Replication origin regions on:
[Wikipedia]
[Google]
[Amazon]
 In
In


 DNA replication, like all biological polymerization processes, proceeds in three enzymatically catalyzed and coordinated steps: initiation, elongation and termination.
DNA replication, like all biological polymerization processes, proceeds in three enzymatically catalyzed and coordinated steps: initiation, elongation and termination.

 For a cell to divide, it must first replicate its DNA. DNA replication is an all-or-none process; once replication begins, it proceeds to completion. Once replication is complete, it does not occur again in the same cell cycle. This is made possible by the division of initiation of the
For a cell to divide, it must first replicate its DNA. DNA replication is an all-or-none process; once replication begins, it proceeds to completion. Once replication is complete, it does not occur again in the same cell cycle. This is made possible by the division of initiation of the

 The replication fork is a structure that forms within the long helical DNA during DNA replication. It is created by helicases, which break the hydrogen bonds holding the two DNA strands together in the helix. The resulting structure has two branching "prongs", each one made up of a single strand of DNA. These two strands serve as the template for the leading and lagging strands, which will be created as DNA polymerase matches complementary nucleotides to the templates; the templates may be properly referred to as the leading strand template and the lagging strand template.
DNA is read by DNA polymerase in the 3′ to 5′ direction, meaning the new strand is synthesized in the 5' to 3' direction. Since the leading and lagging strand templates are oriented in opposite directions at the replication fork, a major issue is how to achieve synthesis of new lagging strand DNA, whose direction of synthesis is opposite to the direction of the growing replication fork.
The replication fork is a structure that forms within the long helical DNA during DNA replication. It is created by helicases, which break the hydrogen bonds holding the two DNA strands together in the helix. The resulting structure has two branching "prongs", each one made up of a single strand of DNA. These two strands serve as the template for the leading and lagging strands, which will be created as DNA polymerase matches complementary nucleotides to the templates; the templates may be properly referred to as the leading strand template and the lagging strand template.
DNA is read by DNA polymerase in the 3′ to 5′ direction, meaning the new strand is synthesized in the 5' to 3' direction. Since the leading and lagging strand templates are oriented in opposite directions at the replication fork, a major issue is how to achieve synthesis of new lagging strand DNA, whose direction of synthesis is opposite to the direction of the growing replication fork.
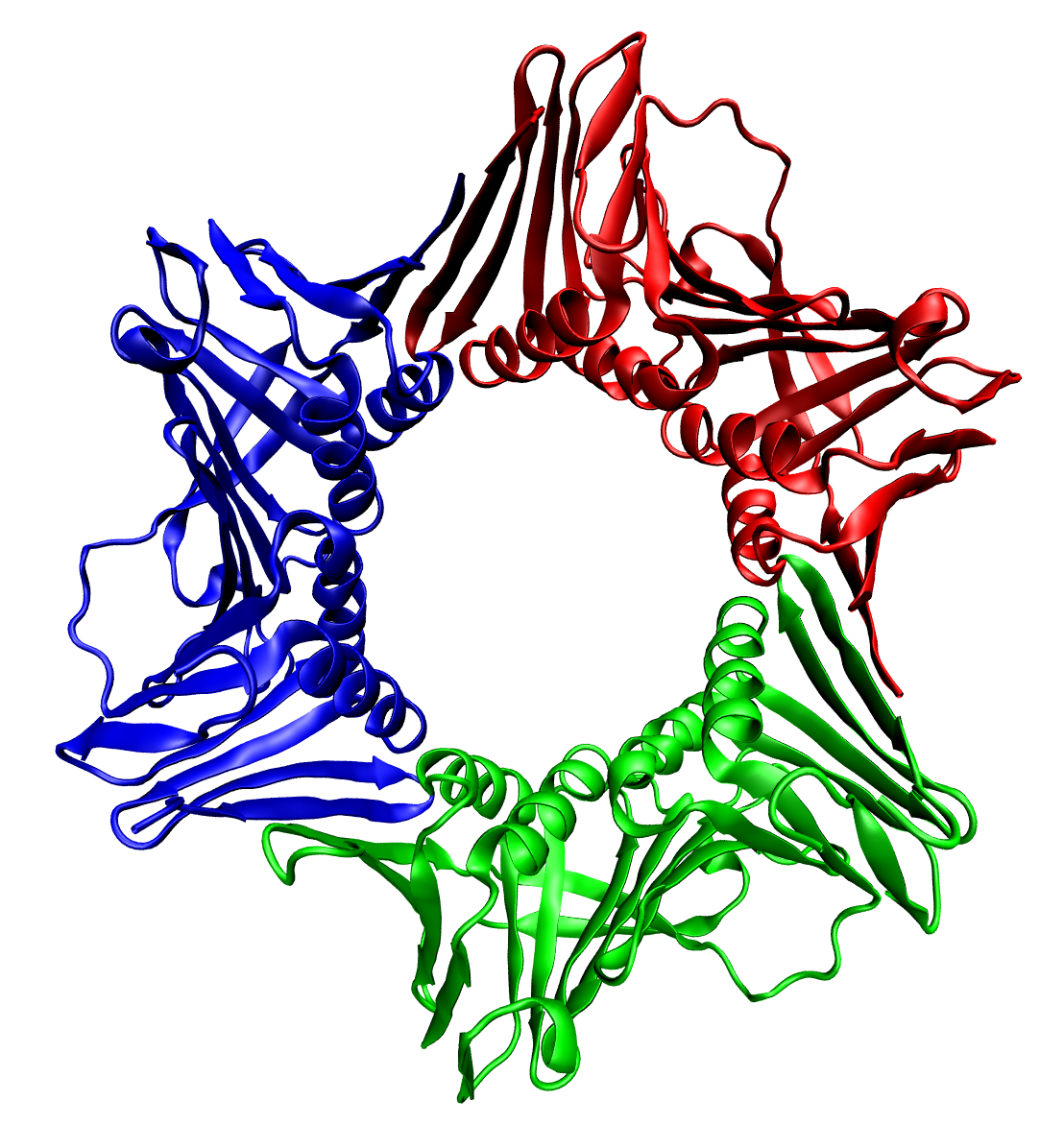 In all cases the helicase is composed of six polypeptides that wrap around only one strand of the DNA being replicated. The two polymerases are bound to the helicase heximer. In eukaryotes the helicase wraps around the leading strand, and in prokaryotes it wraps around the lagging strand.
As helicase unwinds DNA at the replication fork, the DNA ahead is forced to rotate. This process results in a build-up of twists in the DNA ahead. This build-up forms a torsional resistance that would eventually halt the progress of the replication fork. Topoisomerases are enzymes that temporarily break the strands of DNA, relieving the tension caused by unwinding the two strands of the DNA helix; topoisomerases (including DNA gyrase) achieve this by adding negative DNA supercoil, supercoils to the DNA helix.
Bare single-stranded DNA tends to fold back on itself forming Biomolecular structure#Secondary structure, secondary structures; these structures can interfere with the movement of DNA polymerase. To prevent this, single-strand binding proteins bind to the DNA until a second strand is synthesized, preventing secondary structure formation.
Double-stranded DNA is coiled around histones that play an important role in regulating gene expression so the replicated DNA must be coiled around histones at the same places as the original DNA. To ensure this, histone Chaperone (protein), chaperones disassemble the chromatin before it is replicated and replace the histones in the correct place. Some steps in this reassembly are somewhat speculative.
