Short tandem repeat on:
[Wikipedia]
[Google]
[Amazon]
A microsatellite is a tract of repetitive DNA in which certain DNA motifs (ranging in length from one to six or more
 Unlike
Unlike

 Microsatellites were popularized in
Microsatellites were popularized in
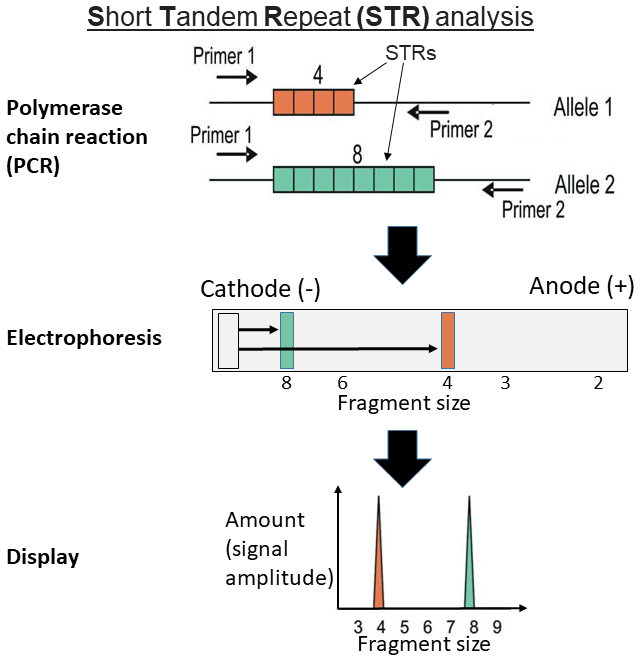 Repetitive DNA is not easily analysed by next generation DNA sequencing methods, which struggle with homopolymeric tracts. Therefore, microsatellites are normally analysed by conventional PCR amplification and amplicon size determination, sometimes followed by Sanger DNA sequencing.
In forensics, the analysis is performed by extracting
Repetitive DNA is not easily analysed by next generation DNA sequencing methods, which struggle with homopolymeric tracts. Therefore, microsatellites are normally analysed by conventional PCR amplification and amplicon size determination, sometimes followed by Sanger DNA sequencing.
In forensics, the analysis is performed by extracting
repeat masker
Once the potentially useful microsatellites are determined, the flanking sequences can be used to design
All known disease-causing short tandem repeatsMicroSatellite DataBase
* Search tools: *
*
IMEx
*
Imperfect SSR Finder
Ćöfind perfect or imperfect SSRs in
JSTRINGŌĆöJava Search for Tandem Repeats In Genomes
*
Microsatellite repeats finder
*
*
MREPATT
*
Mreps
*
Ćöa tandem repeat search tool for perfect and imperfect repeatsŌĆöthe maximum pattern size depends only on computational power *
Poly
*
*
SSR Finder
*
STAR
*
*
*
TRED
*
TROLL
*
Zebrafish Repeats
{{Authority control Genetics Forensic genetics Repetitive DNA sequences
base pairs
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA ...
) are repeated, typically 5ŌĆō50 times. Microsatellites occur at thousands of locations within an organism's genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding ge ...
. They have a higher mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA or viral replication, mi ...
rate than other areas of DNA leading to high genetic diversity
Genetic diversity is the total number of genetic characteristics in the genetic makeup of a species, it ranges widely from the number of species to differences within species and can be attributed to the span of survival for a species. It is dis ...
. Microsatellites are often referred to as short tandem repeats (STRs) by forensic geneticists and in genetic genealogy
Genetic genealogy is the use of genealogical DNA tests, i.e., DNA profiling and DNA testing, in combination with traditional genealogical methods, to infer genetic relationships between individuals. This application of genetics came to be used b ...
, or as simple sequence repeats (SSRs) by plant geneticists.
Microsatellites and their longer cousins, the minisatellite
A minisatellite is a tract of repetitive DNA in which certain DNA motifs (ranging in length from 10ŌĆō60 base pairs) are typically repeated 5-50 times. Minisatellites occur at more than 1,000 locations in the human genome and they are notable for ...
s, together are classified as VNTR
A variable number tandem repeat (or VNTR) is a location in a genome where a short nucleotide sequence is organized as a tandem repeat. These can be found on many chromosomes, and often show Polymorphism (biology), variations in length (number of ...
(variable number of tandem repeat
Tandem repeats occur in DNA when a pattern of one or more nucleotides is repeated and the repetitions are directly adjacent to each other. Several protein domains also form tandem repeats within their amino acid primary structure, such as armadil ...
s) DNA. The name "satellite" DNA refers to the early observation that centrifugation of genomic DNA in a test tube separates a prominent layer of bulk DNA from accompanying "satellite" layers of repetitive DNA.
They are widely used for DNA profiling
DNA profiling (also called DNA fingerprinting) is the process of determining an individual's DNA characteristics. DNA analysis intended to identify a species, rather than an individual, is called DNA barcoding.
DNA profiling is a forensic tec ...
in cancer diagnosis
Cancer is a group of diseases involving abnormal cell growth with the potential to invade or spread to other parts of the body. These contrast with benign tumors, which do not spread. Possible signs and symptoms include a lump, abnormal bl ...
, in kinship
In anthropology, kinship is the web of social relationships that form an important part of the lives of all humans in all societies, although its exact meanings even within this discipline are often debated. Anthropologist Robin Fox says that ...
analysis (especially paternity testing
DNA paternity testing is the use of DNA profiles to determine whether an individual is the biological parent of another individual. Paternity testing can be especially important when the rights and duties of the father are in issue and a child ...
