Polymerase chain reaction on:
[Wikipedia]
[Google]
[Amazon]
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) to a large enough amount to study in detail. PCR was invented in 1983 by the American biochemist Kary Mullis at

 PCR amplifies a specific region of a DNA strand (the DNA target). Most PCR methods amplify DNA fragments of between 0.1 and 10 kilo base pairs (kbp) in length, although some techniques allow for amplification of fragments up to 40 kbp. The amount of amplified product is determined by the available substrates in the reaction, which becomes limiting as the reaction progresses.
A basic PCR set-up requires several components and reagents, Chapter 8: In vitro Amplification of DNA by the Polymerase Chain Reaction including:
* a ''DNA template'' that contains the DNA target region to amplify
* a '' DNA polymerase''; an enzyme that polymerizes new DNA strands; heat-resistant ''Taq'' polymerase is especially common, as it is more likely to remain intact during the high-temperature DNA denaturation process
* two DNA '' primers'' that are complementary to the 3' (three prime) ends of each of the sense and anti-sense strands of the DNA target (DNA polymerase can only bind to and elongate from a double-stranded region of DNA; without primers, there is no double-stranded initiation site at which the polymerase can bind); specific primers that are complementary to the DNA target region are selected beforehand, and are often custom-made in a laboratory or purchased from commercial biochemical suppliers
* ''deoxynucleoside triphosphates'', or dNTPs (sometimes called "deoxynucleotide triphosphates";
PCR amplifies a specific region of a DNA strand (the DNA target). Most PCR methods amplify DNA fragments of between 0.1 and 10 kilo base pairs (kbp) in length, although some techniques allow for amplification of fragments up to 40 kbp. The amount of amplified product is determined by the available substrates in the reaction, which becomes limiting as the reaction progresses.
A basic PCR set-up requires several components and reagents, Chapter 8: In vitro Amplification of DNA by the Polymerase Chain Reaction including:
* a ''DNA template'' that contains the DNA target region to amplify
* a '' DNA polymerase''; an enzyme that polymerizes new DNA strands; heat-resistant ''Taq'' polymerase is especially common, as it is more likely to remain intact during the high-temperature DNA denaturation process
* two DNA '' primers'' that are complementary to the 3' (three prime) ends of each of the sense and anti-sense strands of the DNA target (DNA polymerase can only bind to and elongate from a double-stranded region of DNA; without primers, there is no double-stranded initiation site at which the polymerase can bind); specific primers that are complementary to the DNA target region are selected beforehand, and are often custom-made in a laboratory or purchased from commercial biochemical suppliers
* ''deoxynucleoside triphosphates'', or dNTPs (sometimes called "deoxynucleotide triphosphates";
 To check whether the PCR successfully generated the anticipated DNA target region (also sometimes referred to as the amplimer or amplicon),
To check whether the PCR successfully generated the anticipated DNA target region (also sometimes referred to as the amplimer or amplicon), 
 As with other chemical reactions, the reaction rate and efficiency of PCR are affected by limiting factors. Thus, the entire PCR process can further be divided into three stages based on reaction progress:
* ''Exponential amplification'': At every cycle, the amount of product is doubled (assuming 100% reaction efficiency). After 30 cycles, a single copy of DNA can be increased up to 1,000,000,000 (one billion) copies. In a sense, then, the replication of a discrete strand of DNA is being manipulated in a tube under controlled conditions. The reaction is very sensitive: only minute quantities of DNA must be present.
* ''Leveling off stage'': The reaction slows as the DNA polymerase loses activity and as consumption of reagents, such as dNTPs and primers, causes them to become more limited.
* ''Plateau'': No more product accumulates due to exhaustion of reagents and enzyme.
As with other chemical reactions, the reaction rate and efficiency of PCR are affected by limiting factors. Thus, the entire PCR process can further be divided into three stages based on reaction progress:
* ''Exponential amplification'': At every cycle, the amount of product is doubled (assuming 100% reaction efficiency). After 30 cycles, a single copy of DNA can be increased up to 1,000,000,000 (one billion) copies. In a sense, then, the replication of a discrete strand of DNA is being manipulated in a tube under controlled conditions. The reaction is very sensitive: only minute quantities of DNA must be present.
* ''Leveling off stage'': The reaction slows as the DNA polymerase loses activity and as consumption of reagents, such as dNTPs and primers, causes them to become more limited.
* ''Plateau'': No more product accumulates due to exhaustion of reagents and enzyme.
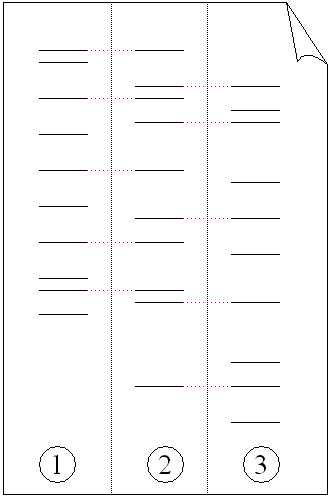 Some PCR fingerprint methods have high discriminative power and can be used to identify genetic relationships between individuals, such as parent-child or between siblings, and are used in paternity testing (Fig. 4). This technique may also be used to determine evolutionary relationships among organisms when certain molecular clocks are used (i.e. the
Some PCR fingerprint methods have high discriminative power and can be used to identify genetic relationships between individuals, such as parent-child or between siblings, and are used in paternity testing (Fig. 4). This technique may also be used to determine evolutionary relationships among organisms when certain molecular clocks are used (i.e. the
 In its most discriminating form, ''
In its most discriminating form, ''
 The heat-resistant enzymes that are a key component in polymerase chain reaction were discovered in the 1960s as a product of a microbial life form that lived in the superheated waters of Yellowstone's Mushroom Spring.
A 1971 paper in the '' Journal of Molecular Biology'' by Kjell Kleppe and co-workers in the laboratory of H. Gobind Khorana first described a method of using an enzymatic assay to replicate a short DNA template with primers ''in vitro''. However, this early manifestation of the basic PCR principle did not receive much attention at the time and the invention of the polymerase chain reaction in 1983 is generally credited to Kary Mullis.
When Mullis developed the PCR in 1983, he was working in Emeryville, California for
The heat-resistant enzymes that are a key component in polymerase chain reaction were discovered in the 1960s as a product of a microbial life form that lived in the superheated waters of Yellowstone's Mushroom Spring.
A 1971 paper in the '' Journal of Molecular Biology'' by Kjell Kleppe and co-workers in the laboratory of H. Gobind Khorana first described a method of using an enzymatic assay to replicate a short DNA template with primers ''in vitro''. However, this early manifestation of the basic PCR principle did not receive much attention at the time and the invention of the polymerase chain reaction in 1983 is generally credited to Kary Mullis.
When Mullis developed the PCR in 1983, he was working in Emeryville, California for
US Patent for PCR
What is PCR plateau effect?
YouTube tutorial video
History of the Polymerase Chain Reaction
from the Smithsonian Institution Archives {{DEFAULTSORT:Polymerase Chain Reaction Molecular biology Laboratory techniques DNA profiling techniques Amplifiers Roche Biotechnology Molecular biology techniques American inventions
Cetus Corporation
Cetus Corporation was one of the first biotechnology companies. It was established in Berkeley, California, in 1971, but conducted most of its operations in nearby Emeryville. Before merging with Chiron Corporation in 1991 (now a part of Novart ...
; Mullis and biochemist Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993.