DNA clamp, Clamp proteins form a sliding clamp around DNA, helping the DNA polymerase maintain contact with its template, thereby assisting with processivity. The inner face of the clamp enables DNA to be threaded through it. Once the polymerase reaches the end of the template or detects double-stranded DNA, the sliding clamp undergoes a conformational change that releases the DNA polymerase. Clamp-loading proteins are used to initially load the clamp, recognizing the junction between template and RNA primers.:274-5
In all cases the helicase is composed of six polypeptides that wrap around only one strand of the DNA being replicated. The two polymerases are bound to the helicase heximer. In eukaryotes the helicase wraps around the leading strand, and in prokaryotes it wraps around the lagging strand.
As helicase unwinds DNA at the replication fork, the DNA ahead is forced to rotate. This process results in a build-up of twists in the DNA ahead. This build-up forms a torsional resistance that would eventually halt the progress of the replication fork. Topoisomerases are enzymes that temporarily break the strands of DNA, relieving the tension caused by unwinding the two strands of the DNA helix; topoisomerases (including DNA gyrase) achieve this by adding negative DNA supercoil, supercoils to the DNA helix.
Bare single-stranded DNA tends to fold back on itself forming Biomolecular structure#Secondary structure, secondary structures; these structures can interfere with the movement of DNA polymerase. To prevent this, single-strand binding proteins bind to the DNA until a second strand is synthesized, preventing secondary structure formation.
Double-stranded DNA is coiled around histones that play an important role in regulating gene expression so the replicated DNA must be coiled around histones at the same places as the original DNA. To ensure this, histone Chaperone (protein), chaperones disassemble the chromatin before it is replicated and replace the histones in the correct place. Some steps in this reassembly are somewhat speculative.
DNA clamp, Clamp proteins form a sliding clamp around DNA, helping the DNA polymerase maintain contact with its template, thereby assisting with processivity. The inner face of the clamp enables DNA to be threaded through it. Once the polymerase reaches the end of the template or detects double-stranded DNA, the sliding clamp undergoes a conformational change that releases the DNA polymerase. Clamp-loading proteins are used to initially load the clamp, recognizing the junction between template and RNA primers.:274-5
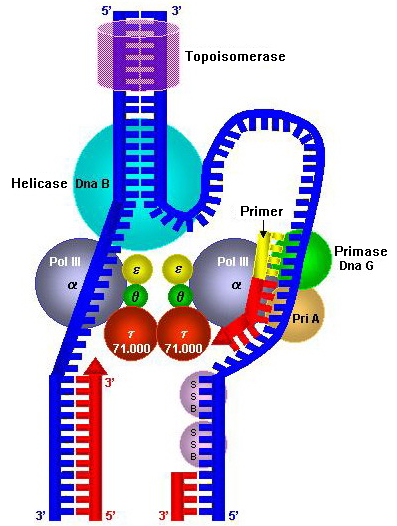 Replication machineries consist of factors involved in DNA replication and appearing on template ssDNAs. Replication machineries include primosotors are replication enzymes; DNA polymerase, DNA helicases, DNA clamps and DNA topoisomerases, and replication proteins; e.g. single-stranded DNA binding proteins (SSB). In the replication machineries these components coordinate. In most of the bacteria, all of the factors involved in DNA replication are located on replication forks and the complexes stay on the forks during DNA replication. These replication machineries are called replisomes or DNA replicase systems. These terms are generic terms for proteins located on replication forks. In eukaryotic and some bacterial cells the replisomes are not formed.
Since replication machineries do not move relatively to template DNAs such as factories, they are called a replication factory. In an alternative figure, DNA factories are similar to projectors and DNAs are like as cinematic films passing constantly into the projectors. In the replication factory model, after both DNA helicases for leading strands and lagging strands are loaded on the template DNAs, the helicases run along the DNAs into each other. The helicases remain associated for the remainder of replication process. Peter Meister et al. observed directly replication sites in
Replication machineries consist of factors involved in DNA replication and appearing on template ssDNAs. Replication machineries include primosotors are replication enzymes; DNA polymerase, DNA helicases, DNA clamps and DNA topoisomerases, and replication proteins; e.g. single-stranded DNA binding proteins (SSB). In the replication machineries these components coordinate. In most of the bacteria, all of the factors involved in DNA replication are located on replication forks and the complexes stay on the forks during DNA replication. These replication machineries are called replisomes or DNA replicase systems. These terms are generic terms for proteins located on replication forks. In eukaryotic and some bacterial cells the replisomes are not formed.
Since replication machineries do not move relatively to template DNAs such as factories, they are called a replication factory. In an alternative figure, DNA factories are similar to projectors and DNAs are like as cinematic films passing constantly into the projectors. In the replication factory model, after both DNA helicases for leading strands and lagging strands are loaded on the template DNAs, the helicases run along the DNAs into each other. The helicases remain associated for the remainder of replication process. Peter Meister et al. observed directly replication sites in

 Most bacteria do not go through a well-defined cell cycle but instead continuously copy their DNA; during rapid growth, this can result in the concurrent occurrence of multiple rounds of replication. In ''E. coli'', the best-characterized bacteria, DNA replication is regulated through several mechanisms, including: the hemimethylation and sequestering of the origin sequence, the ratio of Adenosine triphosphate, adenosine triphosphate (ATP) to Adenosine diphosphate, adenosine diphosphate (ADP), and the levels of protein DnaA. All these control the binding of initiator proteins to the origin sequences.
Because ''E. coli'' DNA methylation, methylates GATC DNA sequences, DNA synthesis results in hemimethylated sequences. This hemimethylated DNA is recognized by the protein SeqA protein domain, SeqA, which binds and sequesters the origin sequence; in addition, DnaA (required for initiation of replication) binds less well to hemimethylated DNA. As a result, newly replicated origins are prevented from immediately initiating another round of DNA replication.
ATP builds up when the cell is in a rich medium, triggering DNA replication once the cell has reached a specific size. ATP competes with ADP to bind to DnaA, and the DnaA-ATP complex is able to initiate replication. A certain number of DnaA proteins are also required for DNA replication — each time the origin is copied, the number of binding sites for DnaA doubles, requiring the synthesis of more DnaA to enable another initiation of replication.
In fast-growing bacteria, such as ''E. coli'', chromosome replication takes more time than dividing the cell. The bacteria solve this by initiating a new round of replication before the previous one has been terminated. The new round of replication will form the chromosome of the cell that is born two generations after the dividing cell. This mechanism creates overlapping replication cycles.
Most bacteria do not go through a well-defined cell cycle but instead continuously copy their DNA; during rapid growth, this can result in the concurrent occurrence of multiple rounds of replication. In ''E. coli'', the best-characterized bacteria, DNA replication is regulated through several mechanisms, including: the hemimethylation and sequestering of the origin sequence, the ratio of Adenosine triphosphate, adenosine triphosphate (ATP) to Adenosine diphosphate, adenosine diphosphate (ADP), and the levels of protein DnaA. All these control the binding of initiator proteins to the origin sequences.