) and in forensic identification. They are also used in genetic linkage
Genetic linkage is the tendency of DNA sequences that are close together on a chromosome to be inherited together during the meiosis phase of sexual reproduction. Two genetic markers that are physically near to each other are unlikely to be separ ...
analysis to locate a gene or a mutation responsible for a given trait or disease. Microsatellites are also used in population genetics
Population genetics is a subfield of genetics that deals with genetic differences within and between populations, and is a part of evolutionary biology. Studies in this branch of biology examine such phenomena as adaptation, speciation, and pop ...
to measure levels of relatedness between subspecies, groups and individuals.
History
Although the first microsatellite was characterised in 1984 at theUniversity of Leicester
, mottoeng = So that they may have life
, established =
, type = public research university
, endowment = ┬Ż20.0 million
, budget = ┬Ż326 million
, chancellor = David Willetts
, vice_chancellor = Nishan Canagarajah
, head_labe ...
by Weller, Jeffreys Jeffreys is a surname, which may refer to:
People: (See also the common variants Jeffries and Jefferies)
* Alec Jeffreys (born 1950), British biologist and discoverer of DNA fingerprinting
* Anne Jeffreys (1923ŌĆō2017), American actress and sing ...
and colleagues as a polymorphic GGAT repeat in the human myoglobin gene, the term "microsatellite" was introduced later, in 1989, by Litt and Luty. The name "satellite" DNA refers to the early observation that centrifugation of genomic DNA in a test tube separates a prominent layer of bulk DNA from accompanying "satellite" layers of repetitive DNA. The increasing availability of DNA amplification by PCR at the beginning of the 1990s triggered a large number of studies using the amplification of microsatellites as genetic markers for forensic medicine, for paternity testing, and for positional cloning to find the gene underlying a trait or disease. Prominent early applications include the identifications by microsatellite genotyping of the eight-year-old skeletal remains of a British murder victim (Hagelberg et al. 1991), and of the Auschwitz concentration camp doctor Josef Mengele
, allegiance =
, branch = Schutzstaffel
, serviceyears = 1938ŌĆō1945
, rank = ''Schutzstaffel, SS''-''Hauptsturmf├╝hrer'' (Captain)
, servicenumber =
, battles =
, unit =
, awards =
, command ...
who escaped to South America following World War II (Jeffreys et al. 1992).
Structures, locations, and functions
A microsatellite is a tract of tandemly repeated (i.e. adjacent) DNA motifs that range in length from one to six or up to ten nucleotides (the exact definition and delineation to the longer minisatellites varies from author to author), and are typically repeated 5ŌĆō50 times. For example, the sequence TATATATATA is a dinucleotide microsatellite, and GTCGTCGTCGTCGTC is a trinucleotide microsatellite (with A beingAdenine
Adenine () ( symbol A or Ade) is a nucleobase (a purine derivative). It is one of the four nucleobases in the nucleic acid of DNA that are represented by the letters GŌĆōCŌĆōAŌĆōT. The three others are guanine, cytosine and thymine. Its derivati ...
, G Guanine
Guanine () ( symbol G or Gua) is one of the four main nucleobases found in the nucleic acids DNA and RNA, the others being adenine, cytosine, and thymine (uracil in RNA). In DNA, guanine is paired with cytosine. The guanine nucleoside is called ...
, C Cytosine
Cytosine () ( symbol C or Cyt) is one of the four nucleobases found in DNA and RNA, along with adenine, guanine, and thymine (uracil in RNA). It is a pyrimidine derivative, with a heterocyclic aromatic ring and two substituents attached (an am ...
, and T Thymine
Thymine () ( symbol T or Thy) is one of the four nucleobases in the nucleic acid of DNA that are represented by the letters GŌĆōCŌĆōAŌĆōT. The others are adenine, guanine, and cytosine. Thymine is also known as 5-methyluracil, a pyrimidine nu ...
). Repeat units of four and five nucleotides are referred to as tetra- and pentanucleotide motifs, respectively. Most eukaryotes have microsatellites, with the notable exception of some yeast species. Microsatellites are distributed throughout the genome. The human genome for example contains 50,000ŌĆō100,000 dinucleotide microsatellites, and lesser numbers of tri-, tetra- and pentanucleotide microsatellites. Many are located in non-coding parts of the human genome and therefore do not produce proteins, but they can also be located in regulatory regions and coding region
The coding region of a gene, also known as the coding sequence (CDS), is the portion of a gene's DNA or RNA that codes for protein. Studying the length, composition, regulation, splicing, structures, and functions of coding regions compared to no ...
s.
Microsatellites in non-coding regions may not have any specific function, and therefore might not be selected against; this allows them to accumulate mutations unhindered over the generations and gives rise to variability that can be used for DNA fingerprinting and identification purposes. Other microsatellites are located in regulatory flanking or intronic
An intron is any nucleotide sequence within a gene that is not expressed or operative in the final RNA product. The word ''intron'' is derived from the term ''intragenic region'', i.e. a region inside a gene."The notion of the cistron .e., gene ...
regions of genes, or directly in codon
The genetic code is the set of rules used by living cells to translate information encoded within genetic material ( DNA or RNA sequences of nucleotide triplets, or codons) into proteins. Translation is accomplished by the ribosome, which links ...
s of genes ŌĆō microsatellite mutations in such cases can lead to phenotypic changes and diseases, notably in triplet expansion diseases such as fragile X syndrome
Fragile X syndrome (FXS) is a genetic disorder characterized by mild-to-moderate intellectual disability. The average IQ in males with FXS is under 55, while about two thirds of affected females are intellectually disabled. Physical features may ...
and Huntington's disease
Huntington's disease (HD), also known as Huntington's chorea, is a neurodegenerative disease that is mostly inherited. The earliest symptoms are often subtle problems with mood or mental abilities. A general lack of coordination and an unst ...
