Metabolism on:
[Wikipedia]
[Google]
[Amazon]

 Metabolism (, from el, μεταβολή ''metabolē'', "change") is the set of
Metabolism (, from el, μεταβολή ''metabolē'', "change") is the set of
 Most of the structures that make up animals, plants and microbes are made from four basic classes of
Most of the structures that make up animals, plants and microbes are made from four basic classes of
 Carbohydrates are
Carbohydrates are
 Metabolism involves a vast array of chemical reactions, but most fall under a few basic types of reactions that involve the transfer of functional groups of atoms and their bonds within molecules. This common chemistry allows cells to use a small set of metabolic intermediates to carry chemical groups between different reactions. These group-transfer intermediates are called coenzymes. Each class of group-transfer reactions is carried out by a particular coenzyme, which is the substrate for a set of enzymes that produce it, and a set of enzymes that consume it. These coenzymes are therefore continuously made, consumed and then recycled.
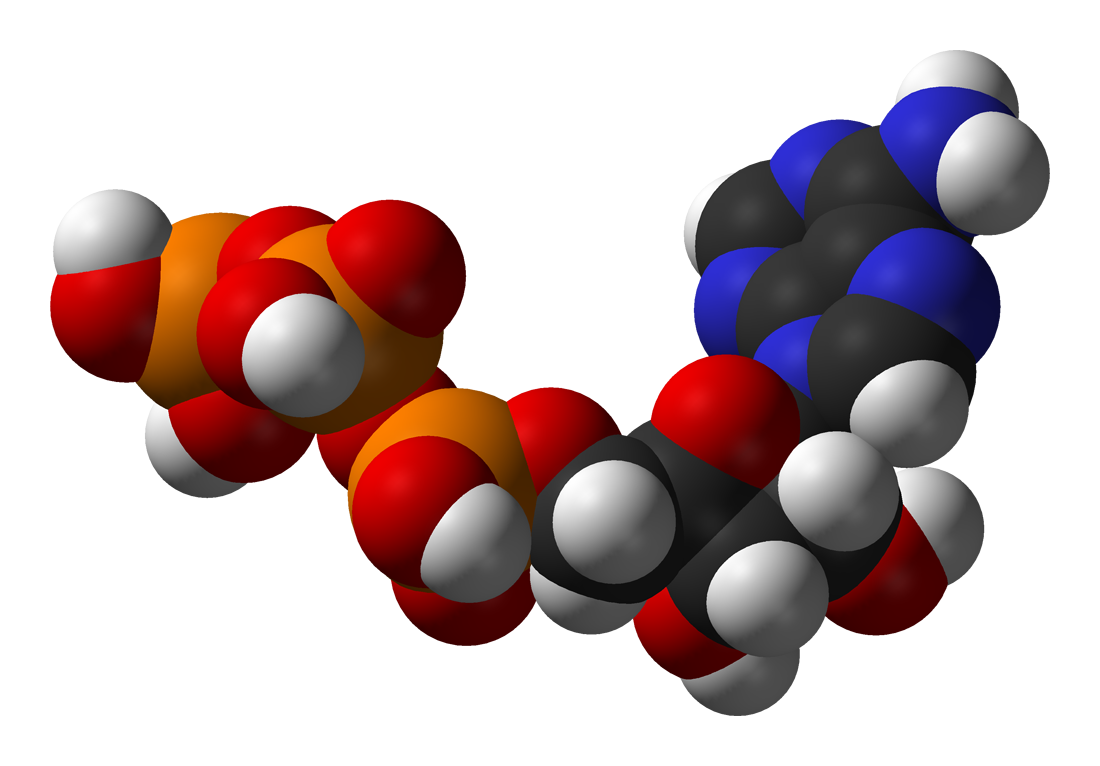
One central coenzyme is adenosine triphosphate (ATP), the universal energy currency of cells. This nucleotide is used to transfer chemical energy between different chemical reactions. There is only a small amount of ATP in cells, but as it is continuously regenerated, the human body can use about its own weight in ATP per day. ATP acts as a bridge between catabolism and anabolism. Catabolism breaks down molecules, and anabolism puts them together. Catabolic reactions generate ATP, and anabolic reactions consume it. It also serves as a carrier of phosphate groups in phosphorylation reactions.
A vitamin is an organic compound needed in small quantities that cannot be made in cells. In
Metabolism involves a vast array of chemical reactions, but most fall under a few basic types of reactions that involve the transfer of functional groups of atoms and their bonds within molecules. This common chemistry allows cells to use a small set of metabolic intermediates to carry chemical groups between different reactions. These group-transfer intermediates are called coenzymes. Each class of group-transfer reactions is carried out by a particular coenzyme, which is the substrate for a set of enzymes that produce it, and a set of enzymes that consume it. These coenzymes are therefore continuously made, consumed and then recycled.
One central coenzyme is adenosine triphosphate (ATP), the universal energy currency of cells. This nucleotide is used to transfer chemical energy between different chemical reactions. There is only a small amount of ATP in cells, but as it is continuously regenerated, the human body can use about its own weight in ATP per day. ATP acts as a bridge between catabolism and anabolism. Catabolism breaks down molecules, and anabolism puts them together. Catabolic reactions generate ATP, and anabolic reactions consume it. It also serves as a carrier of phosphate groups in phosphorylation reactions.
