. If transferring by suction, 20X SSC buffer is used to ensure a seal and prevent drying of the gel. Buffer transfer by capillary action
Capillary action (sometimes called capillarity, capillary motion, capillary rise, capillary effect, or wicking) is the process of a liquid flowing in a narrow space without the assistance of, or even in opposition to, any external forces li ...
from a region of high water potential
Water potential is the potential energy of water per unit volume relative to pure water in reference conditions. Water potential quantifies the tendency of water to move from one area to another due to osmosis, gravity, mechanical pressure and mat ...
to a region of low water potential (usually filter paper and paper tissues) is then used to move the DNA from the gel onto the membrane; ion exchange
Ion exchange is a reversible interchange of one kind of ion present in an insoluble solid with another of like charge present in a solution surrounding the solid with the reaction being used especially for softening or making water demineralised, ...
interactions bind the DNA to the membrane due to the negative charge of the DNA and positive charge of the membrane.
# The membrane is then baked in a vacuum or regular oven at 80 °C for 2 hours (standard conditions; nitrocellulose or nylon membrane) or exposed to ultraviolet radiation
Ultraviolet (UV) is a form of electromagnetic radiation with wavelength from 10 nm (with a corresponding frequency around 30 PHz) to 400 nm (750 THz), shorter than that of visible light, but longer than X-rays. UV radiation i ...
(nylon membrane) to permanently attach the transferred DNA to the membrane.
# The membrane is then exposed to a hybridization probe
In molecular biology, a hybridization probe (HP) is a fragment of DNA or RNA of usually 15–10000 nucleotide long which can be radioactively or fluorescently labeled. HP can be used to detect the presence of nucleotide sequences in analyzed RNA ...
—a single DNA fragment with a specific sequence whose presence in the target DNA is to be determined. The probe DNA is labelled so that it can be detected, usually by incorporating radioactivity or tagging the molecule with a fluorescent
Fluorescence is the emission of light by a substance that has absorbed light or other electromagnetic radiation. It is a form of luminescence. In most cases, the emitted light has a longer wavelength, and therefore a lower photon energy, tha ...
or chromogenic dye. In some cases, the hybridization probe may be made from RNA, rather than DNA. To ensure the specificity of the binding of the probe to the sample DNA, most common hybridization methods use salmon or herring sperm DNA for blocking of the membrane surface and target DNA, deionized formamide
Formamide is an amide derived from formic acid. It is a colorless liquid which is miscible with water and has an ammonia-like odor. It is chemical feedstock for the manufacture of sulfa drugs and other pharmaceuticals, herbicides and pesticide ...
, and detergents such as SDS to reduce non-specific binding of the probe.
# After hybridization, excess probe is washed from the membrane (typically using SSC buffer In biochemistry and molecular biology, saline-sodium citrate (SSC) buffer is used as a hybridization buffer (chemistry), buffer, to control stringency for washing steps in protocols for Southern blotting, in situ hybridization, DNA Microarray or Nor ...
), and the pattern of hybridization is visualized on X-ray
An X-ray, or, much less commonly, X-radiation, is a penetrating form of high-energy electromagnetic radiation. Most X-rays have a wavelength ranging from 10 picometers to 10 nanometers, corresponding to frequencies in the range 30&nb ...
film by autoradiography in the case of a radioactive or fluorescent probe, or by development of colour on the membrane if a chromogenic detection method is used.
Result
Hybridization of the probe to a specific DNA fragment on the filter membrane indicates that this fragment contains DNA sequence that is complementary to the probe. The transfer step of the DNA from the electrophoresis gel to a membrane permits easy binding of the labeled hybridization probe to the size-fractionated DNA. It also allows for the fixation of the target-probe hybrids, required for analysis by autoradiography or other detection methods. Southern blots performed with restriction enzyme-digested genomic DNA may be used to determine the number of sequences (e.g., gene copies) in agenome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding ge ...
. A probe that hybridizes only to a single DNA segment that has not been cut by the restriction enzyme will produce a single band on a Southern blot, whereas multiple bands will likely be observed when the probe hybridizes to several highly similar sequences (e.g., those that may be the result of sequence duplication). Modification of the hybridization conditions (for example, increasing the hybridization temperature or decreasing salt concentration) may be used to increase specificity and decrease hybridization of the probe to sequences that are less than 100% similar.
Applications
Southern blotting transfer may be used for homology-based cloning on the basis of amino acid sequence of the protein product of the target gene.Oligonucleotide
Oligonucleotides are short DNA or RNA molecules, oligomers, that have a wide range of applications in genetic testing, research, and forensics. Commonly made in the laboratory by solid-phase chemical synthesis, these small bits of nucleic acids c ...
s are designed so that they are complementary to the target sequence. The oligonucleotides are chemically synthesized, radiolabeled, and used to screen a DNA library, or other collections of cloned DNA fragments. Sequences that hybridize with the hybridization probe are further analysed, for example, to obtain the full length sequence of the targeted gene.
Southern blotting can also be used to identify methylated sites in particular genes. Particularly useful are the restriction nucleases ''MspI'' and ''HpaII'', both of which recognize and cleave within the same sequence. However, ''HpaII'' requires that a C within that site be methylated, whereas ''MspI'' cleaves only DNA unmethylated at that site. Therefore, any methylated sites within a sequence analyzed with a particular probe will be cleaved by the former, but not the latter, enzyme.Biochemistry 3rd Edition, Matthews, Van Holde et al, Addison Wesley Publishing, pg 977
See also
*Gel electrophoresis of nucleic acids
Nucleic acid electrophoresis is an analytical technique used to separate DNA or RNA fragments by size and reactivity. Nucleic acid molecules which are to be analyzed are set upon a viscous medium, the gel, where an electric field induces the nu ...
* Restriction fragment
*Genetic fingerprint
DNA profiling (also called DNA fingerprinting) is the process of determining an individual's DNA characteristics. DNA analysis intended to identify a species, rather than an individual, is called DNA barcoding.
DNA profiling is a forensic tec ...
*Northern blot
The northern blot, or RNA blot,Gilbert, S. F. (2000) Developmental Biology, 6th Ed. Sunderland MA, Sinauer Associates. is a technique used in molecular biology research to study gene expression by detection of RNA (or isolated mRNA) in a sample.K ...
*Western blot
The western blot (sometimes called the protein immunoblot), or western blotting, is a widely used analytical technique in molecular biology and immunogenetics to detect specific proteins in a sample of tissue homogenate or extract. Besides detect ...
*Eastern blot
The eastern blot, or eastern blotting, is a biochemical technique used to analyze protein post-translational modifications including the addition of lipids, phosphates, and glycoconjugates. It is most often used to detect carbohydrate epitopes. Thu ...
*Southwestern blot
The southwestern blot, is a lab technique that involves identifying as well as characterizing DNA-binding proteins by their ability to bind to specific oligonucleotide probes. Determination of molecular weight of proteins binding to DNA is also ...
* Northwestern blot
References
External links
OpenWetWare
{{Molecular probes Molecular biology techniques