.
Telomeres
A telomere (; ) is a region of repetitive nucleotide sequences associated with specialized proteins at the ends of linear chromosomes. Although there are different architectures, telomeres, in a broad sense, are a widespread genetic feature mos ...
are linear sequences of DNA that sit at the very ends of chromosomes and protect the integrity of genomic material (not unlike an aglet
An aglet ( ) or aiglet is a small sheath, often made of plastic or metal, attached at each end of a shoelace, a cord, or a drawstring. An aglet keeps the fibers of the lace or cord from unraveling; its firmness and narrow profile make it easier ...
on the end of a shoelace) during successive rounds of cell division due to the "end replication problem". In white blood cells, the gradual shortening of telomeric DNA has been shown to inversely correlate with aging in several sample types. Telomeres consist of repetitive DNA, with the hexanucleotide repeat motif TTAGGG in vertebrates. They are thus classified as minisatellite
A minisatellite is a tract of repetitive DNA in which certain DNA motifs (ranging in length from 10ŌĆō60 base pairs) are typically repeated 5-50 times. Minisatellites occur at more than 1,000 locations in the human genome and they are notable for ...
s. Similarly, insects have shorter repeat motifs in their telomeres that could arguably be considered microsatellites.
Mutation mechanisms and mutation rates
 Unlike
Unlike point mutations
A point mutation is a genetic mutation where a single nucleotide base is changed, inserted or deleted from a DNA or RNA sequence of an organism's genome. Point mutations have a variety of effects on the downstream protein productŌĆöconsequences ...
, which affect only a single nucleotide, microsatellite mutations lead to the gain or loss of an entire repeat unit, and sometimes two or more repeats simultaneously. Thus, the mutation rate
In genetics, the mutation rate is the frequency of new mutations in a single gene or organism over time. Mutation rates are not constant and are not limited to a single type of mutation; there are many different types of mutations. Mutation rates ...
at microsatellite loci is expected to differ from other mutation rates, such as base substitution rates. The actual cause of mutations in microsatellites is debated.
One proposed cause of such length changes is replication slippage, caused by mismatches between DNA strands while being replicated during meiosis. DNA polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create ...
, the enzyme responsible for reading DNA during replication, can slip while moving along the template strand and continue at the wrong nucleotide. DNA polymerase slippage is more likely to occur when a repetitive sequence (such as CGCGCG) is replicated. Because microsatellites consist of such repetitive sequences, DNA polymerase may make errors at a higher rate in these sequence regions. Several studies have found evidence that slippage is the cause of microsatellite mutations. Typically, slippage in each microsatellite occurs about once per 1,000 generations. Thus, slippage changes in repetitive DNA are three orders of magnitude more common than point mutations in other parts of the genome. Most slippage results in a change of just one repeat unit, and slippage rates vary for different allele lengths and repeat unit sizes, and within different species. If there is a large size difference between individual alleles, then there may be increased instability during recombination at meiosis.
Another possible cause of microsatellite mutations are point mutations, where only one nucleotide is incorrectly copied during replication. A study comparing human and primate genomes found that most changes in repeat number in short microsatellites appear due to point mutations rather than slippage.
Microsatellite mutation rates
Microsatellite mutation rates vary with base position relative to the microsatellite, repeat type, and base identity. Mutation rate rises specifically with repeat number, peaking around six to eight repeats and then decreasing again. Increased heterozygosity in a population will also increase microsatellite mutation rates, especially when there is a large length difference between alleles. This is likely due tohomologous chromosomes
A couple of homologous chromosomes, or homologs, are a set of one maternal and one paternal chromosome that pair up with each other inside a cell during fertilization. Homologs have the same genes in the same loci where they provide points alon ...
with arms of unequal lengths causing instability during meiosis.
Direct estimates of microsatellite mutation rates have been made in numerous organisms, from insects to humans. In the desert locust
The desert locust (''Schistocerca gregaria'') is a species of locust, a periodically swarming, short-horned grasshopper in the family Acrididae. They are found primarily in the deserts and dry areas of northern and eastern Africa, Arabia, and sou ...
''Schistocerca gregaria'', the microsatellite mutation rate was estimated at 2.1 x 10ŌłÆ4 per generation per locus. The microsatellite mutation rate in human male germ lines is five to six times higher than in female germ lines and ranges from 0 to 7 x 10ŌłÆ3 per locus per gamete per generation. In the nematode ''Pristionchus pacificus
''Pristionchus pacificus'' is a species of free-living nematodes (roundworms) in the family Diplogastridae. The species has been established as a satellite model organism to '' Caenorhabditis elegans'', with which it shared a common ancestor ...
'', the estimated microsatellite mutation rate ranges from 8.9 ├Ś 10ŌłÆ5 to 7.5 ├Ś 10ŌłÆ4 per locus per generation.