A vitamin is an organic compound needed in small quantities that cannot be made in cells. In 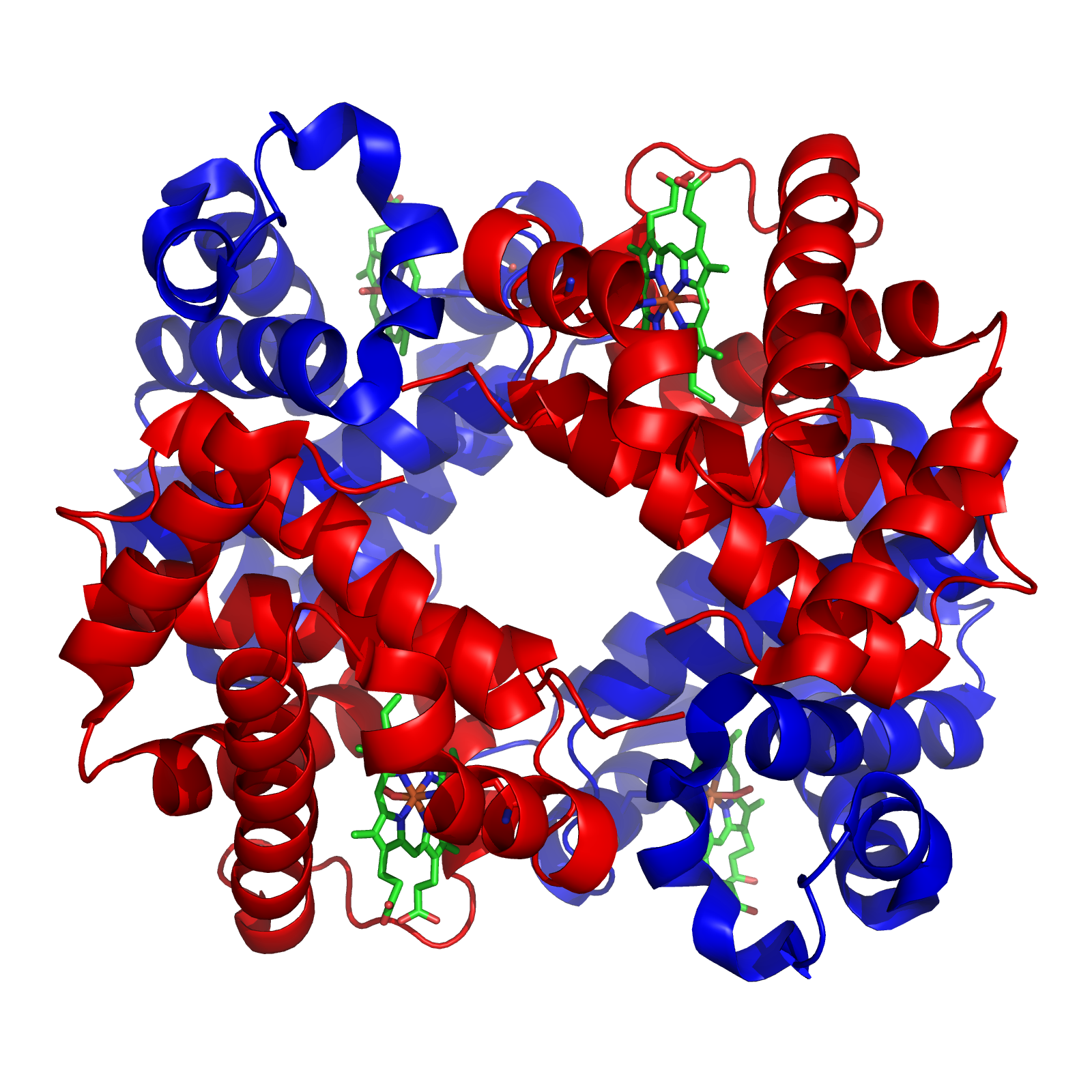

 Pumping protons out of the mitochondria creates a proton concentration difference across the membrane and generates an
Pumping protons out of the mitochondria creates a proton concentration difference across the membrane and generates an
 Photosynthesis is the synthesis of carbohydrates from sunlight and
Photosynthesis is the synthesis of carbohydrates from sunlight and
 Fatty acids are made by fatty acid synthases that polymerize and then reduce acetyl-CoA units. The acyl chains in the fatty acids are extended by a cycle of reactions that add the acyl group, reduce it to an alcohol, dehydrate it to an alkene group and then reduce it again to an alkane group. The enzymes of fatty acid biosynthesis are divided into two groups: in animals and fungi, all these fatty acid synthase reactions are carried out by a single multifunctional type I protein, while in plant plastids and bacteria separate type II enzymes perform each step in the pathway.
Terpenes and isoprenoids are a large class of lipids that include the carotenoids and form the largest class of plant natural products. These compounds are made by the assembly and modification of isoprene units donated from the reactive precursors
Fatty acids are made by fatty acid synthases that polymerize and then reduce acetyl-CoA units. The acyl chains in the fatty acids are extended by a cycle of reactions that add the acyl group, reduce it to an alcohol, dehydrate it to an alkene group and then reduce it again to an alkane group. The enzymes of fatty acid biosynthesis are divided into two groups: in animals and fungi, all these fatty acid synthase reactions are carried out by a single multifunctional type I protein, while in plant plastids and bacteria separate type II enzymes perform each step in the pathway.
Terpenes and isoprenoids are a large class of lipids that include the carotenoids and form the largest class of plant natural products. These compounds are made by the assembly and modification of isoprene units donated from the reactive precursors
 There are multiple levels of metabolic regulation. In intrinsic regulation, the metabolic pathway self-regulates to respond to changes in the levels of substrates or products; for example, a decrease in the amount of product can increase the flux through the pathway to compensate. This type of regulation often involves allosteric regulation of the activities of multiple enzymes in the pathway. Extrinsic control involves a cell in a multicellular organism changing its metabolism in response to signals from other cells. These signals are usually in the form of water-soluble messengers such as hormones and
There are multiple levels of metabolic regulation. In intrinsic regulation, the metabolic pathway self-regulates to respond to changes in the levels of substrates or products; for example, a decrease in the amount of product can increase the flux through the pathway to compensate. This type of regulation often involves allosteric regulation of the activities of multiple enzymes in the pathway. Extrinsic control involves a cell in a multicellular organism changing its metabolism in response to signals from other cells. These signals are usually in the form of water-soluble messengers such as hormones and
 The central pathways of metabolism described above, such as glycolysis and the citric acid cycle, are present in all three domains of living things and were present in the
The central pathways of metabolism described above, such as glycolysis and the citric acid cycle, are present in all three domains of living things and were present in the
 Classically, metabolism is studied by a reductionist approach that focuses on a single metabolic pathway. Particularly valuable is the use of radioactive tracers at the whole-organism, tissue and cellular levels, which define the paths from precursors to final products by identifying radioactively labelled intermediates and products. The enzymes that catalyze these chemical reactions can then be purified and their kinetics and responses to inhibitors investigated. A parallel approach is to identify the small molecules in a cell or tissue; the complete set of these molecules is called the
Classically, metabolism is studied by a reductionist approach that focuses on a single metabolic pathway. Particularly valuable is the use of radioactive tracers at the whole-organism, tissue and cellular levels, which define the paths from precursors to final products by identifying radioactively labelled intermediates and products. The enzymes that catalyze these chemical reactions can then be purified and their kinetics and responses to inhibitors investigated. A parallel approach is to identify the small molecules in a cell or tissue; the complete set of these molecules is called the

 In these early studies, the mechanisms of these metabolic processes had not been identified and a vital force was thought to animate living tissue. In the 19th century, when studying the fermentation of sugar to alcohol by yeast, Louis Pasteur concluded that fermentation was catalyzed by substances within the yeast cells he called "ferments". He wrote that "alcoholic fermentation is an act correlated with the life and organization of the yeast cells, not with the death or putrefaction of the cells." This discovery, along with the publication by Friedrich Wöhler in 1828 of a paper on the chemical synthesis of urea, and is notable for being the first organic compound prepared from wholly inorganic precursors. This proved that the organic compounds and chemical reactions found in cells were no different in principle than any other part of chemistry.
It was the discovery of
In these early studies, the mechanisms of these metabolic processes had not been identified and a vital force was thought to animate living tissue. In the 19th century, when studying the fermentation of sugar to alcohol by yeast, Louis Pasteur concluded that fermentation was catalyzed by substances within the yeast cells he called "ferments". He wrote that "alcoholic fermentation is an act correlated with the life and organization of the yeast cells, not with the death or putrefaction of the cells." This discovery, along with the publication by Friedrich Wöhler in 1828 of a paper on the chemical synthesis of urea, and is notable for being the first organic compound prepared from wholly inorganic precursors. This proved that the organic compounds and chemical reactions found in cells were no different in principle than any other part of chemistry.
It was the discovery of
The Biochemistry of Metabolism
Sparknotes SAT biochemistry
Overview of biochemistry. School level.
Undergraduate-level guide to molecular biology. Human metabolism
Guide to human metabolic pathways. School level.