Because ''E. coli'' DNA methylation, methylates GATC DNA sequences, DNA synthesis results in hemimethylated sequences. This hemimethylated DNA is recognized by the protein SeqA protein domain, SeqA, which binds and sequesters the origin sequence; in addition, DnaA (required for initiation of replication) binds less well to hemimethylated DNA. As a result, newly replicated origins are prevented from immediately initiating another round of DNA replication.
ATP builds up when the cell is in a rich medium, triggering DNA replication once the cell has reached a specific size. ATP competes with ADP to bind to DnaA, and the DnaA-ATP complex is able to initiate replication. A certain number of DnaA proteins are also required for DNA replication — each time the origin is copied, the number of binding sites for DnaA doubles, requiring the synthesis of more DnaA to enable another initiation of replication.
In fast-growing bacteria, such as ''E. coli'', chromosome replication takes more time than dividing the cell. The bacteria solve this by initiating a new round of replication before the previous one has been terminated. The new round of replication will form the chromosome of the cell that is born two generations after the dividing cell. This mechanism creates overlapping replication cycles.

 There are many events that contribute to replication stress, including:
* Misincorporation of ribonucleotides
* Unusual Nucleic acid secondary structure, DNA structures
* Conflicts between replication and transcription
* Insufficiency of essential replication factors
* Chromosomal fragile site, Common fragile sites
* Overexpression or constitutive activation of oncogenes
* Chromatin inaccessibility
There are many events that contribute to replication stress, including:
* Misincorporation of ribonucleotides
* Unusual Nucleic acid secondary structure, DNA structures
* Conflicts between replication and transcription
* Insufficiency of essential replication factors
* Chromosomal fragile site, Common fragile sites
* Overexpression or constitutive activation of oncogenes
* Chromatin inaccessibility
molecular biology
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions. The study of chemical and physi ...
, DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule. DNA replication occurs in all living organisms
In biology, an organism () is any living system that functions as an individual entity. All organisms are composed of cells (cell theory). Organisms are classified by taxonomy into groups such as multicellular animals, plants, and fun ...
acting as the most essential part for biological inheritance
Heredity, also called inheritance or biological inheritance, is the passing on of traits from parents to their offspring; either through asexual reproduction or sexual reproduction, the offspring cells or organisms acquire the genetic informa ...
. This is essential for cell division during growth and repair of damaged tissues, while it also ensures that each of the new cells receives its own copy of the DNA. The cell possesses the distinctive property of division, which makes replication of DNA essential.
DNA is made up of a double helix
A double is a look-alike or doppelgänger; one person or being that resembles another.
Double, The Double or Dubble may also refer to:
Film and television
* Double (filmmaking), someone who substitutes for the credited actor of a character
* ...
of two complementary strands. The double helix describes the appearance of a double-stranded DNA which is thus composed of two linear strands that run opposite to each other and twist together to form. During replication, these strands are separated. Each strand of the original DNA molecule then serves as a template for the production of its counterpart, a process referred to as semiconservative replication
Semiconservative replication describe the mechanism of DNA replication in all known cells. DNA replication occurs on multiple origins of replication along the DNA template strand (antinsense strand). As the DNA double helix is unwound by helicase, ...
. As a result of semi-conservative replication, the new helix will be composed of an original DNA strand as well as a newly synthesized strand. Cellular proofreading and error-checking mechanisms ensure near perfect fidelity
Fidelity is the quality of faithfulness or loyalty. Its original meaning regarded duty in a broader sense than the related concept of ''fealty''. Both derive from the Latin word ''fidēlis'', meaning "faithful or loyal". In the City of London fin ...
for DNA replication.Imperfect DNA replication results in mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA replication, DNA or viral repl ...
s.
In a cell
Cell most often refers to:
* Cell (biology), the functional basic unit of life
Cell may also refer to:
Locations
* Monastic cell, a small room, hut, or cave in which a religious recluse lives, alternatively the small precursor of a monastery ...
, DNA replication begins at specific locations, or origins of replication, in the genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding g ...
which contains the genetic material of an organism. Unwinding of DNA at the origin and synthesis of new strands, accommodated by an enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products ...
known as helicase
Helicases are a class of enzymes thought to be vital to all organisms. Their main function is to unpack an organism's genetic material. Helicases are motor proteins that move directionally along a nucleic acid phosphodiester backbone, separatin ...
, results in replication fork
In molecular biology, DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule. DNA replication occurs in all living organisms acting as the most essential part for biological inheritanc ...
s growing bi-directionally from the origin. A number of protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, res ...
s are associated with the replication fork to help in the initiation and continuation of DNA synthesis
DNA synthesis is the natural or artificial creation of deoxyribonucleic acid (DNA) molecules. DNA is a macromolecule made up of nucleotide units, which are linked by covalent bonds and hydrogen bonds, in a repeating structure. DNA synthesis occurs ...
. Most prominently, DNA polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create ...
synthesizes the new strands by adding nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecule ...
s that complement each (template) strand. DNA replication occurs during the S-stage of interphase
Interphase is the portion of the cell cycle that is not accompanied by visible changes under the microscope, and includes the G1, S and G2 phases. During interphase, the cell grows (G1), replicates its DNA (S) and prepares for mitosis (G2). A c ...
.
DNA replication (DNA amplification) can also be performed ''in vitro
''In vitro'' (meaning in glass, or ''in the glass'') studies are performed with microorganisms, cells, or biological molecules outside their normal biological context. Colloquially called " test-tube experiments", these studies in biology ...
'' (artificially, outside a cell). DNA polymerases isolated from cells and artificial DNA primers can be used to start DNA synthesis at known sequences in a template DNA molecule. Polymerase chain reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) ...
(PCR), ligase chain reaction The ligase chain reaction (LCR) is a method of DNA amplification. The ligase chain reaction (LCR) is an amplification process that differs from PCR in that it involves a thermostable ligase to join two probes or other molecules together which can t ...
(LCR), and transcription-mediated amplification
Transcription-mediated amplification (TMA) is an isothermal (does not change the nucleic acid temperature), single-tube nucleic acid amplification system utilizing two enzymes, RNA polymerase and reverse transcriptase.
"Amplification" means crea ...
(TMA) are examples. In March 2021, researchers reported evidence suggesting that a preliminary form of transfer RNA, a necessary component of translation
Translation is the communication of the meaning of a source-language text by means of an equivalent target-language text. The English language draws a terminological distinction (which does not exist in every language) between ''transla ...
, the biological synthesis of new protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, res ...
s in accordance with the genetic code
The genetic code is the set of rules used by living cells to translate information encoded within genetic material ( DNA or RNA sequences of nucleotide triplets, or codons) into proteins. Translation is accomplished by the ribosome, which links ...
, could have been a replicator molecule itself in the very early development of life, or abiogenesis
In biology, abiogenesis (from a- 'not' + Greek bios 'life' + genesis 'origin') or the origin of life is the natural process by which life has arisen from non-living matter, such as simple organic compounds. The prevailing scientific hypothes ...
.
DNA structure
DNA exists as a double-stranded structure, with both strands coiled together to form the characteristic double-helix. Each single strand of DNA is a chain of four types ofnucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecule ...
s. Nucleotides in DNA contain a deoxyribose
Deoxyribose, or more precisely 2-deoxyribose, is a monosaccharide with idealized formula H−(C=O)−(CH2)−(CHOH)3−H. Its name indicates that it is a deoxy sugar, meaning that it is derived from the sugar ribose by loss of a hydroxy group. D ...
sugar, a phosphate
In chemistry, a phosphate is an anion, salt, functional group or ester derived from a phosphoric acid. It most commonly means orthophosphate, a derivative of orthophosphoric acid .