Biological effects of microsatellite mutations
Many microsatellites are located in non-coding DNA and are biologically silent. Others are located in regulatory or even coding DNA ŌĆō microsatellite mutations in such cases can lead to phenotypic changes and diseases. A genome-wide study estimates that microsatellite variation contributes 10ŌĆō15% of heritable gene expression variation in humans.Effects on proteins
In mammals, 20% to 40% of proteins contain repeating sequences of amino acids encoded by short sequence repeats. Most of the short sequence repeats within protein-coding portions of the genome have a repeating unit of three nucleotides, since that length will not cause frame-shifts when mutating. Each trinucleotide repeating sequence is transcribed into a repeating series of the same amino acid. In yeasts, the most common repeated amino acids are glutamine, glutamic acid, asparagine, aspartic acid and serine. Mutations in these repeating segments can affect the physical and chemical properties of proteins, with the potential for producing gradual and predictable changes in protein action. For example, length changes in tandemly repeating regions in the Runx2 gene lead to differences in facial length in domesticated dogs (''Canis familiaris''), with an association between longer sequence lengths and longer faces. This association also applies to a wider range of Carnivora species. Length changes in polyalanine tracts within the HoxA13 gene are linked to Hand-Foot-Genital Syndrome, a developmental disorder in humans. Length changes in other triplet repeats are linked to more than 40 neurological diseases in humans, notably triplet expansion diseases such asfragile X syndrome
Fragile X syndrome (FXS) is a genetic disorder characterized by mild-to-moderate intellectual disability. The average IQ in males with FXS is under 55, while about two thirds of affected females are intellectually disabled. Physical features may ...
and Huntington's disease
Huntington's disease (HD), also known as Huntington's chorea, is a neurodegenerative disease that is mostly inherited. The earliest symptoms are often subtle problems with mood or mental abilities. A general lack of coordination and an unst ...
. Evolutionary changes from replication slippage also occur in simpler organisms. For example, microsatellite length changes are common within surface membrane proteins in yeast, providing rapid evolution in cell properties. Specifically, length changes in the FLO1 gene control the level of adhesion to substrates. Short sequence repeats also provide rapid evolutionary change to surface proteins in pathenogenic bacteria; this may allow them to keep up with immunological changes in their hosts. Length changes in short sequence repeats in a fungus (''Neurospora crassa'') control the duration of its circadian clock
A circadian clock, or circadian oscillator, is a biochemical oscillator that cycles with a stable phase (waves), phase and is synchronized with solar time.
Such a clock's ''in vivo'' period is necessarily almost exactly 24 hours (the earth's curre ...
cycles.
Effects on gene regulation
Length changes of microsatellites within promoters and other cis-regulatory regions can change gene expression quickly, between generations. The human genome contains many (>16,000) short sequence repeats in regulatory regions, which provide ŌĆśtuning knobsŌĆÖ on the expression of many genes. Length changes in bacterial SSRs can affect fimbriae formation in ''Haemophilus influenzae'', by altering promoter spacing. Dinucleotide microsatellites are linked to abundant variation in cis-regulatory control regions in the human genome. Microsatellites in control regions of the Vasopressin 1a receptor gene in voles influence their social behavior, and level of monogamy. InEwing's sarcoma
Ewing sarcoma is a type of cancer that forms in bone or soft tissue. Symptoms may include swelling and pain at the site of the tumor, fever, and a bone fracture. The most common areas where it begins are the legs, pelvis, and chest wall. In about ...
(a type of painful bone cancer in young humans), a point mutation has created an extended GGAA microsatellite which binds a transcription factor, which in turn activates the EGR2 gene which drives the cancer. In addition, other GGAA microsatellites may influence the expression of genes that contribute to the clinical outcome of Ewing sarcoma patients.
Effects within introns
Microsatellites withinintron
An intron is any nucleotide sequence within a gene that is not expressed or operative in the final RNA product. The word ''intron'' is derived from the term ''intragenic region'', i.e. a region inside a gene."The notion of the cistron .e., gene. ...
s also influence phenotype, through means that are not currently understood. For example, a GAA triplet expansion in the first intron of the X25 gene appears to interfere with transcription, and causes Friedreich Ataxia
Friedreich's ataxia (FRDA or FA) is an autosomal-recessive genetic disease that causes difficulty walking, a loss of sensation in the arms and legs, and impaired speech that worsens over time. Symptoms generally start between 5 and 20 year ...
. Tandem repeats in the first intron of the Asparagine synthetase gene are linked to acute lymphoblastic leukaemia. A repeat polymorphism in the fourth intron of the NOS3 gene is linked to hypertension in a Tunisian population. Reduced repeat lengths in the EGFR gene are linked with osteosarcomas.
An archaic form of splicing preserved in Zebrafish
The zebrafish (''Danio rerio'') is a freshwater fish belonging to the minnow family ( Cyprinidae) of the order Cypriniformes. Native to South Asia, it is a popular aquarium fish, frequently sold under the trade name zebra danio (and thus often ...
is known to use microsatellite sequences within intronic mRNA for the removal of introns in the absence of U2AF2 and other splicing machinery. It is theorized that these sequences form highly stable cloverleaf configurations that bring the 3' and 5' intron splice sites into close proximity, effectively replacing the spliceosome
A spliceosome is a large ribonucleoprotein (RNP) complex found primarily within the nucleus of eukaryotic cells. The spliceosome is assembled from small nuclear RNAs (snRNA) and numerous proteins. Small nuclear RNA (snRNA) molecules bind to specifi ...
. This method of RNA splicing is believed to have diverged from human evolution at the formation of tetrapod
Tetrapods (; ) are four-limbed vertebrate animals constituting the superclass Tetrapoda (). It includes extant and extinct amphibians, sauropsids ( reptiles, including dinosaurs and therefore birds) and synapsids (pelycosaurs, extinct theraps ...
s and to represent an artifact of an RNA world
The RNA world is a hypothetical stage in the evolutionary history of life on Earth, in which self-replicating RNA molecules proliferated before the evolution of DNA and proteins. The term also refers to the hypothesis that posits the existence ...
.