PCR is fundamental to many of the procedures used in genetic testing and research, including analysis of ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences
A nucleic acid sequence is a succession of bases signified by a series of a set of five different letters that indicate the order of nucleotides forming alleles within a DNA (using GACT) or RNA (GACU) molecule. By convention, sequences are us ...
are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory
A medical laboratory or clinical laboratory is a laboratory where tests are conducted out on clinical specimens to obtain information about the health of a patient to aid in diagnosis, treatment, and prevention of disease. Clinical Medical labor ...
research for a broad variety of applications including biomedical research and criminal forensics.
The majority of PCR methods rely on thermal cycling. Thermal cycling exposes reactants to repeated cycles of heating and cooling to permit different temperature-dependent reactions—specifically, DNA melting and enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products ...
-driven DNA replication. PCR employs two main reagents— primers (which are short single strand DNA fragments known as oligonucleotide
Oligonucleotides are short DNA or RNA molecules, oligomers, that have a wide range of applications in genetic testing, research, and forensics. Commonly made in the laboratory by solid-phase chemical synthesis, these small bits of nucleic acids ...
s that are a complementary sequence to the target DNA region) and a DNA polymerase. In the first step of PCR, the two strands of the DNA double helix are physically separated at a high temperature in a process called nucleic acid denaturation. In the second step, the temperature is lowered and the primers bind to the complementary sequences of DNA. The two DNA strands then become templates
Template may refer to:
Tools
* Die (manufacturing), used to cut or shape material
* Mold, in a molding process
* Stencil, a pattern or overlay used in graphic arts (drawing, painting, etc.) and sewing to replicate letters, shapes or designs
Co ...
for DNA polymerase to enzymatically assemble a new DNA strand from free nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecule ...
s, the building blocks of DNA. As PCR progresses, the DNA generated is itself used as a template for replication, setting in motion a chain reaction
A chain reaction is a sequence of reactions where a reactive product or by-product causes additional reactions to take place. In a chain reaction, positive feedback leads to a self-amplifying chain of events.
Chain reactions are one way that sys ...
in which the original DNA template is exponentially
Exponential may refer to any of several mathematical topics related to exponentiation, including:
*Exponential function, also:
**Matrix exponential, the matrix analogue to the above
*Exponential decay, decrease at a rate proportional to value
*Expo ...
amplified.
Almost all PCR applications employ a heat-stable DNA polymerase, such as ''Taq'' polymerase, an enzyme originally isolated from the thermophilic
A thermophile is an organism—a type of extremophile—that thrives at relatively high temperatures, between . Many thermophiles are archaea, though they can be bacteria or fungi. Thermophilic eubacteria are suggested to have been among the earl ...
bacterium ''Thermus aquaticus
''Thermus aquaticus'' is a species of bacteria that can tolerate high temperatures, one of several thermophilic bacteria that belong to the ''Deinococcota'' phylum. It is the source of the heat-resistant enzyme ''Taq'' DNA polymerase, one of th ...
''. If the polymerase used was heat-susceptible, it would denature under the high temperatures of the denaturation step. Before the use of ''Taq'' polymerase, DNA polymerase had to be manually added every cycle, which was a tedious and costly process.
Applications of the technique include DNA cloning
Molecular cloning is a set of experimental methods in molecular biology that are used to assemble recombinant DNA molecules and to direct their replication within host organisms. The use of the word ''cloning'' refers to the fact that the metho ...
for sequencing, gene cloning and manipulation, gene mutagenesis; construction of DNA-based phylogenies, or functional analysis of gene
In biology, the word gene (from , ; "... Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a b ...
s; diagnosis and monitoring of genetic disorders; amplification of ancient DNA; analysis of genetic fingerprints for DNA profiling (for example, in forensic science and parentage testing); and detection of pathogens in nucleic acid test
A nucleic acid test (NAT) is a technique used to detect a particular nucleic acid sequence and thus usually to detect and identify a particular species or subspecies of organism, often a virus or bacterium that acts as a pathogen in blood, tissu ...
s for the diagnosis of infectious diseases.
Principles

nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecule ...
s containing triphosphate groups), the building blocks from which the DNA polymerase synthesizes a new DNA strand
* a ''buffer solution
A buffer solution (more precisely, pH buffer or hydrogen ion buffer) is an aqueous solution consisting of a mixture of a weak acid and its conjugate base, or vice versa. Its pH changes very little when a small amount of strong acid or base is ...
'' providing a suitable chemical environment for optimum activity and stability of the DNA polymerase
* '' bivalent cations
An ion () is an atom or molecule with a net electrical charge.
The charge of an electron is considered to be negative by convention and this charge is equal and opposite to the charge of a proton, which is considered to be positive by con ...
'', typically magnesium (Mg) or manganese
Manganese is a chemical element with the symbol Mn and atomic number 25. It is a hard, brittle, silvery metal, often found in minerals in combination with iron. Manganese is a transition metal with a multifaceted array of industrial alloy use ...
(Mn) ions; Mg2+ is the most common, but Mn2+ can be used for PCR-mediated DNA mutagenesis, as a higher Mn2+ concentration increases the error rate during DNA synthesis; and ''monovalent cations'', typically potassium (K) ions
The reaction is commonly carried out in a volume of 10–200 μL in small reaction tubes (0.2–0.5 mL volumes) in a thermal cycler
The thermal cycler (also known as a thermocycler, PCR machine or DNA amplifier) is a laboratory apparatus most commonly used to amplify segments of DNA via the polymerase chain reaction (PCR). Thermal cyclers may also be used in laboratories to fa ...
. The thermal cycler heats and cools the reaction tubes to achieve the temperatures required at each step of the reaction (see below). Many modern thermal cyclers make use of the Peltier effect
The thermoelectric effect is the direct conversion of temperature differences to electric voltage and vice versa via a thermocouple. A thermoelectric device creates a voltage when there is a different temperature on each side. Conversely, when ...
, which permits both heating and cooling of the block holding the PCR tubes simply by reversing the electric current. Thin-walled reaction tubes permit favorable thermal conductivity to allow for rapid thermal equilibrium. Most thermal cyclers have heated lids to prevent condensation at the top of the reaction tube. Older thermal cyclers lacking a heated lid require a layer of oil on top of the reaction mixture or a ball of wax inside the tube.
Procedure
Typically, PCR consists of a series of 20–40 repeated temperature changes, called thermal cycles, with each cycle commonly consisting of two or three discrete temperature steps (see figure below). The cycling is often preceded by a single temperature step at a very high temperature (>), and followed by one hold at the end for final product extension or brief storage. The temperatures used and the length of time they are applied in each cycle depend on a variety of parameters, including the enzyme used for DNA synthesis, the concentration of bivalent ions and dNTPs in the reaction, and the melting temperature (''Tm'') of the primers. The individual steps common to most PCR methods are as follows: * ''Initialization'': This step is only required for DNA polymerases that require heat activation by hot-start PCR. It consists of heating the reaction chamber to a temperature of , or if extremely thermostable polymerases are used, which is then held for 1–10 minutes. * '' Denaturation'': This step is the first regular cycling event and consists of heating the reaction chamber to for 20–30 seconds. This causes DNA melting, or denaturation, of the double-stranded DNA template by breaking the hydrogen bonds between complementary bases, yielding two single-stranded DNA molecules. * '' Annealing'': In the next step, the reaction temperature is lowered to for 20–40 seconds, allowing annealing of the primers to each of the single-stranded DNA templates. Two different primers are typically included in the reaction mixture: one for each of the two single-stranded complements containing the target region. The primers are single-stranded sequences themselves, but are much shorter than the length of the target region, complementing only very short sequences at the 3' end of each strand. : It is critical to determine a proper temperature for the annealing step because efficiency and specificity are strongly affected by the annealing temperature. This temperature must be low enough to allow for hybridization of the primer to the strand, but high enough for the hybridization to be specific, i.e., the primer should bind ''only'' to a perfectly complementary part of the strand, and nowhere else. If the temperature is too low, the primer may bind imperfectly. If it is too high, the primer may not bind at all. A typical annealing temperature is about 3–5 °C below the ''Tm'' of the primers used. Stable hydrogen bonds between complementary bases are formed only when the primer sequence very closely matches the template sequence. During this step, the polymerase binds to the primer-template hybrid and begins DNA formation. * ''Extension/elongation'': The temperature at this step depends on the DNA polymerase used; the optimum activity temperature for the thermostable DNA polymerase of ''Taq'' polymerase is approximately , though a temperature of is commonly used with this enzyme. In this step, the DNA polymerase synthesizes a new DNA strand complementary to the DNA template strand by adding free dNTPs from the reaction mixture that is complementary to the template in the 5'-to-3' direction, condensing the 5'-phosphate group
In chemistry, a phosphate is an anion, salt, functional group or ester derived from a phosphoric acid. It most commonly means orthophosphate, a derivative of orthophosphoric acid .