THE Medical Biochemistry Page
Comprehensive resource on human metabolism. Databases
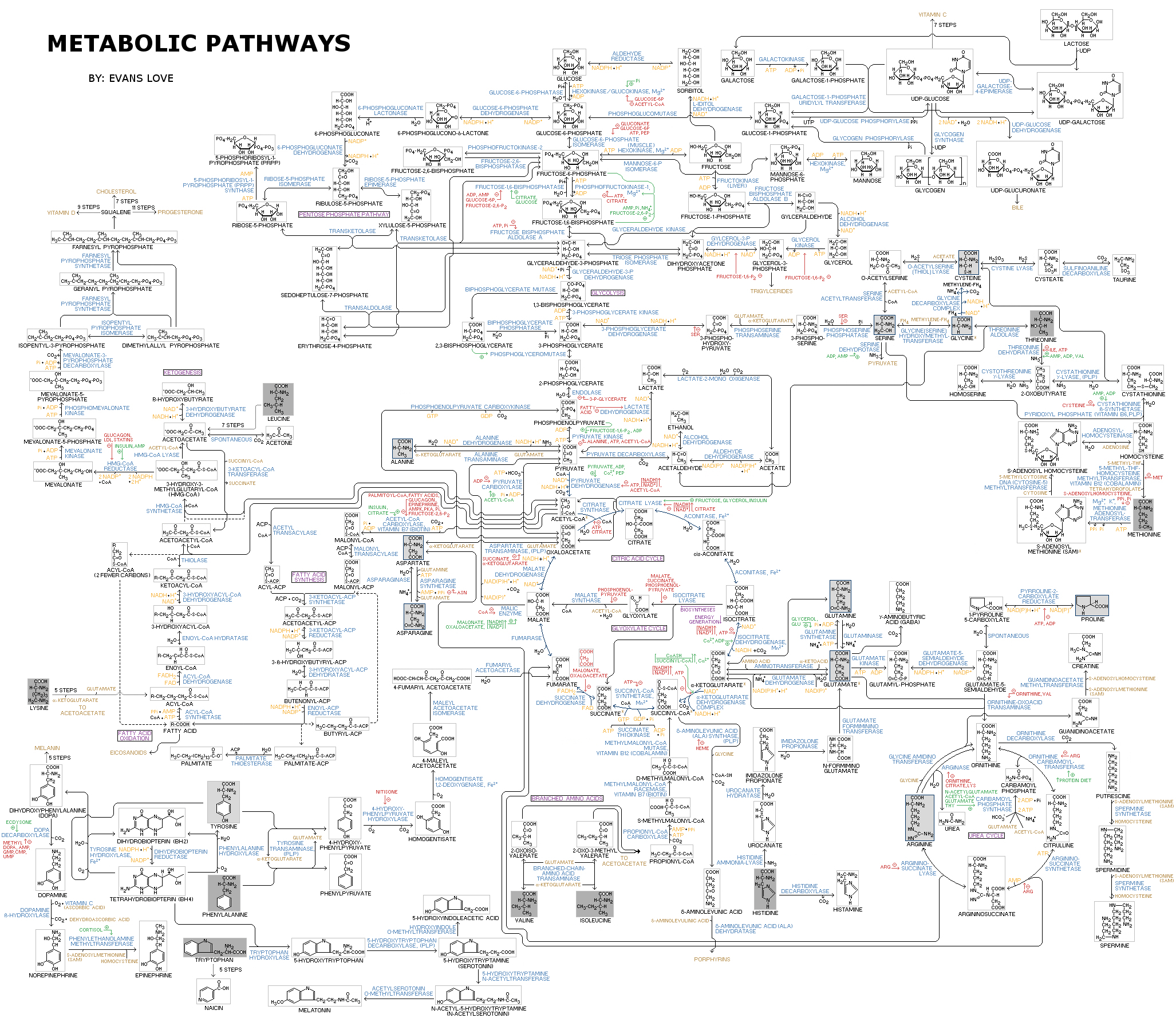
Flow Chart of Metabolic Pathways
at ExPASy
IUBMB-Nicholson Metabolic Pathways Chart
SuperCYP: Database for Drug-Cytochrome-Metabolism
Metabolic pathways
* {{Authority control, state=collapsed Underwater diving physiology

 Metabolism (, from el, μεταβολή ''metabolē'', "change") is the set of
Metabolism (, from el, μεταβολή ''metabolē'', "change") is the set of life
Life is a quality that distinguishes matter that has biological processes, such as Cell signaling, signaling and self-sustaining processes, from that which does not, and is defined by the capacity for Cell growth, growth, reaction to Stimu ...
-sustaining chemical reactions in organisms. The three main functions of metabolism are: the conversion of the energy in food to energy available to run cellular processes; the conversion of food to building blocks for protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, res ...
s, lipids, nucleic acids, and some carbohydrate
In organic chemistry, a carbohydrate () is a biomolecule consisting of carbon (C), hydrogen (H) and oxygen (O) atoms, usually with a hydrogen–oxygen atom ratio of 2:1 (as in water) and thus with the empirical formula (where ''m'' may or m ...
s; and the elimination of metabolic wastes. These enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products ...
-catalyzed reactions allow organisms to grow and reproduce, maintain their structures, and respond to their environments. The word metabolism can also refer to the sum of all chemical reactions that occur in living organisms, including digestion
Digestion is the breakdown of large insoluble food molecules into small water-soluble food molecules so that they can be absorbed into the watery blood plasma. In certain organisms, these smaller substances are absorbed through the small intest ...
and the transportation of substances into and between different cells, in which case the above described set of reactions within the cells is called intermediary (or intermediate) metabolism.
Metabolic reactions may be categorized as '' catabolic'' – the ''breaking down'' of compounds (for example, of glucose to pyruvate by cellular respiration); or '' anabolic'' – the ''building up'' ( synthesis) of compounds (such as proteins, carbohydrates, lipids, and nucleic acids). Usually, catabolism releases energy, and anabolism consumes energy.
The chemical reactions of metabolism are organized into metabolic pathways, in which one chemical is transformed through a series of steps into another chemical, each step being facilitated by a specific enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products ...
. Enzymes are crucial to metabolism because they allow organisms to drive desirable reactions that require energy and will not occur by themselves, by coupling
A coupling is a device used to connect two shafts together at their ends for the purpose of transmitting power. The primary purpose of couplings is to join two pieces of rotating equipment while permitting some degree of misalignment or end mov ...
them to spontaneous reactions that release energy. Enzymes act as catalysts
Catalysis () is the process of increasing the rate of a chemical reaction by adding a substance known as a catalyst (). Catalysts are not consumed in the reaction and remain unchanged after it. If the reaction is rapid and the catalyst recyc ...
– they allow a reaction to proceed more rapidly – and they also allow the regulation
Regulation is the management of complex systems according to a set of rules and trends. In systems theory, these types of rules exist in various fields of biology and society, but the term has slightly different meanings according to context. Fo ...
of the rate of a metabolic reaction, for example in response to changes in the cell's environment or to signals
In signal processing, a signal is a function that conveys information about a phenomenon. Any quantity that can vary over space or time can be used as a signal to share messages between observers. The ''IEEE Transactions on Signal Processing'' ...
from other cells.