Effects within transposons
Almost 50% of the human genome is contained in various types of transposable elements (also called transposons, or ŌĆśjumping genesŌĆÖ), and many of them contain repetitive DNA. It is probable that short sequence repeats in those locations are also involved in the regulation of gene expression.Applications
Microsatellites are used for assessing chromosomal DNA deletions in cancer diagnosis. Microsatellites are widely used forDNA profiling
DNA profiling (also called DNA fingerprinting) is the process of determining an individual's DNA characteristics. DNA analysis intended to identify a species, rather than an individual, is called DNA barcoding.
DNA profiling is a forensic tec ...
, also known as "genetic fingerprinting", of crime stains (in forensics) and of tissues (in transplant patients). They are also widely used in kinship
In anthropology, kinship is the web of social relationships that form an important part of the lives of all humans in all societies, although its exact meanings even within this discipline are often debated. Anthropologist Robin Fox says that ...
analysis (most commonly in paternity testing). Also, microsatellites are used for mapping locations within the genome, specifically in genetic linkage
Genetic linkage is the tendency of DNA sequences that are close together on a chromosome to be inherited together during the meiosis phase of sexual reproduction. Two genetic markers that are physically near to each other are unlikely to be separ ...
analysis to locate a gene or a mutation responsible for a given trait or disease. As a special case of mapping, they can be used for studies of gene duplication
Gene duplication (or chromosomal duplication or gene amplification) is a major mechanism through which new genetic material is generated during molecular evolution. It can be defined as any duplication of a region of DNA that contains a gene. ...
or deletion. Researchers use microsatellites in population genetics
Population genetics is a subfield of genetics that deals with genetic differences within and between populations, and is a part of evolutionary biology. Studies in this branch of biology examine such phenomena as adaptation, speciation, and pop ...
and in species conservation projects. Plant geneticists have proposed the use of microsatellites for marker assisted selection
Marker assisted selection or marker aided selection (MAS) is an indirect selection process where a trait of interest is selected based on a marker ( morphological, biochemical or DNA/ RNA variation) linked to a trait of interest (e.g. producti ...
of desirable traits in plant breeding.
Cancer diagnosis
Intumour
A neoplasm () is a type of abnormal and excessive growth of tissue. The process that occurs to form or produce a neoplasm is called neoplasia. The growth of a neoplasm is uncoordinated with that of the normal surrounding tissue, and persists ...
cells, whose controls on replication are damaged, microsatellites may be gained or lost at an especially high frequency during each round of mitosis
In cell biology, mitosis () is a part of the cell cycle in which replicated chromosomes are separated into two new nuclei. Cell division by mitosis gives rise to genetically identical cells in which the total number of chromosomes is mainta ...
. Hence a tumour cell line might show a different genetic fingerprint
DNA profiling (also called DNA fingerprinting) is the process of determining an individual's DNA characteristics. DNA analysis intended to identify a species, rather than an individual, is called DNA barcoding.
DNA profiling is a forensic tec ...
from that of the host tissue, and, especially in colorectal cancer
Colorectal cancer (CRC), also known as bowel cancer, colon cancer, or rectal cancer, is the development of cancer from the colon or rectum (parts of the large intestine). Signs and symptoms may include blood in the stool, a change in bowel m ...
, might present with loss of heterozygosity
Loss of heterozygosity (LOH) is a type of genetic abnormality in diploid organisms in which one copy of an entire gene and its surrounding chromosomal region are lost. Since diploid cells have two copies of their genes, one from each parent, a sing ...
. Microsatellites have therefore been routinely used in cancer diagnosis to assess tumour progression.

Forensic and medical fingerprinting
Microsatellite analysis became popular in the field offorensics
Forensic science, also known as criminalistics, is the application of science to criminal and civil laws, mainlyŌĆöon the criminal sideŌĆöduring criminal investigation, as governed by the legal standards of admissible evidence and crimina ...
in the 1990s. It is used for the genetic fingerprinting
DNA profiling (also called DNA fingerprinting) is the process of determining an individual's DNA characteristics. DNA analysis intended to identify a species, rather than an individual, is called DNA barcoding.
DNA profiling is a forensic tec ...
of individuals where it permits forensic identification (typically matching a crime stain to a victim or perpetrator). It is also used to follow up bone marrow transplant
Hematopoietic stem-cell transplantation (HSCT) is the transplantation of multipotent hematopoietic stem cells, usually derived from bone marrow, peripheral blood, or umbilical cord blood in order to replicate inside of a patient and to produce ...
patients.
The microsatellites in use today for forensic analysis are all tetra- or penta-nucleotide repeats, as these give a high degree of error-free data while being short enough to survive degradation in non-ideal conditions. Even shorter repeat sequences would tend to suffer from artifacts such as PCR stutter and preferential amplification, while longer repeat sequences would suffer more highly from environmental degradation and would amplify less well by PCR. Another forensic consideration is that the person's medical privacy
Medical privacy or health privacy is the practice of maintaining the security and confidentiality of patient records. It involves both the conversational discretion of health care providers and the security of medical records. The terms can also r ...
must be respected, so that forensic STRs are chosen which are non-coding, do not influence gene regulation, and are not usually trinucleotide STRs which could be involved in triplet expansion diseases such as Huntington's disease
Huntington's disease (HD), also known as Huntington's chorea, is a neurodegenerative disease that is mostly inherited. The earliest symptoms are often subtle problems with mood or mental abilities. A general lack of coordination and an unst ...
. Forensic STR profiles are stored in DNA databanks such as the UK National DNA Database
The United Kingdom National DNA Database (NDNAD; officially the ''UK National Criminal Intelligence DNA Database'') is a national DNA Database that was set up in 1995. In 2005 it had 3.1 million profiles and in 2020 it had 6.6 million profiles ...