The phosphate or orthophosphate ion is derived from phosph ...
, and a nucleobase
Nucleobases, also known as ''nitrogenous bases'' or often simply ''bases'', are nitrogen-containing biological compounds that form nucleosides, which, in turn, are components of nucleotides, with all of these monomers constituting the basic b ...
. The four types of nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecule ...
correspond to the four nucleobase
Nucleobases, also known as ''nitrogenous bases'' or often simply ''bases'', are nitrogen-containing biological compounds that form nucleosides, which, in turn, are components of nucleotides, with all of these monomers constituting the basic b ...
s adenine
Adenine () ( symbol A or Ade) is a nucleobase (a purine derivative). It is one of the four nucleobases in the nucleic acid of DNA that are represented by the letters G–C–A–T. The three others are guanine, cytosine and thymine. Its deri ...
, cytosine
Cytosine () ( symbol C or Cyt) is one of the four nucleobases found in DNA and RNA, along with adenine, guanine, and thymine (uracil in RNA). It is a pyrimidine derivative, with a heterocyclic aromatic ring and two substituents attached (an ...
, guanine
Guanine () ( symbol G or Gua) is one of the four main nucleobases found in the nucleic acids DNA and RNA, the others being adenine, cytosine, and thymine (uracil in RNA). In DNA, guanine is paired with cytosine. The guanine nucleoside is c ...
, and thymine
Thymine () ( symbol T or Thy) is one of the four nucleobases in the nucleic acid of DNA that are represented by the letters G–C–A–T. The others are adenine, guanine, and cytosine. Thymine is also known as 5-methyluracil, a pyrimidi ...
, commonly abbreviated as A, C, G and T. Adenine and guanine are purine
Purine is a heterocyclic aromatic organic compound that consists of two rings ( pyrimidine and imidazole) fused together. It is water-soluble. Purine also gives its name to the wider class of molecules, purines, which include substituted purines ...
bases, while cytosine and thymine are pyrimidines. These nucleotides form phosphodiester bonds
In chemistry, a phosphodiester bond occurs when exactly two of the hydroxyl groups () in phosphoric acid react with hydroxyl groups on other molecules to form two ester bonds. The "bond" involves this linkage . Discussion of phosphodiesters is ...
, creating the phosphate-deoxyribose backbone of the DNA double helix with the nucleobases pointing inward (i.e., toward the opposing strand). Nucleobases are matched between strands through hydrogen bonds to form base pairs. Adenine pairs with thymine (two hydrogen bonds), and guanine pairs with cytosine (three hydrogen bonds).
DNA strands have a directionality, and the different ends of a single strand are called the "3′ (three-prime) end" and the "5′ (five-prime) end". By convention, if the base sequence of a single strand of DNA is given, the left end of the sequence is the 5′ end, while the right end of the sequence is the 3′ end. The strands of the double helix are anti-parallel with one being 5′ to 3′, and the opposite strand 3′ to 5′. These terms refer to the carbon atom in deoxyribose to which the next phosphate in the chain attaches. Directionality has consequences in DNA synthesis, because DNA polymerase can synthesize DNA in only one direction by adding nucleotides to the 3′ end of a DNA strand.
The pairing of complementary bases in DNA (through hydrogen bonding
In chemistry, a hydrogen bond (or H-bond) is a primarily electrostatic force of attraction between a hydrogen (H) atom which is covalently bound to a more electronegative "donor" atom or group (Dn), and another electronegative atom bearing a l ...
) means that the information contained within each strand is redundant. Phosphodiester (intra-strand) bonds are stronger than hydrogen (inter-strand) bonds. The actual job of the phosphodiester bonds is where in DNA polymers connect the 5' carbon atom of one nucleotide to the 3' carbon atom of another nucleotide, while the hydrogen bonds stabilize DNA double helices across the helix axis but not in the direction of the axis 1. This allows the strands to be separated from one another. The nucleotides on a single strand can therefore be used to reconstruct nucleotides on a newly synthesized partner strand.
DNA polymerase
DNA polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create ...
s are a family of enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products ...
s that carry out all forms of DNA replication. DNA polymerases in general cannot initiate synthesis of new strands, but can only extend an existing DNA or RNA strand paired with a template strand. To begin synthesis, a short fragment of RNA, called a primer, must be created and paired with the template DNA strand.
DNA polymerase adds a new strand of DNA by extending the 3′ end of an existing nucleotide chain, adding new nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecule ...
s matched to the template strand one at a time via the creation of phosphodiester bond
In chemistry, a phosphodiester bond occurs when exactly two of the hydroxyl groups () in phosphoric acid react with hydroxyl groups on other molecules to form two ester bonds. The "bond" involves this linkage . Discussion of phosphodiesters is d ...
s. The energy for this process of DNA polymerization comes from hydrolysis of the high-energy phosphate
High-energy phosphate can mean one of two things:
* The phosphate-phosphate (phosphoanhydride/phosphoric anhydride/macroergic/phosphagen) bonds formed when compounds such as adenosine diphosphate (ADP) and adenosine triphosphate (ATP) are created ...
(phosphoanhydride) bonds between the three phosphates attached to each unincorporated base. Free bases with their attached phosphate groups are called nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecule ...
s; in particular, bases with three attached phosphate groups are called nucleoside triphosphate
A nucleoside triphosphate is a nucleoside containing a nitrogenous base bound to a 5-carbon sugar (either ribose or deoxyribose), with three phosphate groups bound to the sugar. They are the molecular precursors of both DNA and RNA, which are ch ...
s. When a nucleotide is being added to a growing DNA strand, the formation of a phosphodiester bond between the proximal phosphate of the nucleotide to the growing chain is accompanied by hydrolysis of a high-energy phosphate bond with release of the two distal phosphate groups as a pyrophosphate
In chemistry, pyrophosphates are phosphorus oxyanions that contain two phosphorus atoms in a P–O–P linkage. A number of pyrophosphate salts exist, such as disodium pyrophosphate (Na2H2P2O7) and tetrasodium pyrophosphate (Na4P2O7), among othe ...
. Enzymatic hydrolysis of the resulting pyrophosphate
In chemistry, pyrophosphates are phosphorus oxyanions that contain two phosphorus atoms in a P–O–P linkage. A number of pyrophosphate salts exist, such as disodium pyrophosphate (Na2H2P2O7) and tetrasodium pyrophosphate (Na4P2O7), among othe ...
into inorganic phosphate consumes a second high-energy phosphate bond and renders the reaction effectively irreversible.The energetics of this process may also help explain the directionality of synthesis—if DNA were synthesized in the 3′ to 5′ direction, the energy for the process would come from the 5′ end of the growing strand rather than from free nucleotides. The problem is that if the high energy triphosphates were on the growing strand and not on the free nucleotides, proof-reading by removing a mismatched terminal nucleotide would be problematic: Once a nucleotide is added, the triphosphate is lost and a single phosphate remains on the backbone between the new nucleotide and the rest of the strand. If the added nucleotide were mismatched, removal would result in a DNA strand terminated by a monophosphate at the end of the "growing strand" rather than a high energy triphosphate. So strand would be stuck and wouldn't be able to grow anymore. In actuality, the high energy triphosphates hydrolyzed at each step originate from the free nucleotides, not the polymerized strand, so this issue does not exist.