The metabolic system of a particular organism determines which substances it will find nutritious
Nutrition is the biochemical and physiological process by which an organism uses food to support its life. It provides organisms with nutrients, which can be metabolized to create energy and chemical structures. Failure to obtain sufficient nu ...
and which poisonous. For example, some prokaryotes use hydrogen sulfide as a nutrient, yet this gas is poisonous to animals. The basal metabolic rate of an organism is the measure of the amount of energy consumed by all of these chemical reactions.
A striking feature of metabolism is the similarity of the basic metabolic pathways among vastly different species. For example, the set of carboxylic acids that are best known as the intermediates in the citric acid cycle are present in all known organisms, being found in species as diverse as the unicellular
A unicellular organism, also known as a single-celled organism, is an organism that consists of a single cell, unlike a multicellular organism that consists of multiple cells. Organisms fall into two general categories: prokaryotic organisms and ...
bacterium '' Escherichia coli'' and huge multicellular organisms like elephant
Elephants are the largest existing land animals. Three living species are currently recognised: the African bush elephant, the African forest elephant, and the Asian elephant. They are the only surviving members of the family Elephantidae ...
s. These similarities in metabolic pathways are likely due to their early appearance in evolutionary history
The history of life on Earth traces the processes by which living and fossil organisms evolved, from the earliest emergence of life to present day. Earth formed about 4.5 billion years ago (abbreviated as ''Ga'', for ''gigaannum'') and ev ...
, and their retention is likely due to their efficacy. In various diseases, such as type II diabetes, metabolic syndrome, and cancer
Cancer is a group of diseases involving abnormal cell growth with the potential to invade or spread to other parts of the body. These contrast with benign tumors, which do not spread. Possible signs and symptoms include a lump, abnormal b ...
, normal metabolism is disrupted. The metabolism of cancer cells is also different from the metabolism of normal cells, and these differences can be used to find targets for therapeutic intervention in cancer.
Key biochemicals

 Most of the structures that make up animals, plants and microbes are made from four basic classes of
Most of the structures that make up animals, plants and microbes are made from four basic classes of molecule
A molecule is a group of two or more atoms held together by attractive forces known as chemical bonds; depending on context, the term may or may not include ions which satisfy this criterion. In quantum physics, organic chemistry, and bioche ...
s: amino acids, carbohydrate
In organic chemistry, a carbohydrate () is a biomolecule consisting of carbon (C), hydrogen (H) and oxygen (O) atoms, usually with a hydrogen–oxygen atom ratio of 2:1 (as in water) and thus with the empirical formula (where ''m'' may or m ...
s, nucleic acid and lipids (often called fat
In nutrition, biology, and chemistry, fat usually means any ester of fatty acids, or a mixture of such compounds, most commonly those that occur in living beings or in food.
The term often refers specifically to triglycerides (triple est ...
s). As these molecules are vital for life, metabolic reactions either focus on making these molecules during the construction of cells and tissues, or on breaking them down and using them to obtain energy, by their digestion. These biochemicals can be joined to make polymer
A polymer (; Greek '' poly-'', "many" + ''-mer'', "part")
is a substance or material consisting of very large molecules called macromolecules, composed of many repeating subunits. Due to their broad spectrum of properties, both synthetic a ...
s such as DNA and protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, res ...
s, essential macromolecules
A macromolecule is a very large molecule important to biophysical processes, such as a protein or nucleic acid. It is composed of thousands of covalently bonded atoms. Many macromolecules are polymers of smaller molecules called monomers. The ...
of life.
Amino acids and proteins
Proteins are made of amino acids arranged in a linear chain joined by peptide bonds. Many proteins areenzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products ...
s that catalyze
Catalysis () is the process of increasing the rate of a chemical reaction by adding a substance known as a catalyst (). Catalysts are not consumed in the reaction and remain unchanged after it. If the reaction is rapid and the catalyst recyc ...
the chemical reactions in metabolism. Other proteins have structural or mechanical functions, such as those that form the cytoskeleton
The cytoskeleton is a complex, dynamic network of interlinking protein filaments present in the cytoplasm of all cells, including those of bacteria and archaea. In eukaryotes, it extends from the cell nucleus to the cell membrane and is com ...
, a system of scaffolding
Scaffolding, also called scaffold or staging, is a temporary structure used to support a work crew and materials to aid in the construction, maintenance and repair of buildings, bridges and all other man-made structures. Scaffolds are widely use ...
that maintains the cell shape. Proteins are also important in cell signaling
In biology, cell signaling (cell signalling in British English) or cell communication is the ability of a cell to receive, process, and transmit signals with its environment and with itself. Cell signaling is a fundamental property of all cellula ...
, immune responses, cell adhesion, active transport
In cellular biology, ''active transport'' is the movement of molecules or ions across a cell membrane from a region of lower concentration to a region of higher concentration—against the concentration gradient. Active transport requires cellul ...
across membranes, and the cell cycle
The cell cycle, or cell-division cycle, is the series of events that take place in a cell that cause it to divide into two daughter cells. These events include the duplication of its DNA (DNA replication) and some of its organelles, and sub ...
. Amino acids also contribute to cellular energy metabolism by providing a carbon source for entry into the citric acid cycle ( tricarboxylic acid cycle), especially when a primary source of energy, such as glucose
Glucose is a simple sugar with the molecular formula . Glucose is overall the most abundant monosaccharide, a subcategory of carbohydrates. Glucose is mainly made by plants and most algae during photosynthesis from water and carbon dioxide, u ...
, is scarce, or when cells undergo metabolic stress.
Lipids
Lipids are the most diverse group of biochemicals. Their main structural uses are as part of biological membranes both internal and external, such as the cell membrane. Their chemical energy can also be used. Lipids are the polymers of fatty acids that contain a long, non-polar hydrocarbon chain with a small polar region containing oxygen. Lipids are usually defined as hydrophobic oramphipathic
An amphiphile (from the Greek αμφις amphis, both, and φιλíα philia, love, friendship), or amphipath, is a chemical compound possessing both hydrophilic (''water-loving'', polar) and lipophilic (''fat-loving'') properties. Such a compoun ...
biological molecules but will dissolve in organic solvent
A solvent (s) (from the Latin '' solvō'', "loosen, untie, solve") is a substance that dissolves a solute, resulting in a solution. A solvent is usually a liquid but can also be a solid, a gas, or a supercritical fluid. Water is a solvent for ...
s such as ethanol
Ethanol (abbr. EtOH; also called ethyl alcohol, grain alcohol, drinking alcohol, or simply alcohol) is an organic compound. It is an alcohol with the chemical formula . Its formula can be also written as or (an ethyl group linked to a ...
, benzene
Benzene is an organic chemical compound with the molecular formula C6H6. The benzene molecule is composed of six carbon atoms joined in a planar ring with one hydrogen atom attached to each. Because it contains only carbon and hydrogen atoms ...
or chloroform. The fat
In nutrition, biology, and chemistry, fat usually means any ester of fatty acids, or a mixture of such compounds, most commonly those that occur in living beings or in food.
The term often refers specifically to triglycerides (triple est ...