(NDNAD), the American CODIS
The Combined DNA Index System (CODIS) is the United States national DNA database created and maintained by the Federal Bureau of Investigation. CODIS consists of three levels of information; Local DNA Index Systems (LDIS) where DNA profiles orig ...
or the Australian NCIDD.
Kinship analysis (paternity testing)
Autosomal microsatellites are widely used forDNA profiling
DNA profiling (also called DNA fingerprinting) is the process of determining an individual's DNA characteristics. DNA analysis intended to identify a species, rather than an individual, is called DNA barcoding.
DNA profiling is a forensic tec ...
in kinship
In anthropology, kinship is the web of social relationships that form an important part of the lives of all humans in all societies, although its exact meanings even within this discipline are often debated. Anthropologist Robin Fox says that ...
analysis (most commonly in paternity testing). Paternally inherited Y-STR
A Y-STR is a short tandem repeat (STR) on the Y-chromosome. Y-STRs are often used in forensics, paternity, and genealogical DNA testing.
Y-STRs are taken specifically from the male Y chromosome. These Y-STRs provide a weaker analysis than autosoma ...
s (microsatellites on the Y chromosome
The Y chromosome is one of two sex chromosomes (allosomes) in therian mammals, including humans, and many other animals. The other is the X chromosome. Y is normally the sex-determining chromosome in many species, since it is the presence or abse ...
) are often used in genealogical DNA test
A genealogical DNA test is a DNA-based test used in genetic genealogy that looks at specific locations of a person's genome in order to find or verify ancestral genealogical relationships, or (with lower reliability) to estimate the ethnic mixt ...
ing.
Genetic linkage analysis
During the 1990s and the first several years of this millennium, microsatellites were the workhorse genetic markers for genome-wide scans to locate any gene responsible for a given phenotype or disease, usingsegregation Segregation may refer to:
Separation of people
* Geographical segregation, rates of two or more populations which are not homogenous throughout a defined space
* School segregation
* Housing segregation
* Racial segregation, separation of humans ...
observations across generations of a sampled pedigree. Although the rise of higher throughput and cost-effective single-nucleotide polymorphism
In genetics, a single-nucleotide polymorphism (SNP ; plural SNPs ) is a germline substitution of a single nucleotide at a specific position in the genome. Although certain definitions require the substitution to be present in a sufficiently lar ...
(SNP) platforms led to the era of the SNP for genome scans, microsatellites remain highly informative measures of genomic variation for linkage and association studies. Their continued advantage lies in their greater allelic diversity than biallelic SNPs, thus microsatellites can differentiate alleles within a SNP-defined linkage disequilibrium block of interest. Thus, microsatellites have successfully led to discoveries of type 2 diabetes (TCF7L2) and prostate cancer genes (the 8q21 region).
Population genetics
population genetics
Population genetics is a subfield of genetics that deals with genetic differences within and between populations, and is a part of evolutionary biology. Studies in this branch of biology examine such phenomena as adaptation, speciation, and pop ...
during the 1990s because as PCR became ubiquitous in laboratories researchers were able to design primers and amplify sets of microsatellites at low cost. Their uses are wide-ranging. A microsatellite with a neutral evolutionary history makes it applicable for measuring or inferring bottlenecks, local adaptation Local adaptation is a mechanism in evolutionary biology whereby a population of organisms evolves to be more well-suited to its local environment than other members of the same species that live elsewhere. Local adaptation requires that different po ...
, the allelic fixation index
The fixation index (FST) is a measure of population differentiation due to genetic structure. It is frequently estimated from genetic polymorphism data, such as single-nucleotide polymorphisms (SNP) or microsatellites. Developed as a special case ...
(FST), population size
In population genetics and population ecology, population size (usually denoted ''N'') is the number of individual organisms in a population. Population size is directly associated with amount of genetic drift, and is the underlying cause of effect ...
, and gene flow
In population genetics, gene flow (also known as gene migration or geneflow and allele flow) is the transfer of genetic material from one population to another. If the rate of gene flow is high enough, then two populations will have equivalent a ...
. As next generation sequencing
DNA sequencing is the process of determining the nucleic acid sequence ŌĆō the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. The ...
becomes more affordable the use of microsatellites has decreased, however they remain a crucial tool in the field.
Plant breeding
Marker assisted selection
Marker assisted selection or marker aided selection (MAS) is an indirect selection process where a trait of interest is selected based on a marker ( morphological, biochemical or DNA/ RNA variation) linked to a trait of interest (e.g. producti ...
or marker aided selection (MAS) is an indirect selection process where a trait of interest is selected based on a marker ( morphological, biochemical
Biochemistry or biological chemistry is the study of chemical processes within and relating to living organisms. A sub-discipline of both chemistry and biology, biochemistry may be divided into three fields: structural biology, enzymology an ...
or DNA/RNA
Ribonucleic acid (RNA) is a polymeric molecule essential in various biological roles in coding, decoding, regulation and expression of genes. RNA and deoxyribonucleic acid ( DNA) are nucleic acids. Along with lipids, proteins, and carbohydra ...
variation) linked to a trait of interest (e.g. productivity, disease resistance, stress tolerance, and quality), rather than on the trait itself. Microsatellites have been proposed to be used as such markers to assist plant breeding.
Analysis
 Repetitive DNA is not easily analysed by next generation DNA sequencing methods, which struggle with homopolymeric tracts. Therefore, microsatellites are normally analysed by conventional PCR amplification and amplicon size determination, sometimes followed by Sanger DNA sequencing.