In general, DNA polymerases are highly accurate, with an intrinsic error rate of less than one mistake for every 107 nucleotides added. In addition, some DNA polymerases also have proofreading ability; they can remove nucleotides from the end of a growing strand in order to correct mismatched bases. Finally, post-replication mismatch repair mechanisms monitor the DNA for errors, being capable of distinguishing mismatches in the newly synthesized DNA strand from the original strand sequence. Together, these three discrimination steps enable replication fidelity of less than one mistake for every 109 nucleotides added.
The rate of DNA replication in a living cell was first measured as the rate of phage T4 DNA elongation in phage-infected ''E. coli''. During the period of exponential DNA increase at 37 °C, the rate was 749 nucleotides per second. The mutation rate per base pair per replication during phage T4 DNA synthesis is 1.7 per 108.
Replication process
Initiation

 For a cell to divide, it must first replicate its DNA. DNA replication is an all-or-none process; once replication begins, it proceeds to completion. Once replication is complete, it does not occur again in the same cell cycle. This is made possible by the division of initiation of the
For a cell to divide, it must first replicate its DNA. DNA replication is an all-or-none process; once replication begins, it proceeds to completion. Once replication is complete, it does not occur again in the same cell cycle. This is made possible by the division of initiation of the pre-replication complex
A pre-replication complex (pre-RC) is a protein complex that forms at the origin of replication during the initiation step of DNA replication. Formation of the pre-RC is required for DNA replication to occur. Complete and faithful replication of ...
.
Pre-replication complex
In late mitosis and earlyG1 phase
The G1 phase, gap 1 phase, or growth 1 phase, is the first of four phases of the cell cycle that takes place in eukaryotic cell division. In this part of interphase, the cell synthesizes mRNA and proteins in preparation for subsequent steps lead ...
, a large complex of initiator proteins assembles into the pre-replication complex at particular points in the DNA, known as "origins
Origin(s) or The Origin may refer to:
Arts, entertainment, and media
Comics and manga
* ''Origin'' (comics), a Wolverine comic book mini-series published by Marvel Comics in 2002
* ''The Origin'' (Buffy comic), a 1999 ''Buffy the Vampire Sl ...
". In '' E. coli'' the primary initiator protein is DnaA
Introduction
Based on the Replicon Model, a positively active initiator molecule contacts with a particular spot on a circular chromosome called the replicator to start DNA replication. DnaA is a protein that activates initiation of DNA replica ...
; in yeast
Yeasts are eukaryotic, single-celled microorganisms classified as members of the fungus kingdom. The first yeast originated hundreds of millions of years ago, and at least 1,500 species are currently recognized. They are estimated to constit ...
, this is the origin recognition complex
In molecular biology, origin recognition complex (ORC) is a multi-subunit DNA binding complex (6 subunits) that binds in all eukaryotes and archaea in an ATP-dependent manner to origins of replication. The subunits of this complex are encoded ...
. Sequences used by initiator proteins tend to be "AT-rich" (rich in adenine and thymine bases), because A-T base pairs have two hydrogen bonds (rather than the three formed in a C-G pair) and thus are easier to strand-separate. In eukaryotes, the origin recognition complex catalyzes the assembly of initiator proteins into the pre-replication complex. In addition, a recent report suggests that budding yeast ORC dimerizes in a cell cycle dependent manner to control licensing. ORC and Noc3p are continuously bound to the chromatin throughout the cell cycle. Cdc6
Cell division control protein 6 homolog is a protein that in humans is encoded by the ''CDC6'' gene.
The protein encoded by this gene is highly similar to Saccharomyces cerevisiae Cdc6, a protein essential for the initiation of DNA replication. ...
and Cdt1
CDT1 (Chromatin licensing and DNA replication factor 1) is a protein that in humans is encoded by the ''CDT1'' gene. It is a licensing factor that functions to limit DNA from replicating more than once per cell cycle.
Role in pre-replication com ...
then associate with the bound origin recognition complex at the origin in order to form a larger complex necessary to load the Mcm complex onto the DNA. The Mcm complex is the helicase that will unravel the DNA helix at the replication origins and replication fork
In molecular biology, DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule. DNA replication occurs in all living organisms acting as the most essential part for biological inheritanc ...
s in eukaryotes. The Mcm complex is recruited at late G1 phase and loaded by the ORC-Cdc6-Cdt1 complex onto the DNA via ATP-dependent protein remodeling. The loading of the Mcm complex onto the origin DNA marks the completion of pre-replication complex formation.
If environmental conditions are right in late G1 phase, the G1 and G1/S cyclin- Cdk complexes are activated, which stimulate expression of genes that encode components of the DNA synthetic machinery. G1/S-Cdk activation also promotes the expression and activation of S-Cdk complexes, which may play a role in activating replication origins depending on species and cell type. Control of these Cdks vary depending on cell type and stage of development. This regulation is best understood in budding yeast
''Saccharomyces cerevisiae'' () (brewer's yeast or baker's yeast) is a species of yeast (single-celled fungus microorganisms). The species has been instrumental in winemaking, baking, and brewing since ancient times. It is believed to have been ...
, where the S cyclins Clb5 and Clb6 are primarily responsible for DNA replication. Clb5,6-Cdk1 complexes directly trigger the activation of replication origins and are therefore required throughout S phase to directly activate each origin.
In a similar manner, Cdc7 is also required through S phase to activate replication origins. Cdc7 is not active throughout the cell cycle, and its activation is strictly timed to avoid premature initiation of DNA replication. In late G1, Cdc7 activity rises abruptly as a result of association with the regulatory subunit DBF4
Protein DBF4 homolog A is a protein that is encoded by the ''DBF4'' gene in humans.
Interactions
DBF4 has been shown to interact with:
* Cell division cycle 7-related protein kinase,
* MCM3,
* MCM7,
* ORC2L, and
* ORC6L
Origin recogni ...
, which binds Cdc7 directly and promotes its protein kinase activity. Cdc7 has been found to be a rate-limiting regulator of origin activity. Together, the G1/S-Cdks and/or S-Cdks and Cdc7 collaborate to directly activate the replication origins, leading to initiation of DNA synthesis.
Preinitiation complex
In early S phase, S-Cdk and Cdc7 activation lead to the assembly of the preinitiation complex, a massive protein complex formed at the origin. Formation of the preinitiation complex displaces Cdc6 and Cdt1 from the origin replication complex, inactivating and disassembling the pre-replication complex. Loading the preinitiation complex onto the origin activates the Mcm helicase, causing unwinding of the DNA helix. The preinitiation complex also loads α-primase and other DNA polymerases onto the DNA. After α-primase synthesizes the first primers, the primer-template junctions interact with the clamp loader, which loads the sliding clamp onto the DNA to begin DNA synthesis. The components of the preinitiation complex remain associated with replication forks as they move out from the origin.Elongation
DNA polymerase has 5′–3′ activity. All known DNA replication systems require a free 3′hydroxyl
In chemistry, a hydroxy or hydroxyl group is a functional group with the chemical formula and composed of one oxygen atom covalently bonded to one hydrogen atom. In organic chemistry, alcohols and carboxylic acids contain one or more hydro ...
group before synthesis can be initiated (note: the DNA template is read in 3′ to 5′ direction whereas a new strand is synthesized in the 5′ to 3′ direction—this is often confused). Four distinct mechanisms for DNA synthesis are recognized:
# All cellular life forms and many DNA virus
A virus is a submicroscopic infectious agent that replicates only inside the living cells of an organism. Viruses infect all life forms, from animals and plants to microorganisms, including bacteria and archaea.