The phosphate or orthophosphate ion is derived from phosph ...
of the dNTPs with the 3'- hydroxy group at the end of the nascent (elongating) DNA strand. The precise time required for elongation depends both on the DNA polymerase used and on the length of the DNA target region to amplify. As a rule of thumb, at their optimal temperature, most DNA polymerases polymerize a thousand bases per minute. Under optimal conditions (i.e., if there are no limitations due to limiting substrates or reagents), at each extension/elongation step, the number of DNA target sequences is doubled. With each successive cycle, the original template strands plus all newly generated strands become template strands for the next round of elongation, leading to exponential (geometric) amplification of the specific DNA target region.
: The processes of denaturation, annealing and elongation constitute a single cycle. Multiple cycles are required to amplify the DNA target to millions of copies. The formula used to calculate the number of DNA copies formed after a given number of cycles is 2n, where ''n'' is the number of cycles. Thus, a reaction set for 30 cycles results in 230, or , copies of the original double-stranded DNA target region.
* ''Final elongation'': This single step is optional, but is performed at a temperature of (the temperature range required for optimal activity of most polymerases used in PCR) for 5–15 minutes after the last PCR cycle to ensure that any remaining single-stranded DNA is fully elongated.
* ''Final hold'': The final step cools the reaction chamber to for an indefinite time, and may be employed for short-term storage of the PCR products.
agarose gel electrophoresis
Agarose gel electrophoresis is a method of gel electrophoresis used in biochemistry, molecular biology, genetics, and clinical chemistry to separate a mixed population of macromolecules such as DNA or proteins in a matrix of agarose, one of the ...
may be employed for size separation of the PCR products. The size of the PCR products is determined by comparison with a DNA ladder, a molecular weight marker which contains DNA fragments of known sizes, which runs on the gel alongside the PCR products.

Stages
Optimization
In practice, PCR can fail for various reasons, such as sensitivity or contamination. Contamination with extraneous DNA can lead to spurious products and is addressed with lab protocols and procedures that separate pre-PCR mixtures from potential DNA contaminants. For instance, if DNA from a crime scene is analyzed, a single DNA molecule from lab personnel could be amplified and misguide the investigation. Hence the PCR-setup areas is separated from the analysis or purification of other PCR products, disposable plasticware used, and the work surface between reaction setups needs to be thoroughly cleaned. Specificity can be adjusted by experimental conditions so that no spurious products are generated. Primer-design techniques are important in improving PCR product yield and in avoiding the formation of unspecific products. The usage of alternate buffer components or polymerase enzymes can help with amplification of long or otherwise problematic regions of DNA. For instance, Q5 polymerase is said to be ~280 times less error-prone than Taq polymerase. Both the running parameters (e.g. temperature and duration of cycles), or the addition of reagents, such as formamide, may increase the specificity and yield of PCR. Computer simulations of theoretical PCR results ( Electronic PCR) may be performed to assist in primer design.Applications
Selective DNA isolation
PCR allows isolation of DNA fragments from genomic DNA by selective amplification of a specific region of DNA. This use of PCR augments many ways, such as generatinghybridization probe
In molecular biology, a hybridization probe (HP) is a fragment of DNA or RNA of usually 15–10000 nucleotide long which can be radioactively or fluorescently labeled. HP can be used to detect the presence of nucleotide sequences in analyzed RNA ...
s for Southern or northern hybridization and DNA cloning
Molecular cloning is a set of experimental methods in molecular biology that are used to assemble recombinant DNA molecules and to direct their replication within host organisms. The use of the word ''cloning'' refers to the fact that the metho ...
, which require larger amounts of DNA, representing a specific DNA region. PCR supplies these techniques with high amounts of pure DNA, enabling analysis of DNA samples even from very small amounts of starting material.
Other applications of PCR include DNA sequencing to determine unknown PCR-amplified sequences in which one of the amplification primers may be used in Sanger sequencing, isolation of a DNA sequence to expedite recombinant DNA technologies involving the insertion of a DNA sequence into a plasmid, phage
A bacteriophage (), also known informally as a ''phage'' (), is a duplodnaviria virus that infects and replicates within bacteria and archaea. The term was derived from "bacteria" and the Greek φαγεῖν ('), meaning "to devour". Bacter ...
, or cosmid (depending on size) or the genetic material of another organism. Bacterial colonies ''(such as E. coli)'' can be rapidly screened by PCR for correct DNA vector
Vector most often refers to:
*Euclidean vector, a quantity with a magnitude and a direction
*Vector (epidemiology), an agent that carries and transmits an infectious pathogen into another living organism
Vector may also refer to:
Mathematic ...
constructs. PCR may also be used for genetic fingerprinting
DNA profiling (also called DNA fingerprinting) is the process of determining an individual's DNA characteristics. DNA analysis intended to identify a species, rather than an individual, is called DNA barcoding.
DNA profiling is a forensic t ...
; a forensic technique used to identify a person or organism by comparing experimental DNAs through different PCR-based methods.
 Some PCR fingerprint methods have high discriminative power and can be used to identify genetic relationships between individuals, such as parent-child or between siblings, and are used in paternity testing (Fig. 4). This technique may also be used to determine evolutionary relationships among organisms when certain molecular clocks are used (i.e. the
Some PCR fingerprint methods have high discriminative power and can be used to identify genetic relationships between individuals, such as parent-child or between siblings, and are used in paternity testing (Fig. 4). This technique may also be used to determine evolutionary relationships among organisms when certain molecular clocks are used (i.e. the 16S rRNA 16S rRNA may refer to:
* 16S ribosomal RNA
16 S ribosomal RNA (or 16 S rRNA) is the RNA component of the 30S subunit of a prokaryotic ribosome ( SSU rRNA). It binds to the Shine-Dalgarno sequence and provides most of the SSU structure.
The g ...
and recA genes of microorganisms).
Amplification and quantification of DNA
Because PCR amplifies the regions of DNA that it targets, PCR can be used to analyze extremely small amounts of sample. This is often critical forforensic analysis
Forensic science, also known as criminalistics, is the application of science to criminal and civil laws, mainly—on the criminal side—during criminal investigation, as governed by the legal standards of admissible evidence and criminal p ...
, when only a trace amount of DNA is available as evidence. PCR may also be used in the analysis of ancient DNA
Ancient DNA (aDNA) is DNA isolated from ancient specimens. Due to degradation processes (including cross-linking, deamination and fragmentation) ancient DNA is more degraded in comparison with contemporary genetic material. Even under the bes ...
that is tens of thousands of years old. These PCR-based techniques have been successfully used on animals, such as a forty-thousand-year-old mammoth, and also on human DNA, in applications ranging from the analysis of Egyptian mummies to the identification of a Russian tsar and the body of English king Richard III.