In forensics, the analysis is performed by extracting
Repetitive DNA is not easily analysed by next generation DNA sequencing methods, which struggle with homopolymeric tracts. Therefore, microsatellites are normally analysed by conventional PCR amplification and amplicon size determination, sometimes followed by Sanger DNA sequencing.
In forensics, the analysis is performed by extracting nuclear DNA
Nuclear DNA (nDNA), or nuclear deoxyribonucleic acid, is the DNA contained within each cell nucleus of a eukaryotic organism. It encodes for the majority of the genome in eukaryotes, with mitochondrial DNA and plastid DNA coding for the rest. It ...
from the cells of a sample of interest, then amplifying specific polymorphic regions of the extracted DNA by means of the polymerase chain reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) t ...
. Once these sequences have been amplified, they are resolved either through gel electrophoresis
Gel electrophoresis is a method for separation and analysis of biomacromolecules ( DNA, RNA, proteins, etc.) and their fragments, based on their size and charge. It is used in clinical chemistry to separate proteins by charge or size (IEF ...
or capillary electrophoresis
Capillary electrophoresis (CE) is a family of electrokinetic separation methods performed in submillimeter diameter capillaries and in micro- and nanofluidic channels. Very often, CE refers to capillary zone electrophoresis (CZE), but other electr ...
, which will allow the analyst to determine how many repeats of the microsatellites sequence in question there are. If the DNA was resolved by gel electrophoresis, the DNA can be visualized either by silver stain In pathology, silver staining is the use of silver to selectively alter the appearance of a target in microscopy of histological sections; in temperature gradient gel electrophoresis; and in polyacrylamide gels.
In traditional stained glass, silve ...
ing (low sensitivity, safe, inexpensive), or an intercalating dye such as ethidium bromide
Ethidium bromide (or homidium bromide, chloride salt homidium chloride) is an intercalating agent commonly used as a fluorescent tag (nucleic acid stain) in molecular biology laboratories for techniques such as agarose gel electrophoresis. It i ...
(fairly sensitive, moderate health risks, inexpensive), or as most modern forensics labs use, fluorescent dyes
A fluorophore (or fluorochrome, similarly to a chromophore) is a fluorescent chemical compound that can re-emit light upon light excitation. Fluorophores typically contain several combined aromatic groups, or planar or cyclic molecules with se ...
(highly sensitive, safe, expensive). Instruments built to resolve microsatellite fragments by capillary electrophoresis also use fluorescent dyes. Forensic profiles are stored in major databanks. The British
British may refer to:
Peoples, culture, and language
* British people, nationals or natives of the United Kingdom, British Overseas Territories, and Crown Dependencies.
** Britishness, the British identity and common culture
* British English, ...
data base for microsatellite loci identification was originally based on the British SGM+ system using 10 loci and a sex marker. The Americans increased this number to 13 loci. The Australian database is called the NCIDD, and since 2013 it has been using 18 core markers for DNA profiling.
Amplification
Microsatellites can be amplified for identification by thepolymerase chain reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) t ...
(PCR) process, using the unique sequences of flanking regions as primers. DNA is repeatedly denatured at a high temperature to separate the double strand, then cooled to allow annealing of primers and the extension of nucleotide sequences through the microsatellite. This process results in production of enough DNA to be visible on agarose
Agarose is a heteropolysaccharide, generally extracted from certain red seaweed. It is a linear polymer made up of the repeating unit of agarobiose, which is a disaccharide made up of D-galactose and 3,6-anhydro-L-galactopyranose. Agarose is o ...
or polyacrylamide
Polyacrylamide (abbreviated as PAM) is a polymer with the formula (-CH2CHCONH2-). It has a linear-chain structure. PAM is highly water-absorbent, forming a soft gel when hydrated. In 2008, an estimated 750,000,000 kg were produced, mainly fo ...
gels; only small amounts of DNA are needed for amplification because in this way thermocycling creates an exponential increase in the replicated segment. With the abundance of PCR technology, primers that flank microsatellite loci are simple and quick to use, but the development of correctly functioning primers is often a tedious and costly process.
Design of microsatellite primers
If searching for microsatellite markers in specific regions of a genome, for example within a particularintron
An intron is any nucleotide sequence within a gene that is not expressed or operative in the final RNA product. The word ''intron'' is derived from the term ''intragenic region'', i.e. a region inside a gene."The notion of the cistron .e., gene. ...
, primers can be designed manually. This involves searching the genomic DNA sequence for microsatellite repeats, which can be done by eye or by using automated tools such arepeat masker
Once the potentially useful microsatellites are determined, the flanking sequences can be used to design
oligonucleotide
Oligonucleotides are short DNA or RNA molecules, oligomers, that have a wide range of applications in genetic testing, research, and forensics. Commonly made in the laboratory by solid-phase chemical synthesis, these small bits of nucleic acids c ...
primers which will amplify the specific microsatellite repeat in a PCR reaction.
Random microsatellite primers can be developed by cloning
Cloning is the process of producing individual organisms with identical or virtually identical DNA, either by natural or artificial means. In nature, some organisms produce clones through asexual reproduction. In the field of biotechnology, cl ...
random segments of DNA from the focal species. These random segments are inserted into a plasmid
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria; how ...
or bacteriophage
A bacteriophage (), also known informally as a ''phage'' (), is a duplodnaviria virus that infects and replicates within bacteria and archaea. The term was derived from "bacteria" and the Greek Žå╬▒╬│╬Ąß┐¢╬Į ('), meaning "to devour". Bacteri ...
vector
Vector most often refers to:
*Euclidean vector, a quantity with a magnitude and a direction
*Vector (epidemiology), an agent that carries and transmits an infectious pathogen into another living organism
Vector may also refer to:
Mathematic ...