Since Dmitri Ivanovsk ...
es, phage
A bacteriophage (), also known informally as a ''phage'' (), is a duplodnaviria virus that infects and replicates within bacteria and archaea. The term was derived from "bacteria" and the Greek φαγεῖν ('), meaning "to devour". Bacter ...
s and plasmids use a primase
DNA primase is an enzyme involved in the replication of DNA and is a type of RNA polymerase. Primase catalyzes the synthesis of a short RNA (or DNA in some
living organisms) segment called a primer complementary to a ssDNA (single-stranded ...
to synthesize a short RNA primer with a free 3′ OH group which is subsequently elongated by a DNA polymerase.
# The retroelements (including retroviruses) employ a transfer RNA that primes DNA replication by providing a free 3′ OH that is used for elongation by the reverse transcriptase.
# In the adenoviruses and the φ29 family of bacteriophages, the 3′ OH group is provided by the side chain of an amino acid of the genome attached protein (the terminal protein) to which nucleotides are added by the DNA polymerase to form a new strand.
# In the single stranded DNA viruses—a group that includes the circoviruses, the geminiviruses, the parvovirus
Parvoviruses are a family of animal viruses that constitute the family ''Parvoviridae''. They have linear, single-stranded DNA (ssDNA) genomes that typically contain two genes encoding for a replication initiator protein, called NS1, and the p ...
es and others—and also the many phages and plasmids that use the rolling circle replication (RCR) mechanism, the RCR endonuclease creates a nick in the genome strand (single stranded viruses) or one of the DNA strands (plasmids). The 5′ end of the nicked strand is transferred to a tyrosine
-Tyrosine or tyrosine (symbol Tyr or Y) or 4-hydroxyphenylalanine is one of the 20 standard amino acids that are used by cells to synthesize proteins. It is a non-essential amino acid with a polar side group. The word "tyrosine" is from the G ...
residue on the nuclease and the free 3′ OH group is then used by the DNA polymerase to synthesize the new strand.
The first is the best known of these mechanisms and is used by the cellular organisms. In this mechanism, once the two strands are separated, primase
DNA primase is an enzyme involved in the replication of DNA and is a type of RNA polymerase. Primase catalyzes the synthesis of a short RNA (or DNA in some
living organisms) segment called a primer complementary to a ssDNA (single-stranded ...
adds RNA primers to the template strands. The leading strand receives one RNA primer while the lagging strand receives several. The leading strand is continuously extended from the primer by a DNA polymerase with high processivity
In molecular biology and biochemistry, processivity is an enzyme's ability to catalyze "consecutive reactions without releasing its substrate".
For example, processivity is the average number of nucleotides added by a polymerase enzyme, such as ...
, while the lagging strand is extended discontinuously from each primer forming Okazaki fragments
Okazaki fragments are short sequences of DNA nucleotides (approximately 150 to 200 base pairs long in eukaryotes) which are synthesized discontinuously and later linked together by the enzyme DNA ligase to create the lagging strand during DNA ...
. RNase
Ribonuclease (commonly abbreviated RNase) is a type of nuclease that catalyzes the degradation of RNA into smaller components. Ribonucleases can be divided into endoribonucleases and exoribonucleases, and comprise several sub-classes within t ...
removes the primer RNA fragments, and a low processivity DNA polymerase distinct from the replicative polymerase enters to fill the gaps. When this is complete, a single nick on the leading strand and several nicks on the lagging strand can be found. Ligase works to fill these nicks in, thus completing the newly replicated DNA molecule.
The primase used in this process differs significantly between bacteria and archaea/eukaryotes. Bacteria use a primase belonging to the DnaG protein superfamily which contains a catalytic domain of the TOPRIM fold type. The TOPRIM fold contains an α/β core with four conserved strands in a Rossmann fold, Rossmann-like topology. This structure is also found in the catalytic domains of topoisomerase Ia, topoisomerase II, the OLD-family nucleases and DNA repair proteins related to the RecR protein.
The primase used by archaea and eukaryotes, in contrast, contains a highly derived version of the RNA recognition motif (RRM). This primase is structurally similar to many viral RNA-dependent RNA polymerases, reverse transcriptases, cyclic nucleotide generating cyclases and DNA polymerases of the A/B/Y families that are involved in DNA replication and repair. In eukaryotic replication, the primase forms a complex with Pol α.
Multiple DNA polymerases take on different roles in the DNA replication process. In '' E. coli'', Pol III, DNA Pol III is the polymerase enzyme primarily responsible for DNA replication. It assembles into a replication complex at the replication fork that exhibits extremely high processivity, remaining intact for the entire replication cycle. In contrast, Pol I, DNA Pol I is the enzyme responsible for replacing RNA primers with DNA. DNA Pol I has a 5′ to 3′ exonuclease activity in addition to its polymerase activity, and uses its exonuclease activity to degrade the RNA primers ahead of it as it extends the DNA strand behind it, in a process called nick translation. Pol I is much less processive than Pol III because its primary function in DNA replication is to create many short DNA regions rather than a few very long regions.
In eukaryotes, the low-processivity enzyme, Pol α, helps to initiate replication because it forms a complex with primase. In eukaryotes, leading strand synthesis is thought to be conducted by Pol ε; however, this view has recently been challenged, suggesting a role for Pol δ. Primer removal is completed Pol δ while repair of DNA during replication is completed by Pol ε.
As DNA synthesis continues, the original DNA strands continue to unwind on each side of the bubble, forming a replication fork
In molecular biology, DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule. DNA replication occurs in all living organisms acting as the most essential part for biological inheritanc ...
with two prongs. In bacteria, which have a single origin of replication on their circular chromosome, this process creates a "theta structure" (resembling the Greek letter theta: θ). In contrast, eukaryotes have longer linear chromosomes and initiate replication at multiple origins within these.
Replication fork
Leading strand
The leading strand is the strand of new DNA which is synthesized in the same direction as the growing replication fork. This sort of DNA replication is continuous.Lagging strand
The lagging strand is the strand of new DNA whose direction of synthesis is opposite to the direction of the growing replication fork. Because of its orientation, replication of the lagging strand is more complicated as compared to that of the leading strand. As a consequence, the DNA polymerase on this strand is seen to "lag behind" the other strand. The lagging strand is synthesized in short, separated segments. On the lagging strand ''template'', aprimase
DNA primase is an enzyme involved in the replication of DNA and is a type of RNA polymerase. Primase catalyzes the synthesis of a short RNA (or DNA in some
living organisms) segment called a primer complementary to a ssDNA (single-stranded ...
"reads" the template DNA and initiates synthesis of a short complementary RNA primer. A DNA polymerase extends the primed segments, forming Okazaki fragments. The RNA primers are then removed and replaced with DNA, and the fragments of DNA are joined by DNA ligase.