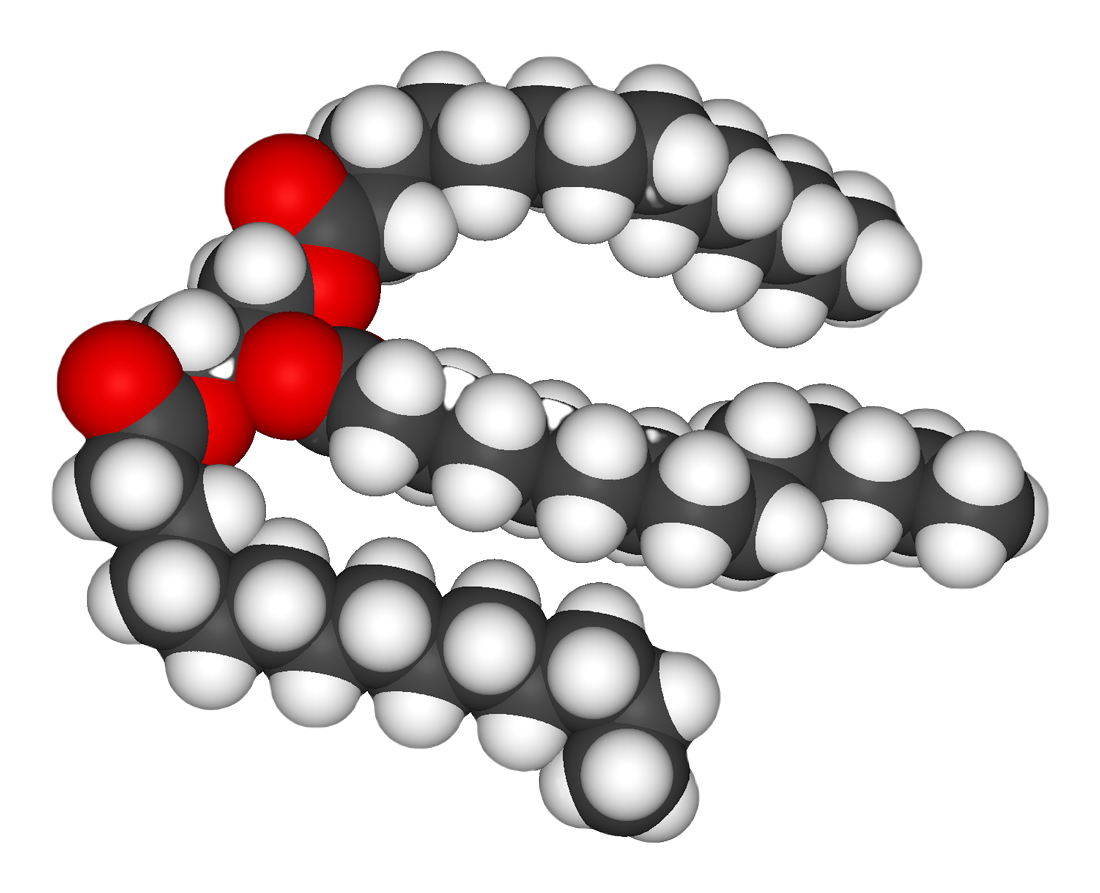
s are a large group of compounds that contain fatty acids and glycerol; a glycerol molecule attached to three fatty acids by ester linkages is called a triacylglyceride
A triglyceride (TG, triacylglycerol, TAG, or triacylglyceride) is an ester derived from glycerol and three fatty acids (from ''tri-'' and ''glyceride'').
Triglycerides are the main constituents of body fat in humans and other vertebrates, as w ...
. Several variations on this basic structure exist, including backbones such as sphingosine
Sphingosine (2-amino-4-trans-octadecene-1,3-diol) is an 18-carbon amino alcohol with an unsaturated hydrocarbon chain, which forms a primary part of sphingolipids, a class of cell membrane lipids that include sphingomyelin, an important phos ...
in sphingomyelin, and hydrophilic
A hydrophile is a molecule or other molecular entity that is attracted to water molecules and tends to be dissolved by water.Liddell, H.G. & Scott, R. (1940). ''A Greek-English Lexicon'' Oxford: Clarendon Press.
In contrast, hydrophobes are ...
groups such as phosphate
In chemistry, a phosphate is an anion, salt, functional group or ester derived from a phosphoric acid. It most commonly means orthophosphate, a derivative of orthophosphoric acid .
The phosphate or orthophosphate ion is derived from phosph ...
as in phospholipids. Steroids such as sterol are another major class of lipids.
Carbohydrates
 Carbohydrates are
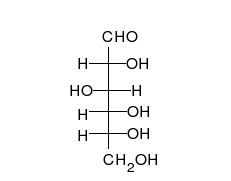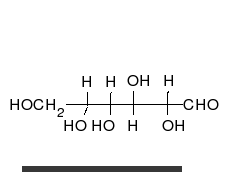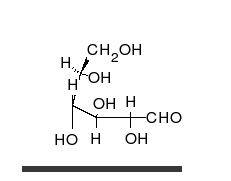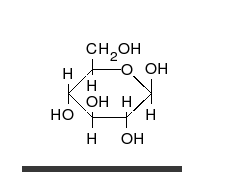
Carbohydrates are aldehyde
In organic chemistry, an aldehyde () is an organic compound containing a functional group with the structure . The functional group itself (without the "R" side chain) can be referred to as an aldehyde but can also be classified as a formyl grou ...
s or ketones, with many hydroxyl groups attached, that can exist as straight chains or rings. Carbohydrates are the most abundant biological molecules, and fill numerous roles, such as the storage and transport of energy ( starch, glycogen) and structural components (cellulose
Cellulose is an organic compound with the formula , a polysaccharide consisting of a linear chain of several hundred to many thousands of β(1→4) linked D-glucose units. Cellulose is an important structural component of the primary cell w ...
in plants, chitin in animals). The basic carbohydrate units are called monosaccharides and include galactose
Galactose (, '' galacto-'' + ''-ose'', "milk sugar"), sometimes abbreviated Gal, is a monosaccharide sugar that is about as sweet as glucose, and about 65% as sweet as sucrose. It is an aldohexose and a C-4 epimer of glucose. A galactose molecu ...
, fructose, and most importantly glucose
Glucose is a simple sugar with the molecular formula . Glucose is overall the most abundant monosaccharide, a subcategory of carbohydrates. Glucose is mainly made by plants and most algae during photosynthesis from water and carbon dioxide, u ...
. Monosaccharides can be linked together to form polysaccharides in almost limitless ways.
Nucleotides
The two nucleic acids, DNA and RNA, are polymers ofnucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecule ...
s. Each nucleotide is composed of a phosphate attached to a ribose or deoxyribose
Deoxyribose, or more precisely 2-deoxyribose, is a monosaccharide with idealized formula H−(C=O)−(CH2)−(CHOH)3−H. Its name indicates that it is a deoxy sugar, meaning that it is derived from the sugar ribose by loss of a hydroxy group. D ...
sugar group which is attached to a nitrogenous base
Nucleobases, also known as ''nitrogenous bases'' or often simply ''bases'', are nitrogen-containing biological compounds that form nucleosides, which, in turn, are components of nucleotides, with all of these monomers constituting the basic b ...