Quantitative PCR
A real-time polymerase chain reaction (real-time PCR, or qPCR) is a laboratory technique of molecular biology based on the polymerase chain reaction (PCR). It monitors the amplification of a targeted DNA molecule during the PCR (i.e., in real ...
or Real Time PCR (qPCR, not to be confused with RT-PCR) methods allow the estimation of the amount of a given sequence present in a sample—a technique often applied to quantitatively determine levels of gene expression. Quantitative PCR is an established tool for DNA quantification that measures the accumulation of DNA product after each round of PCR amplification.
qPCR allows the quantification and detection of a specific DNA sequence in real time since it measures concentration while the synthesis process is taking place. There are two methods for simultaneous detection and quantification. The first method consists of using fluorescent dyes that are retained nonspecifically in between the double strands. The second method involves probes that code for specific sequences and are fluorescently labeled. Detection of DNA using these methods can only be seen after the hybridization of probes with its complementary DNA
In genetics, complementary DNA (cDNA) is DNA synthesized from a single-stranded RNA (e.g., messenger RNA (mRNA) or microRNA (miRNA)) template in a reaction catalyzed by the enzyme reverse transcriptase. cDNA is often used to express a spec ...
(cDNA) takes place. An interesting technique combination is real-time PCR and reverse transcription. This sophisticated technique, called RT-qPCR, allows for the quantification of a small quantity of RNA. Through this combined technique, mRNA is converted to cDNA, which is further quantified using qPCR. This technique lowers the possibility of error at the end point of PCR, increasing chances for detection of genes associated with genetic diseases such as cancer. Laboratories use RT-qPCR for the purpose of sensitively measuring gene regulation. The mathematical foundations for the reliable quantification of the PCR and RT-qPCR facilitate the implementation of accurate fitting procedures of experimental data in research, medical, diagnostic and infectious disease applications.
Medical and diagnostic applications
Prospective parents can be tested for beinggenetic carrier
A hereditary carrier (genetic carrier or just carrier), is a person or other organism that has inherited a recessive allele for a genetic trait or mutation but usually does not display that trait or show symptoms of the disease. Carriers are, ho ...
s, or their children might be tested for actually being affected by a disease
A disease is a particular abnormal condition that negatively affects the structure or function of all or part of an organism, and that is not immediately due to any external injury. Diseases are often known to be medical conditions that a ...
. DNA samples for prenatal testing can be obtained by amniocentesis
Amniocentesis is a medical procedure used primarily in the prenatal diagnosis of genetic conditions. It has other uses such as in the assessment of infection and fetal lung maturity. Prenatal diagnostic testing, which includes amniocentesis, is n ...
, chorionic villus sampling, or even by the analysis of rare fetal cells circulating in the mother's bloodstream. PCR analysis is also essential to preimplantation genetic diagnosis
Preimplantation genetic diagnosis (PGD or PIGD) is the genetic profiling of embryos prior to implantation (as a form of embryo profiling), and sometimes even of oocytes prior to fertilization. PGD is considered in a similar fashion to prenatal ...
, where individual cells of a developing embryo are tested for mutations.
* PCR can also be used as part of a sensitive test for '' tissue typing'', vital to organ transplantation
Organ transplantation is a medical procedure in which an organ is removed from one body and placed in the body of a recipient, to replace a damaged or missing organ. The donor and recipient may be at the same location, or organs may be transpor ...
. there is even a proposal to replace the traditional antibody-based tests for blood type with PCR-based tests.
* Many forms of cancer involve alterations to ''oncogene
An oncogene is a gene that has the potential to cause cancer. In tumor cells, these genes are often mutated, or expressed at high levels.
s''. By using PCR-based tests to study these mutations, therapy regimens can sometimes be individually customized to a patient. PCR permits early diagnosis of malignant
Malignancy () is the tendency of a medical condition to become progressively worse.
Malignancy is most familiar as a characterization of cancer. A ''malignant'' tumor contrasts with a non-cancerous ''benign'' tumor in that a malignancy is not s ...
diseases such as leukemia
Leukemia ( also spelled leukaemia and pronounced ) is a group of blood cancers that usually begin in the bone marrow and result in high numbers of abnormal blood cells. These blood cells are not fully developed and are called ''blasts'' or ...
and lymphomas, which is currently the highest-developed in cancer research and is already being used routinely. PCR assays can be performed directly on genomic DNA samples to detect translocation-specific malignant cells at a sensitivity that is at least 10,000 fold higher than that of other methods. PCR is very useful in the medical field since it allows for the isolation and amplification of tumor suppressors. Quantitative PCR for example, can be used to quantify and analyze single cells, as well as recognize DNA, mRNA and protein confirmations and combinations.
Infectious disease applications
PCR allows for rapid and highly specific diagnosis of infectious diseases, including those caused by bacteria or viruses. PCR also permits identification of non-cultivatable or slow-growing microorganisms such as mycobacteria, anaerobic bacteria, orvirus
A virus is a submicroscopic infectious agent that replicates only inside the living cells of an organism. Viruses infect all life forms, from animals and plants to microorganisms, including bacteria and archaea.
Since Dmitri Ivanovsk ...
es from tissue culture assays and animal model
An animal model (short for animal disease model) is a living, non-human, often genetic-engineered animal used during the research and investigation of human disease, for the purpose of better understanding the disease process without the risk of ha ...
s. The basis for PCR diagnostic applications in microbiology is the detection of infectious agents and the discrimination of non-pathogenic from pathogenic strains by virtue of specific genes.
Characterization and detection of infectious disease organisms have been revolutionized by PCR in the following ways:
* The ''human immunodeficiency virus'' (or ''HIV
The human immunodeficiency viruses (HIV) are two species of ''Lentivirus'' (a subgroup of retrovirus) that infect humans. Over time, they cause acquired immunodeficiency syndrome (AIDS), a condition in which progressive failure of the immune ...
''), is a difficult target to find and eradicate. The earliest tests for infection relied on the presence of antibodies to the virus circulating in the bloodstream. However, antibodies don't appear until many weeks after infection, maternal antibodies mask the infection of a newborn, and therapeutic agents to fight the infection don't affect the antibodies. PCR tests have been developed that can detect as little as one viral genome among the DNA of over 50,000 host cells. Infections can be detected earlier, donated blood can be screened directly for the virus, newborns can be immediately tested for infection, and the effects of antiviral treatments can be quantified.
* Some disease organisms, such as that for '' tuberculosis'', are difficult to sample from patients and slow to be grown in the laboratory. PCR-based tests have allowed detection of small numbers of disease organisms (both live or dead), in convenient samples. Detailed genetic analysis can also be used to detect antibiotic resistance, allowing immediate and effective therapy. The effects of therapy can also be immediately evaluated.
* The spread of a disease organism through populations of domestic or wild animals can be monitored by PCR testing. In many cases, the appearance of new virulent sub-types can be detected and monitored. The sub-types of an organism that were responsible for earlier epidemics can also be determined by PCR analysis.
* Viral DNA can be detected by PCR. The primers used must be specific to the targeted sequences in the DNA of a virus, and PCR can be used for diagnostic analyses or DNA sequencing of the viral genome. The high sensitivity of PCR permits virus detection soon after infection and even before the onset of disease. Such early detection may give physicians a significant lead time in treatment. The amount of virus ("viral load
Viral load, also known as viral burden, is a numerical expression of the quantity of virus in a given volume of fluid, including biological and environmental specimens. It is not to be confused with viral titre or viral titer, which depends on the ...