, which is in turn implanted into ''Escherichia coli
''Escherichia coli'' (),Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. also known as ''E. coli'' (), is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus ''Escher ...
'' bacteria. Colonies are then developed, and screened with fluorescentlyŌĆōlabelled oligonucleotide
Oligonucleotides are short DNA or RNA molecules, oligomers, that have a wide range of applications in genetic testing, research, and forensics. Commonly made in the laboratory by solid-phase chemical synthesis, these small bits of nucleic acids c ...
sequences that will hybridize to a microsatellite repeat, if present on the DNA segment. If positive clones can be obtained from this procedure, the DNA is sequenced and PCR primers are chosen from sequences flanking such regions to determine a specific locus
Locus (plural loci) is Latin for "place". It may refer to:
Entertainment
* Locus (comics), a Marvel Comics mutant villainess, a member of the Mutant Liberation Front
* ''Locus'' (magazine), science fiction and fantasy magazine
** ''Locus Award' ...
. This process involves significant trial and error on the part of researchers, as microsatellite repeat sequences must be predicted and primers that are randomly isolated may not display significant polymorphism. Microsatellite loci are widely distributed throughout the genome and can be isolated from semi-degraded DNA of older specimens, as all that is needed is a suitable substrate for amplification through PCR.
More recent techniques involve using oligonucleotide
Oligonucleotides are short DNA or RNA molecules, oligomers, that have a wide range of applications in genetic testing, research, and forensics. Commonly made in the laboratory by solid-phase chemical synthesis, these small bits of nucleic acids c ...
sequences consisting of repeats complementary to repeats in the microsatellite to "enrich" the DNA extracted ( Microsatellite enrichment). The oligonucleotide probe hybridizes with the repeat in the microsatellite, and the probe/microsatellite complex is then pulled out of solution. The enriched DNA is then cloned as normal, but the proportion of successes will now be much higher, drastically reducing the time required to develop the regions for use. However, which probes to use can be a trial and error process in itself.
ISSR-PCR
ISSR (for inter-simple sequence repeat) is a general term for a genome region between microsatellite loci. The complementary sequences to two neighboring microsatellites are used as PCR primers; the variable region between them gets amplified. The limited length of amplification cycles during PCR prevents excessive replication of overly long contiguous DNA sequences, so the result will be a mix of a variety of amplified DNA strands which are generally short but vary much in length. Sequences amplified by ISSR-PCR can be used for DNA fingerprinting. Since an ISSR may be a conserved or nonconserved region, this technique is not useful for distinguishing individuals, but rather forphylogeography
Phylogeography is the study of the historical processes that may be responsible for the past to present geographic distributions of genealogical lineages. This is accomplished by considering the geographic distribution of individuals in light of ge ...
analyses or maybe delimiting species
In biology, a species is the basic unit of classification and a taxonomic rank of an organism, as well as a unit of biodiversity. A species is often defined as the largest group of organisms in which any two individuals of the appropriate s ...
; sequence diversity is lower than in SSR-PCR, but still higher than in actual gene sequences. In addition, microsatellite sequencing and ISSR sequencing are mutually assisting, as one produces primers for the other.
Limitations
Repetitive DNA is not easily analysed by next generation DNA sequencing methods, which struggle with homopolymeric tracts. Therefore, microsatellites are normally analysed by conventional PCR amplification and amplicon size determination. The use of PCR means that microsatellite length analysis is prone to PCR limitations like any other PCR-amplified DNA locus. A particular concern is the occurrence of ŌĆśnull allele
A null allele is a nonfunctional allele (a variant of a gene) caused by a genetic mutation. Such mutations can cause a complete lack of production of the associated gene product or a product that does not function properly; in either case, the alle ...
sŌĆÖ:
*Occasionally, within a sample of individuals such as in paternity testing casework, a mutation in the DNA flanking the microsatellite can prevent the PCR primer from binding and producing an amplicon (creating a "null allele" in a gel assay), thus only one allele is amplified (from the non-mutated sister chromosome), and the individual may then falsely appear to be homozygous. This can cause confusion in paternity casework. It may then be necessary to amplify the microsatellite using a different set of primers. Null alleles are caused especially by mutations at the 3ŌĆÖ section, where extension commences.
*In species or population analysis, for example in conservation work, PCR primers which amplify microsatellites in one individual or species can work in other species. However, the risk of applying PCR primers across different species is that null alleles become likely, whenever sequence divergence is too great for the primers to bind. The species may then artificially appear to have a reduced diversity. Null alleles in this case can sometimes be indicated by an excessive frequency of homozygotes causing deviations from Hardy-Weinberg equilibrium expectations.
See also
References
Further reading
* * * * * * * * * * *External links
All known disease-causing short tandem repeats
* Search tools: *
*
IMEx
*
Imperfect SSR Finder
Ćöfind perfect or imperfect SSRs in
FASTA
FASTA is a DNA and protein sequence alignment software package first described by David J. Lipman and William R. Pearson in 1985. Its legacy is the FASTA format which is now ubiquitous in bioinformatics.
History
The original FASTA program ...
sequences.
*JSTRINGŌĆöJava Search for Tandem Repeats In Genomes
*
Microsatellite repeats finder
*
*
MREPATT
*
Mreps
*
Ćöa tandem repeat search tool for perfect and imperfect repeatsŌĆöthe maximum pattern size depends only on computational power *
Poly
*
*
SSR Finder
*
STAR
*
*
*
TRED
*
TROLL
*
Zebrafish Repeats
{{Authority control Genetics Forensic genetics Repetitive DNA sequences