. Nucleic acids are critical for the storage and use of genetic information, and its interpretation through the processes of transcription and protein biosynthesis. This information is protected by DNA repair mechanisms and propagated through DNA replication. Many virus
A virus is a submicroscopic infectious agent that replicates only inside the living cells of an organism. Viruses infect all life forms, from animals and plants to microorganisms, including bacteria and archaea.
Since Dmitri Ivanovsk ...
es have an RNA genome, such as HIV
The human immunodeficiency viruses (HIV) are two species of ''Lentivirus'' (a subgroup of retrovirus) that infect humans. Over time, they cause acquired immunodeficiency syndrome (AIDS), a condition in which progressive failure of the immune ...
, which uses reverse transcription to create a DNA template from its viral RNA genome. RNA in ribozyme
Ribozymes (ribonucleic acid enzymes) are RNA molecules that have the ability to catalyze specific biochemical reactions, including RNA splicing in gene expression, similar to the action of protein enzymes. The 1982 discovery of ribozymes demons ...
s such as spliceosomes and ribosomes is similar to enzymes as it can catalyze chemical reactions. Individual nucleoside
Nucleosides are glycosylamines that can be thought of as nucleotides without a phosphate group. A nucleoside consists simply of a nucleobase (also termed a nitrogenous base) and a five-carbon sugar (ribose or 2'-deoxyribose) whereas a nucleoti ...
s are made by attaching a nucleobase to a ribose sugar. These bases are heterocyclic
A heterocyclic compound or ring structure is a cyclic compound that has atoms of at least two different elements as members of its ring(s). Heterocyclic chemistry is the branch of organic chemistry dealing with the synthesis, properties, and ...
rings containing nitrogen, classified as purines or pyrimidines. Nucleotides also act as coenzymes in metabolic-group-transfer reactions.
Coenzymes
human nutrition
Human nutrition deals with the provision of essential nutrients in food that are necessary to support human life and good health. Poor nutrition is a chronic problem often linked to poverty, food security, or a poor understanding of nutritiona ...
, most vitamins function as coenzymes after modification; for example, all water-soluble vitamins are phosphorylated or are coupled to nucleotides when they are used in cells. Nicotinamide adenine dinucleotide (NAD+), a derivative of vitamin B3 ( niacin), is an important coenzyme that acts as a hydrogen acceptor. Hundreds of separate types of dehydrogenase
A dehydrogenase is an enzyme belonging to the group of oxidoreductases that oxidizes a substrate by reducing an electron acceptor, usually NAD+/NADP+ or a flavin coenzyme such as FAD or FMN. Like all catalysts, they catalyze reverse as well as ...
s remove electrons from their substrates and reduce NAD+ into NADH. This reduced form of the coenzyme is then a substrate for any of the reductases in the cell that need to transfer hydrogen atoms to their substrates. Nicotinamide adenine dinucleotide exists in two related forms in the cell, NADH and NADPH. The NAD+/NADH form is more important in catabolic reactions, while NADP+/NADPH is used in anabolic reactions.

Mineral and cofactors
Inorganic elements play critical roles in metabolism; some are abundant (e.g. sodium and potassium) while others function at minute concentrations. About 99% of a human's body weight is made up of the elementscarbon
Carbon () is a chemical element with the symbol C and atomic number 6. It is nonmetallic and tetravalent—its atom making four electrons available to form covalent chemical bonds. It belongs to group 14 of the periodic table. Carbon mak ...
, nitrogen
Nitrogen is the chemical element with the symbol N and atomic number 7. Nitrogen is a nonmetal and the lightest member of group 15 of the periodic table, often called the pnictogens. It is a common element in the universe, estimated at se ...
, calcium, sodium, chlorine
Chlorine is a chemical element with the symbol Cl and atomic number 17. The second-lightest of the halogens, it appears between fluorine and bromine in the periodic table and its properties are mostly intermediate between them. Chlorine i ...
, potassium, hydrogen
Hydrogen is the chemical element with the symbol H and atomic number 1. Hydrogen is the lightest element. At standard conditions hydrogen is a gas of diatomic molecules having the formula . It is colorless, odorless, tasteless, non-toxic ...
, phosphorus, oxygen
Oxygen is the chemical element with the symbol O and atomic number 8. It is a member of the chalcogen group in the periodic table, a highly reactive nonmetal, and an oxidizing agent that readily forms oxides with most elements as ...
and sulfur. Organic compound
In chemistry, organic compounds are generally any chemical compounds that contain carbon-hydrogen or carbon-carbon bonds. Due to carbon's ability to catenate (form chains with other carbon atoms), millions of organic compounds are known. T ...
s (proteins, lipids and carbohydrates) contain the majority of the carbon and nitrogen; most of the oxygen and hydrogen is present as water.
The abundant inorganic elements act as electrolytes. The most important ions are sodium, potassium, calcium, magnesium, chloride
The chloride ion is the anion (negatively charged ion) Cl−. It is formed when the element chlorine (a halogen) gains an electron or when a compound such as hydrogen chloride is dissolved in water or other polar solvents. Chloride sa ...
, phosphate
In chemistry, a phosphate is an anion, salt, functional group or ester derived from a phosphoric acid. It most commonly means orthophosphate, a derivative of orthophosphoric acid .
The phosphate or orthophosphate ion is derived from phosph ...
and the organic ion bicarbonate. The maintenance of precise ion gradients across cell membranes maintains osmotic pressure and pH. Ions are also critical for nerve and muscle function, as action potentials in these tissues are produced by the exchange of electrolytes between the extracellular fluid
In cell biology, extracellular fluid (ECF) denotes all body fluid outside the cells of any multicellular organism. Total body water in healthy adults is about 60% (range 45 to 75%) of total body weight; women and the obese typically have a low ...
and the cell's fluid, the cytosol
The cytosol, also known as cytoplasmic matrix or groundplasm, is one of the liquids found inside cells ( intracellular fluid (ICF)). It is separated into compartments by membranes. For example, the mitochondrial matrix separates the mitochondri ...
. Electrolytes enter and leave cells through proteins in the cell membrane called ion channels. For example, muscle contraction depends upon the movement of calcium, sodium and potassium through ion channels in the cell membrane and T-tubule
T-tubules (transverse tubules) are extensions of the cell membrane that penetrate into the center of skeletal and cardiac muscle cells. With membranes that contain large concentrations of ion channels, transporters, and pumps, T-tubules permi ...
s.
Transition metal
In chemistry, a transition metal (or transition element) is a chemical element in the d-block of the periodic table (groups 3 to 12), though the elements of group 12 (and less often group 3) are sometimes excluded. They are the elements that ca ...
s are usually present as trace element
__NOTOC__
A trace element, also called minor element, is a chemical element whose concentration (or other measure of amount) is very low (a "trace amount"). They are classified into two groups: essential and non-essential. Essential trace elements ...
s in organisms, with zinc
Zinc is a chemical element with the symbol Zn and atomic number 30. Zinc is a slightly brittle metal at room temperature and has a shiny-greyish appearance when oxidation is removed. It is the first element in group 12 (IIB) of the periodi ...
and iron
Iron () is a chemical element with Symbol (chemistry), symbol Fe (from la, Wikt:ferrum, ferrum) and atomic number 26. It is a metal that belongs to the first transition series and group 8 element, group 8 of the periodic table. It is, Abundanc ...
being most abundant of those. Metal cofactors are bound tightly to specific sites in proteins; although enzyme cofactors can be modified during catalysis, they always return to their original state by the end of the reaction catalyzed. Metal micronutrients are taken up into organisms by specific transporters and bind to storage proteins such as ferritin or metallothionein when not in use.