") in a patient can also be quantified by PCR-based DNA quantitation techniques (see below). A variant of PCR ( RT-PCR) is used for detecting viral RNA rather than DNA: in this test the enzyme reverse transcriptase is used to generate a DNA sequence which matches the viral RNA; this DNA is then amplified as per the usual PCR method. RT-PCR is widely used to detect the SARS-CoV-2 viral genome.
* Diseases such as pertussis (or whooping cough
Whooping cough, also known as pertussis or the 100-day cough, is a highly contagious bacterial disease. Initial symptoms are usually similar to those of the common cold with a runny nose, fever, and mild cough, but these are followed by two or t ...
) are caused by the bacteria '' Bordetella pertussis''. This bacteria is marked by a serious acute respiratory infection that affects various animals and humans and has led to the deaths of many young children. The pertussis toxin is a protein exotoxin that binds to cell receptors by two dimers and reacts with different cell types such as T lymphocytes which play a role in cell immunity. PCR is an important testing tool that can detect sequences within the gene for the pertussis toxin. Because PCR has a high sensitivity for the toxin and a rapid turnaround time, it is very efficient for diagnosing pertussis when compared to culture.
Forensic applications
The development of PCR-based genetic (or DNA) fingerprinting protocols has seen widespread application in forensics: * In its most discriminating form, ''
In its most discriminating form, ''genetic fingerprinting
DNA profiling (also called DNA fingerprinting) is the process of determining an individual's DNA characteristics. DNA analysis intended to identify a species, rather than an individual, is called DNA barcoding.
DNA profiling is a forensic t ...
'' can uniquely discriminate any one person from the entire population of the world
In its most general sense, the term "world" refers to the totality of entities, to the whole of reality or to everything that is. The nature of the world has been conceptualized differently in different fields. Some conceptions see the worl ...
. Minute samples of DNA can be isolated from a crime scene
A crime scene is any location that may be associated with a committed crime. Crime scenes contain physical evidence that is pertinent to a criminal investigation. This evidence is collected by crime scene investigators (CSI) and law enforcemen ...
, and compared to that from suspects, or from a DNA database of earlier evidence or convicts. Simpler versions of these tests are often used to rapidly rule out suspects during a criminal investigation. Evidence from decades-old crimes can be tested, confirming or exonerating the people originally convicted.
* Forensic DNA typing has been an effective way of identifying or exonerating criminal suspects due to analysis of evidence discovered at a crime scene. The human genome has many repetitive regions that can be found within gene sequences or in non-coding regions of the genome. Specifically, up to 40% of human DNA is repetitive. There are two distinct categories for these repetitive, non-coding regions in the genome. The first category is called variable number tandem repeats (VNTR), which are 10–100 base pairs long and the second category is called short tandem repeats (STR) and these consist of repeated 2–10 base pair sections. PCR is used to amplify several well-known VNTRs and STRs using primers that flank each of the repetitive regions. The sizes of the fragments obtained from any individual for each of the STRs will indicate which alleles are present. By analyzing several STRs for an individual, a set of alleles for each person will be found that statistically is likely to be unique. Researchers have identified the complete sequence of the human genome. This sequence can be easily accessed through the NCBI website and is used in many real-life applications. For example, the FBI has compiled a set of DNA marker sites used for identification, and these are called the Combined DNA Index System (CODIS) DNA database. Using this database enables statistical analysis to be used to determine the probability that a DNA sample will match. PCR is a very powerful and significant analytical tool to use for forensic DNA typing because researchers only need a very small amount of the target DNA to be used for analysis. For example, a single human hair with attached hair follicle has enough DNA to conduct the analysis. Similarly, a few sperm, skin samples from under the fingernails, or a small amount of blood can provide enough DNA for conclusive analysis.
* Less discriminating forms of DNA fingerprinting can help in '' DNA paternity testing'', where an individual is matched with their close relatives. DNA from unidentified human remains can be tested, and compared with that from possible parents, siblings, or children. Similar testing can be used to confirm the biological parents of an adopted (or kidnapped) child. The actual biological father of a newborn can also be confirmed (or ruled out).
* The PCR AMGX/AMGY design facilitate in amplifying DNA sequences from a very minuscule amount of genome. However it can also be used for real-time sex determination from forensic bone samples. This provides a powerful and effective way to determine gender in forensic cases and ancient specimens.
Research applications
PCR has been applied to many areas of research in molecular genetics: * PCR allows rapid production of short pieces of DNA, even when not more than the sequence of the two primers is known. This ability of PCR augments many methods, such as generating '' hybridization probes'' for Southern ornorthern blot
The northern blot, or RNA blot,Gilbert, S. F. (2000) Developmental Biology, 6th Ed. Sunderland MA, Sinauer Associates. is a technique used in molecular biology research to study gene expression by detection of RNA (or isolated mRNA) in a sample ...
hybridization. PCR supplies these techniques with large amounts of pure DNA, sometimes as a single strand, enabling analysis even from very small amounts of starting material.
* The task of '' DNA sequencing'' can also be assisted by PCR. Known segments of DNA can easily be produced from a patient with a genetic disease mutation. Modifications to the amplification technique can extract segments from a completely unknown genome, or can generate just a single strand of an area of interest.
* PCR has numerous applications to the more traditional process of ''DNA cloning
Molecular cloning is a set of experimental methods in molecular biology that are used to assemble recombinant DNA molecules and to direct their replication within host organisms. The use of the word ''cloning'' refers to the fact that the metho ...
''. It can extract segments for insertion into a vector from a larger genome, which may be only available in small quantities. Using a single set of 'vector primers', it can also analyze or extract fragments that have already been inserted into vectors. Some alterations to the PCR protocol can ''generate mutations'' (general or site-directed) of an inserted fragment.
* ''Sequence-tagged site A sequence-tagged site (or STS) is a short (200 to 500 base pair) DNA sequence that has a single occurrence in the genome and whose location and base sequence are known.
Usage
STSs can be easily detected by the polymerase chain reaction (PCR) usin ...
s'' is a process where PCR is used as an indicator that a particular segment of a genome is present in a particular clone. The Human Genome Project found this application vital to mapping the cosmid clones they were sequencing, and to coordinating the results from different laboratories.
* An application of PCR is the phylogenic analysis of DNA from '' ancient sources'', such as that found in the recovered bones of Neanderthals, from frozen tissues of mammoths, or from the brain of Egyptian mummies. In some cases the highly degraded DNA from these sources might be reassembled during the early stages of amplification.
* A common application of PCR is the study of patterns of '' gene expression''. Tissues (or even individual cells) can be analyzed at different stages to see which genes have become active, or which have been switched off. This application can also use quantitative PCR
A real-time polymerase chain reaction (real-time PCR, or qPCR) is a laboratory technique of molecular biology based on the polymerase chain reaction (PCR). It monitors the amplification of a targeted DNA molecule during the PCR (i.e., in real ...
to quantitate the actual levels of expression
* The ability of PCR to simultaneously amplify several loci from individual sperm has greatly enhanced the more traditional task of '' genetic mapping'' by studying chromosomal crossovers after meiosis
Meiosis (; , since it is a reductional division) is a special type of cell division of germ cells in sexually-reproducing organisms that produces the gametes, such as sperm or egg cells. It involves two rounds of division that ultimately r ...
. Rare crossover events between very close loci have been directly observed by analyzing thousands of individual sperms. Similarly, unusual deletions, insertions, translocations, or inversions can be analyzed, all without having to wait (or pay) for the long and laborious processes of fertilization, embryogenesis, etc.