Dynamics at the replication fork
 In all cases the helicase is composed of six polypeptides that wrap around only one strand of the DNA being replicated. The two polymerases are bound to the helicase heximer. In eukaryotes the helicase wraps around the leading strand, and in prokaryotes it wraps around the lagging strand.
As helicase unwinds DNA at the replication fork, the DNA ahead is forced to rotate. This process results in a build-up of twists in the DNA ahead. This build-up forms a torsional resistance that would eventually halt the progress of the replication fork. Topoisomerases are enzymes that temporarily break the strands of DNA, relieving the tension caused by unwinding the two strands of the DNA helix; topoisomerases (including DNA gyrase) achieve this by adding negative DNA supercoil, supercoils to the DNA helix.
Bare single-stranded DNA tends to fold back on itself forming Biomolecular structure#Secondary structure, secondary structures; these structures can interfere with the movement of DNA polymerase. To prevent this, single-strand binding proteins bind to the DNA until a second strand is synthesized, preventing secondary structure formation.
Double-stranded DNA is coiled around histones that play an important role in regulating gene expression so the replicated DNA must be coiled around histones at the same places as the original DNA. To ensure this, histone Chaperone (protein), chaperones disassemble the chromatin before it is replicated and replace the histones in the correct place. Some steps in this reassembly are somewhat speculative.
DNA clamp, Clamp proteins form a sliding clamp around DNA, helping the DNA polymerase maintain contact with its template, thereby assisting with processivity. The inner face of the clamp enables DNA to be threaded through it. Once the polymerase reaches the end of the template or detects double-stranded DNA, the sliding clamp undergoes a conformational change that releases the DNA polymerase. Clamp-loading proteins are used to initially load the clamp, recognizing the junction between template and RNA primers.:274-5
In all cases the helicase is composed of six polypeptides that wrap around only one strand of the DNA being replicated. The two polymerases are bound to the helicase heximer. In eukaryotes the helicase wraps around the leading strand, and in prokaryotes it wraps around the lagging strand.
As helicase unwinds DNA at the replication fork, the DNA ahead is forced to rotate. This process results in a build-up of twists in the DNA ahead. This build-up forms a torsional resistance that would eventually halt the progress of the replication fork. Topoisomerases are enzymes that temporarily break the strands of DNA, relieving the tension caused by unwinding the two strands of the DNA helix; topoisomerases (including DNA gyrase) achieve this by adding negative DNA supercoil, supercoils to the DNA helix.
Bare single-stranded DNA tends to fold back on itself forming Biomolecular structure#Secondary structure, secondary structures; these structures can interfere with the movement of DNA polymerase. To prevent this, single-strand binding proteins bind to the DNA until a second strand is synthesized, preventing secondary structure formation.
Double-stranded DNA is coiled around histones that play an important role in regulating gene expression so the replicated DNA must be coiled around histones at the same places as the original DNA. To ensure this, histone Chaperone (protein), chaperones disassemble the chromatin before it is replicated and replace the histones in the correct place. Some steps in this reassembly are somewhat speculative.
DNA clamp, Clamp proteins form a sliding clamp around DNA, helping the DNA polymerase maintain contact with its template, thereby assisting with processivity. The inner face of the clamp enables DNA to be threaded through it. Once the polymerase reaches the end of the template or detects double-stranded DNA, the sliding clamp undergoes a conformational change that releases the DNA polymerase. Clamp-loading proteins are used to initially load the clamp, recognizing the junction between template and RNA primers.:274-5
DNA replication proteins
At the replication fork, many replication enzymes assemble on the DNA into a complex molecular machine called the replisome. The following is a list of major DNA replication enzymes that participate in the replisome:Replication machinery
 Replication machineries consist of factors involved in DNA replication and appearing on template ssDNAs. Replication machineries include primosotors are replication enzymes; DNA polymerase, DNA helicases, DNA clamps and DNA topoisomerases, and replication proteins; e.g. single-stranded DNA binding proteins (SSB). In the replication machineries these components coordinate. In most of the bacteria, all of the factors involved in DNA replication are located on replication forks and the complexes stay on the forks during DNA replication. These replication machineries are called replisomes or DNA replicase systems. These terms are generic terms for proteins located on replication forks. In eukaryotic and some bacterial cells the replisomes are not formed.
Since replication machineries do not move relatively to template DNAs such as factories, they are called a replication factory. In an alternative figure, DNA factories are similar to projectors and DNAs are like as cinematic films passing constantly into the projectors. In the replication factory model, after both DNA helicases for leading strands and lagging strands are loaded on the template DNAs, the helicases run along the DNAs into each other. The helicases remain associated for the remainder of replication process. Peter Meister et al. observed directly replication sites in
Replication machineries consist of factors involved in DNA replication and appearing on template ssDNAs. Replication machineries include primosotors are replication enzymes; DNA polymerase, DNA helicases, DNA clamps and DNA topoisomerases, and replication proteins; e.g. single-stranded DNA binding proteins (SSB). In the replication machineries these components coordinate. In most of the bacteria, all of the factors involved in DNA replication are located on replication forks and the complexes stay on the forks during DNA replication. These replication machineries are called replisomes or DNA replicase systems. These terms are generic terms for proteins located on replication forks. In eukaryotic and some bacterial cells the replisomes are not formed.
Since replication machineries do not move relatively to template DNAs such as factories, they are called a replication factory. In an alternative figure, DNA factories are similar to projectors and DNAs are like as cinematic films passing constantly into the projectors. In the replication factory model, after both DNA helicases for leading strands and lagging strands are loaded on the template DNAs, the helicases run along the DNAs into each other. The helicases remain associated for the remainder of replication process. Peter Meister et al. observed directly replication sites in budding yeast
''Saccharomyces cerevisiae'' () (brewer's yeast or baker's yeast) is a species of yeast (single-celled fungus microorganisms). The species has been instrumental in winemaking, baking, and brewing since ancient times. It is believed to have been ...
by monitoring green fluorescent protein (GFP)-tagged DNA polymerases α. They detected DNA replication of pairs of the tagged loci spaced apart symmetrically from a replication origin and found that the distance between the pairs decreased markedly by time.Peter Meister, Angela Taddei1, Susan M. Gasser(June 2006), "In and out of the Replication Factory", ''Cell'' 125 (7): 1233–1235 This finding suggests that the mechanism of DNA replication goes with DNA factories. That is, couples of replication factories are loaded on replication origins and the factories associated with each other. Also, template DNAs move into the factories, which bring extrusion of the template ssDNAs and new DNAs. Meister's finding is the first direct evidence of replication factory model. Subsequent research has shown that DNA helicases form dimers in many eukaryotic cells and bacterial replication machineries stay in single intranuclear location during DNA synthesis.
The replication factories perform disentanglement of sister chromatids. The disentanglement is essential for distributing the chromatids into daughter cells after DNA replication. Because sister chromatids after DNA replication hold each other by Cohesin rings, there is the only chance for the disentanglement in DNA replication. Fixing of replication machineries as replication factories can improve the success rate of DNA replication. If replication forks move freely in chromosomes, catenation of nuclei is aggravated and impedes mitotic segregation.