Catabolism
Catabolism is the set of metabolic processes that break down large molecules. These include breaking down and oxidizing food molecules. The purpose of the catabolic reactions is to provide the energy and components needed by anabolic reactions which build molecules. The exact nature of these catabolic reactions differ from organism to organism, and organisms can be classified based on their sources of energy, hydrogen, and carbon (theirprimary nutritional groups
Primary nutritional groups are groups of organisms, divided in relation to the nutrition mode according to the sources of energy and carbon, needed for living, growth and reproduction. The sources of energy can be light or chemical compounds; the ...
), as shown in the table below. Organic molecules are used as a source of hydrogen atoms or electrons by organotroph
An organotroph is an organism that obtains hydrogen or electrons from organic substrates. This term is used in microbiology to classify and describe organisms based on how they obtain electrons for their respiration processes. Some organotrophs su ...
s, while lithotroph
Lithotrophs are a diverse group of organisms using an inorganic substrate (usually of mineral origin) to obtain reducing equivalents for use in biosynthesis (e.g., carbon dioxide fixation) or energy conservation (i.e., ATP production) via aerobi ...
s use inorganic substrates. Whereas phototrophs convert sunlight to chemical energy, chemotroph
A Chemotroph is an organism that obtains energy by the oxidation of electron donors in their environments. These molecules can be organic (chemoorganotrophs) or inorganic ( chemolithotrophs). The chemotroph designation is in contrast to phototr ...
s depend on redox reactions that involve the transfer of electrons from reduced donor molecules such as organic molecule
In chemistry, organic compounds are generally any chemical compounds that contain carbon-hydrogen or carbon-carbon bonds. Due to carbon's ability to catenate (form chains with other carbon atoms), millions of organic compounds are known. Th ...
s, hydrogen
Hydrogen is the chemical element with the symbol H and atomic number 1. Hydrogen is the lightest element. At standard conditions hydrogen is a gas of diatomic molecules having the formula . It is colorless, odorless, tasteless, non-toxic ...
, hydrogen sulfide or ferrous ions to oxygen
Oxygen is the chemical element with the symbol O and atomic number 8. It is a member of the chalcogen group in the periodic table, a highly reactive nonmetal, and an oxidizing agent that readily forms oxides with most elements as ...
, nitrate or sulfate
The sulfate or sulphate ion is a polyatomic anion with the empirical formula . Salts, acid derivatives, and peroxides of sulfate are widely used in industry. Sulfates occur widely in everyday life. Sulfates are salts of sulfuric acid and many ...
. In animals, these reactions involve complex organic molecule
In chemistry, organic compounds are generally any chemical compounds that contain carbon-hydrogen or carbon-carbon bonds. Due to carbon's ability to catenate (form chains with other carbon atoms), millions of organic compounds are known. Th ...
s that are broken down to simpler molecules, such as carbon dioxide
Carbon dioxide ( chemical formula ) is a chemical compound made up of molecules that each have one carbon atom covalently double bonded to two oxygen atoms. It is found in the gas state at room temperature. In the air, carbon dioxide is trans ...
and water. Photosynthetic organisms, such as plants and cyanobacteria, use similar electron-transfer reactions to store energy absorbed from sunlight.
The most common set of catabolic reactions in animals can be separated into three main stages. In the first stage, large organic molecules, such as protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, res ...
s, polysaccharides or lipids, are digested into their smaller components outside cells. Next, these smaller molecules are taken up by cells and converted to smaller molecules, usually acetyl coenzyme A (acetyl-CoA), which releases some energy. Finally, the acetyl group on acetyl-CoA is oxidized to water and carbon dioxide in the citric acid cycle and electron transport chain, releasing more energy while reducing the coenzyme nicotinamide adenine dinucleotide (NAD+) into NADH.
Digestion
Macromolecules cannot be directly processed by cells. Macromolecules must be broken into smaller units before they can be used in cell metabolism. Different classes of enzymes are used to digest these polymers. Thesedigestive enzyme
Digestive enzymes are a group of enzymes that break down polymeric macromolecules into their smaller building blocks, in order to facilitate their absorption into the cells of the body. Digestive enzymes are found in the digestive tracts of anima ...
s include proteases that digest proteins into amino acids, as well as glycoside hydrolases that digest polysaccharides into simple sugars known as monosaccharides
Monosaccharides (from Greek '' monos'': single, '' sacchar'': sugar), also called simple sugars, are the simplest forms of sugar and the most basic units ( monomers) from which all carbohydrates are built.
They are usually colorless, water ...
.
Microbes simply secrete digestive enzymes into their surroundings, while animals only secrete these enzymes from specialized cells in their guts
The gastrointestinal tract (GI tract, digestive tract, alimentary canal) is the tract or passageway of the digestive system that leads from the mouth to the anus. The GI tract contains all the major organs of the digestive system, in humans and ...
, including the stomach
The stomach is a muscular, hollow organ in the gastrointestinal tract of humans and many other animals, including several invertebrates. The stomach has a dilated structure and functions as a vital organ in the digestive system. The stomach i ...
and pancreas, and in salivary gland
The salivary glands in mammals are exocrine glands that produce saliva through a system of ducts. Humans have three paired major salivary glands ( parotid, submandibular, and sublingual), as well as hundreds of minor salivary glands. Salivary ...
s. The amino acids or sugars released by these extracellular enzymes are then pumped into cells by active transport
In cellular biology, ''active transport'' is the movement of molecules or ions across a cell membrane from a region of lower concentration to a region of higher concentration—against the concentration gradient. Active transport requires cellul ...
proteins.
Energy from organic compounds
Carbohydrate catabolism is the breakdown of carbohydrates into smaller units. Carbohydrates are usually taken into cells after they have been digested into monosaccharides. Once inside, the major route of breakdown is glycolysis, where sugars such asglucose
Glucose is a simple sugar with the molecular formula . Glucose is overall the most abundant monosaccharide, a subcategory of carbohydrates. Glucose is mainly made by plants and most algae during photosynthesis from water and carbon dioxide, u ...
and fructose are converted into pyruvate and some ATP is generated. Pyruvate is an intermediate in several metabolic pathways, but the majority is converted to acetyl-CoA through aerobic (with oxygen) glycolysis and fed into the citric acid cycle. Although some more ATP is generated in the citric acid cycle, the most important product is NADH, which is made from NAD+ as the acetyl-CoA is oxidized. This oxidation releases carbon dioxide
Carbon dioxide ( chemical formula ) is a chemical compound made up of molecules that each have one carbon atom covalently double bonded to two oxygen atoms. It is found in the gas state at room temperature. In the air, carbon dioxide is trans ...
as a waste product. In anaerobic conditions, glycolysis produces lactate, through the enzyme lactate dehydrogenase
Lactate dehydrogenase (LDH or LD) is an enzyme found in nearly all living cells. LDH catalyzes the conversion of lactate to pyruvate and back, as it converts NAD+ to NADH and back. A dehydrogenase is an enzyme that transfers a hydride from one ...
re-oxidizing NADH to NAD+ for re-use in glycolysis.