* Site-directed mutagenesis
Site-directed mutagenesis is a molecular biology method that is used to make specific and intentional mutating changes to the DNA sequence of a gene and any gene products. Also called site-specific mutagenesis or oligonucleotide-directed mutagenesi ...
: PCR can be used to create mutant genes with mutations chosen by scientists at will. These mutations can be chosen in order to understand how proteins accomplish their functions, and to change or improve protein function.
Advantages
PCR has a number of advantages. It is fairly simple to understand and to use, and produces results rapidly. The technique is highly sensitive with the potential to produce millions to billions of copies of a specific product for sequencing, cloning, and analysis. qRT-PCR shares the same advantages as the PCR, with an added advantage of quantification of the synthesized product. Therefore, it has its uses to analyze alterations of gene expression levels in tumors, microbes, or other disease states. PCR is a very powerful and practical research tool. The sequencing of unknown etiologies of many diseases are being figured out by the PCR. The technique can help identify the sequence of previously unknown viruses related to those already known and thus give us a better understanding of the disease itself. If the procedure can be further simplified and sensitive non-radiometric detection systems can be developed, the PCR will assume a prominent place in the clinical laboratory for years to come.Limitations
One major limitation of PCR is that prior information about the target sequence is necessary in order to generate the primers that will allow its selective amplification. This means that, typically, PCR users must know the precise sequence(s) upstream of the target region on each of the two single-stranded templates in order to ensure that the DNA polymerase properly binds to the primer-template hybrids and subsequently generates the entire target region during DNA synthesis. Like all enzymes, DNA polymerases are also prone to error, which in turn causes mutations in the PCR fragments that are generated. Another limitation of PCR is that even the smallest amount of contaminating DNA can be amplified, resulting in misleading or ambiguous results. To minimize the chance of contamination, investigators should reserve separate rooms for reagent preparation, the PCR, and analysis of product. Reagents should be dispensed into single-use aliquots. Pipettors with disposable plungers and extra-long pipette tips should be routinely used. It is moreover recommended to ensure that the lab set-up follows a unidirectional workflow. No materials or reagents used in the PCR and analysis rooms should ever be taken into the PCR preparation room without thorough decontamination. Environmental samples that contain humic acids may inhibit PCR amplification and lead to inaccurate results.Variations
* ''Allele-specific PCR'': a diagnostic or cloning technique based on single-nucleotide variations (SNVs not to be confused withSNPs
In genetics, a single-nucleotide polymorphism (SNP ; plural SNPs ) is a germline substitution of a single nucleotide at a specific position in the genome. Although certain definitions require the substitution to be present in a sufficiently larg ...
) (single-base differences in a patient). It requires prior knowledge of a DNA sequence, including differences between allele
An allele (, ; ; modern formation from Greek ἄλλος ''állos'', "other") is a variation of the same sequence of nucleotides at the same place on a long DNA molecule, as described in leading textbooks on genetics and evolution.
::"The chro ...
s, and uses primers whose 3' ends encompass the SNV (base pair buffer around SNV usually incorporated). PCR amplification under stringent conditions is much less efficient in the presence of a mismatch between template and primer, so successful amplification with an SNP-specific primer signals presence of the specific SNP in a sequence. See SNP genotyping
SNP genotyping is the measurement of genetic variations of single nucleotide polymorphisms (SNPs) between members of a species. It is a form of genotyping, which is the measurement of more general genetic variation. SNPs are one of the most common ...
for more information.
* '' Assembly PCR'' or ''Polymerase Cycling Assembly (PCA)'': artificial synthesis of long DNA sequences by performing PCR on a pool of long oligonucleotides with short overlapping segments. The oligonucleotides alternate between sense and antisense directions, and the overlapping segments determine the order of the PCR fragments, thereby selectively producing the final long DNA product.
* '' Asymmetric PCR'': preferentially amplifies one DNA strand in a double-stranded DNA template. It is used in sequencing and hybridization probing where amplification of only one of the two complementary strands is required. PCR is carried out as usual, but with a great excess of the primer for the strand targeted for amplification. Because of the slow ( arithmetic) amplification later in the reaction after the limiting primer has been used up, extra cycles of PCR are required. A recent modification on this process, known as ''L''inear-''A''fter-''T''he-''E''xponential-PCR (LATE-PCR), uses a limiting primer with a higher melting temperature ( Tm) than the excess primer to maintain reaction efficiency as the limiting primer concentration decreases mid-reaction.
* ''Convective PCR'': a pseudo-isothermal way of performing PCR. Instead of repeatedly heating and cooling the PCR mixture, the solution is subjected to a thermal gradient. The resulting thermal instability driven convective flow automatically shuffles the PCR reagents from the hot and cold regions repeatedly enabling PCR. Parameters such as thermal boundary conditions and geometry of the PCR enclosure can be optimized to yield robust and rapid PCR by harnessing the emergence of chaotic flow fields. Such convective flow PCR setup significantly reduces device power requirement and operation time.
* ''Dial-out PCR'': a highly parallel method for retrieving accurate DNA molecules for gene synthesis. A complex library of DNA molecules is modified with unique flanking tags before massively parallel sequencing. Tag-directed primers then enable the retrieval of molecules with desired sequences by PCR.
* ''Digital PCR
Digital polymerase chain reaction (digital PCR, DigitalPCR, dPCR, or dePCR) is a biotechnological refinement of conventional polymerase chain reaction methods that can be used to directly quantify and clonally amplify nucleic acids strands includ ...
(dPCR)'': used to measure the quantity of a target DNA sequence in a DNA sample. The DNA sample is highly diluted so that after running many PCRs in parallel, some of them do not receive a single molecule of the target DNA. The target DNA concentration is calculated using the proportion of negative outcomes. Hence the name 'digital PCR'.
* ''Helicase-dependent amplification Helicase-dependent amplification (HDA) is a method for ''in vitro'' DNA amplification (like the polymerase chain reaction) that takes place at a constant temperature.
Introduction
The polymerase chain reaction is the most widely used method for '' ...
'': similar to traditional PCR, but uses a constant temperature rather than cycling through denaturation and annealing/extension cycles. DNA helicase
Helicases are a class of enzymes thought to be vital to all organisms. Their main function is to unpack an organism's genetic material. Helicases are motor proteins that move directionally along a nucleic acid phosphodiester backbone, separatin ...
, an enzyme that unwinds DNA, is used in place of thermal denaturation.
* '' Hot start PCR'': a technique that reduces non-specific amplification during the initial set up stages of the PCR. It may be performed manually by heating the reaction components to the denaturation temperature (e.g., 95 °C) before adding the polymerase. Specialized enzyme systems have been developed that inhibit the polymerase's activity at ambient temperature, either by the binding of an antibody or by the presence of covalently bound inhibitors that dissociate only after a high-temperature activation step. Hot-start/cold-finish PCR is achieved with new hybrid polymerases that are inactive at ambient temperature and are instantly activated at elongation temperature.
* '' In silico PCR'' (digital PCR, virtual PCR, electronic PCR, e-PCR) refers to computational tools used to calculate theoretical polymerase chain reaction results using a given set of primers ( probes) to amplify DNA sequences from a sequenced genome or transcriptome. In silico PCR was proposed as an educational tool for molecular biology.
* ''Intersequence-specific PCR'' (ISSR): a PCR method for DNA fingerprinting that amplifies regions between simple sequence repeats to produce a unique fingerprint of amplified fragment lengths.