Termination
Eukaryotes initiate DNA replication at multiple points in the chromosome, so replication forks meet and terminate at many points in the chromosome. Because eukaryotes have linear chromosomes, DNA replication is unable to reach the very end of the chromosomes. Due to this problem, DNA is lost in each replication cycle from the end of the chromosome. Telomeres are regions of repetitive DNA close to the ends and help prevent loss of genes due to this shortening. Shortening of the telomeres is a normal process in somatic cells. This shortens the telomeres of the daughter DNA chromosome. As a result, cells can only divide a certain number of times before the DNA loss prevents further division. (This is known as the Hayflick limit.) Within the germ cell line, which passes DNA to the next generation, telomerase extends the repetitive sequences of the telomere region to prevent degradation. Telomerase can become mistakenly active in somatic cells, sometimes leading to cancer formation. Increased telomerase activity is one of the hallmarks of cancer. Termination requires that the progress of the DNA replication fork must stop or be blocked. Termination at a specific locus, when it occurs, involves the interaction between two components: (1) a termination site sequence in the DNA, and (2) a protein which binds to this sequence to physically stop DNA replication. In various bacterial species, this is named the DNA replication terminus site-binding protein, or Ter protein. Because bacteria have circular chromosomes, termination of replication occurs when the two replication forks meet each other on the opposite end of the parental chromosome. ''E. coli'' regulates this process through the use of termination sequences that, when bound by the Tus protein, enable only one direction of replication fork to pass through. As a result, the replication forks are constrained to always meet within the termination region of the chromosome.Regulation
Eukaryotes
Within eukaryotes, DNA replication is controlled within the context of the cell cycle. As the cell grows and divides, it progresses through stages in the cell cycle; DNA replication takes place during the S phase (synthesis phase). The progress of the eukaryotic cell through the cycle is controlled by cell cycle checkpoints. Progression through checkpoints is controlled through complex interactions between various proteins, including cyclins and cyclin-dependent kinases. Unlike bacteria, eukaryotic DNA replicates in the confines of the nucleus. The G1/S checkpoint (or restriction checkpoint) regulates whether eukaryotic cells enter the process of DNA replication and subsequent division. Cells that do not proceed through this checkpoint remain in the G0 stage and do not replicate their DNA. After passing through the G1/S checkpoint, DNA must be replicated only once in each cell cycle. When the Mcm complex moves away from the origin, the pre-replication complex is dismantled. Because a new Mcm complex cannot be loaded at an origin until the pre-replication subunits are reactivated, one origin of replication can not be used twice in the same cell cycle. Activation of S-Cdks in early S phase promotes the destruction or inhibition of individual pre-replication complex components, preventing immediate reassembly. S and M-Cdks continue to block pre-replication complex assembly even after S phase is complete, ensuring that assembly cannot occur again until all Cdk activity is reduced in late mitosis. In budding yeast, inhibition of assembly is caused by Cdk-dependent phosphorylation of pre-replication complex components. At the onset of S phase, phosphorylation of Cdc6 by Cyclin-dependent kinase 1, Cdk1 causes the binding of Cdc6 to the SCF complex, SCF Ubiquitin--protein ligase, ubiquitin protein ligase, which causes proteolytic destruction of Cdc6. Cdk-dependent phosphorylation of Mcm proteins promotes their export out of the nucleus along with Cdt1 during S phase, preventing the loading of new Mcm complexes at origins during a single cell cycle. Cdk phosphorylation of the origin replication complex also inhibits pre-replication complex assembly. The individual presence of any of these three mechanisms is sufficient to inhibit pre-replication complex assembly. However, mutations of all three proteins in the same cell does trigger reinitiation at many origins of replication within one cell cycle. In animal cells, the protein geminin is a key inhibitor of pre-replication complex assembly. Geminin binds Cdt1, preventing its binding to the origin recognition complex. In G1, levels of geminin are kept low by the APC, which ubiquitinates geminin to target it for degradation. When geminin is destroyed, Cdt1 is released, allowing it to function in pre-replication complex assembly. At the end of G1, the APC is inactivated, allowing geminin to accumulate and bind Cdt1. Replication of chloroplast and mitochondrial genomes occurs independently of the cell cycle, through the process of D-loop replication.Replication focus
In vertebrate cells, replication sites concentrate into positions called replication foci. Replication sites can be detected by immunostaining daughter strands and replication enzymes and monitoring GFP-tagged replication factors. By these methods it is found that replication foci of varying size and positions appear in S phase of cell division and their number per nucleus is far smaller than the number of genomic replication forks. P. Heun et al.,(2001) tracked GFP-tagged replication foci in budding yeast cells and revealed that replication origins move constantly in G1 and S phase and the Molecular dynamics, dynamics decreased significantly in S phase. Traditionally, replication sites were fixed on spatial structure of chromosomes by nuclear matrix or lamins. The Heun's results denied the traditional concepts, budding yeasts do not have lamins, and support that replication origins self-assemble and form replication foci. By firing of replication origins, controlled spatially and temporally, the formation of replication foci is regulated. D. A. Jackson et al.(1998) revealed that neighboring origins fire simultaneously in mammalian cells. Spatial juxtaposition of replication sites brings clustering of replication forks. The clustering do rescue of stalled replication forks and favors normal progress of replication forks. Progress of replication forks is inhibited by many factors; collision with proteins or with complexes binding strongly on DNA, deficiency of dNTPs, nicks on template DNAs and so on. If replication forks stall and the remaining sequences from the stalled forks are not replicated, the daughter strands have nick obtained un-replicated sites. The un-replicated sites on one parent's strand hold the other strand together but not daughter strands. Therefore, the resulting sister chromatids cannot separate from each other and cannot divide into 2 daughter cells. When neighboring origins fire and a fork from one origin is stalled, fork from other origin access on an opposite direction of the stalled fork and duplicate the un-replicated sites. As other mechanism of the rescue there is application of dormant replication origins that excess origins do not fire in normal DNA replication.Bacteria
Problems with DNA replication

 There are many events that contribute to replication stress, including:
* Misincorporation of ribonucleotides
* Unusual Nucleic acid secondary structure, DNA structures
* Conflicts between replication and transcription
* Insufficiency of essential replication factors
* Chromosomal fragile site, Common fragile sites
* Overexpression or constitutive activation of oncogenes
* Chromatin inaccessibility
There are many events that contribute to replication stress, including:
* Misincorporation of ribonucleotides
* Unusual Nucleic acid secondary structure, DNA structures
* Conflicts between replication and transcription
* Insufficiency of essential replication factors
* Chromosomal fragile site, Common fragile sites
* Overexpression or constitutive activation of oncogenes
* Chromatin inaccessibility
Polymerase chain reaction
Researchers commonly replicate DNA ''in vitro'' using the polymerase chain reaction (PCR). PCR uses a pair of primer (molecular biology), primers to span a target region in template DNA, and then polymerizes partner strands in each direction from these primers using a thermostableDNA polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create ...
. Repeating this process through multiple cycles amplifies the targeted DNA region. At the start of each cycle, the mixture of template and primers is heated, separating the newly synthesized molecule and template. Then, as the mixture cools, both of these become templates for annealing of new primers, and the polymerase extends from these. As a result, the number of copies of the target region doubles each round, exponential growth, increasing exponentially.
See also
* Life * Cell (biology) * Cell division * Gene * Gene expression * Epigenetics * Genome * Autopoiesis * Chromosome segregation * Data storage device * Self-replication * Hachimoji DNANotes
References
{{Authority control DNA replication Senescence Cellular processes Molecular biology Copying