* '' Inverse PCR'': is commonly used to identify the flanking sequences around genomic inserts. It involves a series of DNA digestions and self ligation, resulting in known sequences at either end of the unknown sequence.
* ''Ligation-mediated PCR'': uses small DNA linkers ligated to the DNA of interest and multiple primers annealing to the DNA linkers; it has been used for DNA sequencing, genome walking, and DNA footprinting DNA footprinting is a method of investigating the sequence specificity of DNA-binding proteins in vitro. This technique can be used to study protein-DNA interactions both outside and within cells.
The regulation of transcription has been studied ...
.
* '' Methylation-specific PCR'' (MSP): developed by Stephen Baylin and James G. Herman at the Johns Hopkins School of Medicine, and is used to detect methylation of CpG islands in genomic DNA. DNA is first treated with sodium bisulfite, which converts unmethylated cytosine bases to uracil, which is recognized by PCR primers as thymine. Two PCRs are then carried out on the modified DNA, using primer sets identical except at any CpG islands within the primer sequences. At these points, one primer set recognizes DNA with cytosines to amplify methylated DNA, and one set recognizes DNA with uracil or thymine to amplify unmethylated DNA. MSP using qPCR can also be performed to obtain quantitative rather than qualitative information about methylation.
* ''Miniprimer PCR'': uses a thermostable polymerase (S-Tbr) that can extend from short primers ("smalligos") as short as 9 or 10 nucleotides. This method permits PCR targeting to smaller primer binding regions, and is used to amplify conserved DNA sequences, such as the 16S (or eukaryotic 18S) rRNA gene.
* ''Multiplex ligation-dependent probe amplification
Multiplex ligation-dependent probe amplification (MLPA) is a variation of the multiplex polymerase chain reaction that permits amplification of multiple targets with only a single primer pair. It detects copy number changes at the molecular level, ...
'' (''MLPA''): permits amplifying multiple targets with a single primer pair, thus avoiding the resolution limitations of multiplex PCR (see below).
* '' Multiplex-PCR'': consists of multiple primer sets within a single PCR mixture to produce amplicons of varying sizes that are specific to different DNA sequences. By targeting multiple genes at once, additional information may be gained from a single test-run that otherwise would require several times the reagents and more time to perform. Annealing temperatures for each of the primer sets must be optimized to work correctly within a single reaction, and amplicon sizes. That is, their base pair length should be different enough to form distinct bands when visualized by gel electrophoresis.
* ''Nanoparticle-assisted PCR (nanoPCR)'': some nanoparticles (NPs) can enhance the efficiency of PCR (thus being called nanoPCR), and some can even outperform the original PCR enhancers. It was reported that quantum dots (QDs) can improve PCR specificity and efficiency. Single-walled carbon nanotubes (SWCNTs) and multi-walled carbon nanotubes (MWCNTs) are efficient in enhancing the amplification of long PCR. Carbon nanopowder (CNP) can improve the efficiency of repeated PCR and long PCR, while zinc oxide
Zinc oxide is an inorganic compound with the formula . It is a white powder that is insoluble in water. ZnO is used as an additive in numerous materials and products including cosmetics, food supplements, rubbers, plastics, ceramics, glass, cement ...
, titanium dioxide and Ag NPs were found to increase the PCR yield. Previous data indicated that non-metallic NPs retained acceptable amplification fidelity. Given that many NPs are capable of enhancing PCR efficiency, it is clear that there is likely to be great potential for nanoPCR technology improvements and product development.
* '' Nested PCR'': increases the specificity of DNA amplification, by reducing background due to non-specific amplification of DNA. Two sets of primers are used in two successive PCRs. In the first reaction, one pair of primers is used to generate DNA products, which besides the intended target, may still consist of non-specifically amplified DNA fragments. The product(s) are then used in a second PCR with a set of primers whose binding sites are completely or partially different from and located 3' of each of the primers used in the first reaction. Nested PCR is often more successful in specifically amplifying long DNA fragments than conventional PCR, but it requires more detailed knowledge of the target sequences.
* '' Overlap-extension PCR'' or ''Splicing by overlap extension (SOEing) '': a genetic engineering technique that is used to splice together two or more DNA fragments that contain complementary sequences. It is used to join DNA pieces containing genes, regulatory sequences, or mutations; the technique enables creation of specific and long DNA constructs. It can also introduce deletions, insertions or point mutations into a DNA sequence.
* ''PAN-AC'': uses isothermal conditions for amplification, and may be used in living cells.
* ''Quantitative PCR
A real-time polymerase chain reaction (real-time PCR, or qPCR) is a laboratory technique of molecular biology based on the polymerase chain reaction (PCR). It monitors the amplification of a targeted DNA molecule during the PCR (i.e., in real ...
'' (qPCR): used to measure the quantity of a target sequence (commonly in real-time). It quantitatively measures starting amounts of DNA, cDNA, or RNA. Quantitative PCR is commonly used to determine whether a DNA sequence is present in a sample and the number of its copies in the sample. ''Quantitative PCR'' has a very high degree of precision. Quantitative PCR methods use fluorescent dyes, such as Sybr Green, EvaGreen or fluorophore-containing DNA probes, such as TaqMan, to measure the amount of amplified product in real time. It is also sometimes abbreviated to RT-PCR (''real-time'' PCR) but this abbreviation should be used only for reverse transcription PCR. qPCR is the appropriate contractions for quantitative PCR
A real-time polymerase chain reaction (real-time PCR, or qPCR) is a laboratory technique of molecular biology based on the polymerase chain reaction (PCR). It monitors the amplification of a targeted DNA molecule during the PCR (i.e., in real ...
(real-time PCR).
* '' Reverse Complement PCR'' (RC-PCR): Allows the addition of functional domains or sequences of choice to be appended independently to either end of the generated amplicon in a single closed tube reaction. This method generates target specific primers within the reaction by the interaction of universal primers (which contain the desired sequences or domains to be appended) and RC probes.
* ''Reverse Transcription PCR ( RT-PCR)'': for amplifying DNA from RNA. Reverse transcriptase reverse transcribes RNA into cDNA, which is then amplified by PCR. RT-PCR is widely used in expression profiling
In the field of molecular biology, gene expression profiling is the measurement of the activity (the expression) of thousands of genes at once, to create a global picture of cellular function. These profiles can, for example, distinguish between c ...
, to determine the expression of a gene or to identify the sequence of an RNA transcript, including transcription start and termination sites. If the genomic DNA sequence of a gene is known, RT-PCR can be used to map the location of exons and introns
An intron is any nucleotide sequence within a gene that is not expressed or operative in the final RNA product. The word ''intron'' is derived from the term ''intragenic region'', i.e. a region inside a gene."The notion of the cistron .e., gene ...
in the gene. The 5' end of a gene (corresponding to the transcription start site) is typically identified by RACE-PCR (''Rapid Amplification of cDNA Ends'').
* '' RNase H-dependent PCR'' (rhPCR): a modification of PCR that utilizes primers with a 3' extension block that can be removed by a thermostable RNase HII enzyme. This system reduces primer-dimers and allows for multiplexed reactions to be performed with higher numbers of primers.
* ''Single specific primer-PCR'' (SSP-PCR): allows the amplification of double-stranded DNA even when the sequence information is available at one end only. This method permits amplification of genes for which only a partial sequence information is available, and allows unidirectional genome walking from known into unknown regions of the chromosome.
* ''Solid Phase PCR'': encompasses multiple meanings, including Polony Amplification (where PCR colonies are derived in a gel matrix, for example), Bridge PCR (primers are covalently linked to a solid-support surface), conventional Solid Phase PCR (where Asymmetric PCR is applied in the presence of solid support bearing primer with sequence matching one of the aqueous primers) and Enhanced Solid Phase PCR (where conventional Solid Phase PCR can be improved by employing high Tm and nested solid support primer with optional application of a thermal 'step' to favour solid support priming).
* ''Suicide PCR'': typically used in paleogenetics Paleogenetics is the study of the past through the examination of preserved genetic material from the remains of ancient organisms. Emile Zuckerkandl and Linus Pauling introduced the term in 1963, long before the sequencing of DNA, in reference t ...
or other studies where avoiding false positives and ensuring the specificity of the amplified fragment is the highest priority. It was originally described in a study to verify the presence of the microbe Yersinia pestis
''Yersinia pestis'' (''Y. pestis''; formerly '' Pasteurella pestis'') is a gram-negative, non-motile, coccobacillus bacterium without spores that is related to both ''Yersinia pseudotuberculosis'' and ''Yersinia enterocolitica''. It is a facult ...
in dental samples obtained from 14th Century graves of people supposedly killed by the plague during the medieval Black Death epidemic. The method prescribes the use of any primer combination only once in a PCR (hence the term "suicide"), which should never have been used in any positive control PCR reaction, and the primers should always target a genomic region never amplified before in the lab using this or any other set of primers. This ensures that no contaminating DNA from previous PCR reactions is present in the lab, which could otherwise generate false positives.
* ''Thermal asymmetric interlaced PCR (TAIL-PCR)'': for isolation of an unknown sequence flanking a known sequence. Within the known sequence, TAIL-PCR uses a nested pair of primers with differing annealing temperatures; a degenerate primer is used to amplify in the other direction from the unknown sequence.
* '' Touchdown PCR'' (''Step-down PCR''): a variant of PCR that aims to reduce nonspecific background by gradually lowering the annealing temperature as PCR cycling progresses. The annealing temperature at the initial cycles is usually a few degrees (3–5 °C) above the Tm of the primers used, while at the later cycles, it is a few degrees (3–5 °C) below the primer Tm. The higher temperatures give greater specificity for primer binding, and the lower temperatures permit more efficient amplification from the specific products formed during the initial cycles.
* ''Universal Fast Walking'': for genome walking and genetic fingerprinting using a more specific 'two-sided' PCR than conventional 'one-sided' approaches (using only one gene-specific primer and one general primer—which can lead to artefactual 'noise') by virtue of a mechanism involving lariat structure formation. Streamlined derivatives of UFW are LaNe RAGE (lariat-dependent nested PCR for rapid amplification of genomic DNA ends), 5'RACE LaNe and 3'RACE LaNe.
History
Cetus Corporation
Cetus Corporation was one of the first biotechnology companies. It was established in Berkeley, California, in 1971, but conducted most of its operations in nearby Emeryville. Before merging with Chiron Corporation in 1991 (now a part of Novart ...
, one of the first biotechnology companies, where he was responsible for synthesizing short chains of DNA. Mullis has written that he conceived the idea for PCR while cruising along the Pacific Coast Highway one night in his car. He was playing in his mind with a new way of analyzing changes (mutations) in DNA when he realized that he had instead invented a method of amplifying any DNA region through repeated cycles of duplication driven by DNA polymerase. In ''Scientific American
''Scientific American'', informally abbreviated ''SciAm'' or sometimes ''SA'', is an American popular science magazine. Many famous scientists, including Albert Einstein and Nikola Tesla, have contributed articles to it. In print since 1845, it ...
'', Mullis summarized the procedure: "Beginning with a single molecule of the genetic material DNA, the PCR can generate 100 billion similar molecules in an afternoon. The reaction is easy to execute. It requires no more than a test tube, a few simple reagents, and a source of heat." DNA fingerprinting was first used for paternity testing in 1988.
Mullis has credited his use of LSD
Lysergic acid diethylamide (LSD), also known colloquially as acid, is a potent psychedelic drug. Effects typically include intensified thoughts, emotions, and sensory perception. At sufficiently high dosages LSD manifests primarily mental, vi ...
as integral to his development of PCR: "Would I have invented PCR if I hadn't taken LSD? I seriously doubt it. I could sit on a DNA molecule and watch the polymers go by. I learnt that partly on psychedelic drugs."
Mullis and biochemist Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993, seven years after Mullis and his colleagues at Cetus first put his proposal to practice. Mullis's 1985 paper with R. K. Saiki and H. A. Erlich, "Enzymatic Amplification of β-globin Genomic Sequences and Restriction Site Analysis for Diagnosis of Sickle Cell Anemia"—the polymerase chain reaction invention (PCR)—was honored by a Citation for Chemical Breakthrough Award from the Division of History of Chemistry of the American Chemical Society in 2017.
At the core of the PCR method is the use of a suitable DNA polymerase able to withstand the high temperatures of > required for separation of the two DNA strands in the DNA double helix after each replication cycle. The DNA polymerases initially employed for in vitro
''In vitro'' (meaning in glass, or ''in the glass'') studies are performed with microorganisms, cells, or biological molecules outside their normal biological context. Colloquially called " test-tube experiments", these studies in biology ...
experiments presaging PCR were unable to withstand these high temperatures. So the early procedures for DNA replication were very inefficient and time-consuming, and required large amounts of DNA polymerase and continuous handling throughout the process.
The discovery in 1976 of ''Taq'' polymerase—a DNA polymerase purified from the thermophilic bacterium, ''Thermus aquaticus
''Thermus aquaticus'' is a species of bacteria that can tolerate high temperatures, one of several thermophilic bacteria that belong to the ''Deinococcota'' phylum. It is the source of the heat-resistant enzyme ''Taq'' DNA polymerase, one of th ...
'', which naturally lives in hot () environments such as hot springs—paved the way for dramatic improvements of the PCR method. The DNA polymerase isolated from ''T. aquaticus'' is stable at high temperatures remaining active even after DNA denaturation, thus obviating the need to add new DNA polymerase after each cycle. This allowed an automated thermocycler-based process for DNA amplification.
Patent disputes
The PCR technique was patented by Kary Mullis and assigned toCetus Corporation
Cetus Corporation was one of the first biotechnology companies. It was established in Berkeley, California, in 1971, but conducted most of its operations in nearby Emeryville. Before merging with Chiron Corporation in 1991 (now a part of Novart ...
, where Mullis worked when he invented the technique in 1983. The ''Taq'' polymerase enzyme was also covered by patents. There have been several high-profile lawsuits related to the technique, including an unsuccessful lawsuit brought by DuPont. The Swiss pharmaceutical company Hoffmann-La Roche purchased the rights to the patents in 1992. The last of the commercial PCR patents expired in 2017.
A related patent battle over the ''Taq'' polymerase enzyme is still ongoing in several jurisdictions around the world between Roche and Promega. The legal arguments have extended beyond the lives of the original PCR and ''Taq'' polymerase patents, which expired on 28 March 2005.
See also
* COVID-19 testing * DNA spiking *Loop-mediated isothermal amplification
Loop-mediated isothermal amplification (LAMP) is a single-tube technique for the amplification of DNA and a low-cost alternative to detect certain diseases. Reverse transcription loop-mediated isothermal amplification (RT-LAMP) combines LAMP with ...
* Selector technique
* Thermus thermophilus
* Pfu DNA polymerase
References
External links
US Patent for PCR
What is PCR plateau effect?
YouTube tutorial video
History of the Polymerase Chain Reaction
from the Smithsonian Institution Archives {{DEFAULTSORT:Polymerase Chain Reaction Molecular biology Laboratory techniques DNA profiling techniques Amplifiers Roche Biotechnology Molecular biology techniques American inventions