Fats are catabolized by hydrolysis to free fatty acids and glycerol. The glycerol enters glycolysis and the fatty acids are broken down by beta oxidation
In biochemistry and metabolism, beta-oxidation is the catabolic process by which fatty acid molecules are broken down in the cytosol in prokaryotes and in the mitochondria in eukaryotes to generate acetyl-CoA, which enters the citric acid cyc ...
to release acetyl-CoA, which then is fed into the citric acid cycle. Fatty acids release more energy upon oxidation than carbohydrates. Steroids are also broken down by some bacteria in a process similar to beta oxidation, and this breakdown process involves the release of significant amounts of acetyl-CoA, propionyl-CoA, and pyruvate, which can all be used by the cell for energy. ''M. tuberculosis'' can also grow on the lipid cholesterol as a sole source of carbon, and genes involved in the cholesterol-use pathway(s) have been validated as important during various stages of the infection lifecycle of ''M. tuberculosis''.
Amino acids are either used to synthesize proteins and other biomolecules, or oxidized to urea and carbon dioxide to produce energy. The oxidation pathway starts with the removal of the amino group by a transaminase
Transaminases or aminotransferases are enzymes that catalyze a transamination reaction between an amino acid and an α- keto acid. They are important in the synthesis of amino acids, which form proteins.
Function and mechanism
An amino acid ...
. The amino group is fed into the urea cycle, leaving a deaminated carbon skeleton in the form of a keto acid
In organic chemistry, keto acids or ketoacids (also called oxo acids or oxoacids) are organic compounds that contain a carboxylic acid group () and a ketone group ().Franz Dietrich Klingler, Wolfgang Ebertz "Oxocarboxylic Acids" in Ullmann's En ...
. Several of these keto acids are intermediates in the citric acid cycle, for example α-ketoglutarate Ketoglutaric acid or oxoglutaric acid, or its conjugate base, the carboxylate ketoglutarate or oxoglutarate, may refer to the following chemical compounds:
* α-Ketoglutaric acid, an intermediate in the citric acid cycle
The citric acid cycle (CA ...
formed by deamination of glutamate. The glucogenic amino acids can also be converted into glucose, through gluconeogenesis (discussed below).
Energy transformations
Oxidative phosphorylation
In oxidative phosphorylation, the electrons removed from organic molecules in areas such as the citric acid cycle are transferred to oxygen and the energy released is used to make ATP. This is done in eukaryotes by a series of proteins in the membranes of mitochondria called the electron transport chain. In prokaryotes, these proteins are found in the cell's inner membrane. These proteins use the energy from reduced molecules like NADH to pump protons across a membrane. Pumping protons out of the mitochondria creates a proton concentration difference across the membrane and generates an
Pumping protons out of the mitochondria creates a proton concentration difference across the membrane and generates an electrochemical gradient
An electrochemical gradient is a gradient of electrochemical potential, usually for an ion that can move across a membrane. The gradient consists of two parts, the chemical gradient, or difference in solute concentration across a membrane, and ...
. This force drives protons back into the mitochondrion through the base of an enzyme called ATP synthase. The flow of protons makes the stalk subunit rotate, causing the active site of the synthase domain to change shape and phosphorylate adenosine diphosphate – turning it into ATP.
Energy from inorganic compounds
Chemolithotroph
Lithotrophs are a diverse group of organisms using an inorganic substrate (usually of mineral origin) to obtain reducing equivalents for use in biosynthesis (e.g., carbon dioxide fixation) or energy conservation (i.e., ATP production) via aerobi ...
y is a type of metabolism found in prokaryotes where energy is obtained from the oxidation of inorganic compounds. These organisms can use hydrogen
Hydrogen is the chemical element with the symbol H and atomic number 1. Hydrogen is the lightest element. At standard conditions hydrogen is a gas of diatomic molecules having the formula . It is colorless, odorless, tasteless, non-toxic ...
, reduced sulfur compounds (such as sulfide, hydrogen sulfide and thiosulfate), ferrous iron (Fe(II)) or ammonia
Ammonia is an inorganic compound of nitrogen and hydrogen with the formula . A stable binary hydride, and the simplest pnictogen hydride, ammonia is a colourless gas with a distinct pungent smell. Biologically, it is a common nitrogenous wa ...
as sources of reducing power and they gain energy from the oxidation of these compounds. These microbial processes are important in global biogeochemical cycles such as acetogenesis, nitrification and denitrification and are critical for soil fertility.
Energy from light
The energy in sunlight is captured byplant
Plants are predominantly photosynthetic eukaryotes of the kingdom Plantae. Historically, the plant kingdom encompassed all living things that were not animals, and included algae and fungi; however, all current definitions of Plantae exclu ...
s, cyanobacteria, purple bacteria, green sulfur bacteria
The green sulfur bacteria are a phylum of obligately anaerobic photoautotrophic bacteria that metabolize sulfur.
Green sulfur bacteria are nonmotile (except ''Chloroherpeton thalassium'', which may glide) and capable of anoxygenic photosynthe ...
and some protists. This process is often coupled to the conversion of carbon dioxide into organic compounds, as part of photosynthesis, which is discussed below. The energy capture and carbon fixation systems can, however, operate separately in prokaryotes, as purple bacteria and green sulfur bacteria can use sunlight as a source of energy, while switching between carbon fixation and the fermentation of organic compounds.
In many organisms, the capture of solar energy is similar in principle to oxidative phosphorylation, as it involves the storage of energy as a proton concentration gradient. This proton motive force then drives ATP synthesis The electrons needed to drive this electron transport chain come from light-gathering proteins called photosynthetic reaction centre
A photosynthetic reaction center is a complex of several proteins, pigments and other co-factors that together execute the primary energy conversion reactions of photosynthesis. Molecular excitations, either originating directly from sunlight or t ...
s. Reaction centers are classified into two types depending on the nature of photosynthetic pigment present, with most photosynthetic bacteria only having one type, while plants and cyanobacteria have two.
In plants, algae, and cyanobacteria, photosystem II uses light energy to remove electrons from water, releasing oxygen as a waste product. The electrons then flow to the cytochrome b6f complex, which uses their energy to pump protons across the thylakoid membrane in the chloroplast. These protons move back through the membrane as they drive the ATP synthase, as before. The electrons then flow through photosystem I
Photosystem I (PSI, or plastocyanin–ferredoxin oxidoreductase) is one of two photosystems in the photosynthetic light reactions of algae, plants, and cyanobacteria. Photosystem I is an integral membrane protein complex that us ...
and can then be used to reduce the coenzyme NADP+.
Anabolism
Anabolism is the set of constructive metabolic processes where the energy released by catabolism is used to synthesize complex molecules. In general, the complex molecules that make up cellular structures are constructed step-by-step from smaller and simpler precursors. Anabolism involves three basic stages. First, the production of precursors such as amino acids, monosaccharides, isoprenoids andnucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecule ...
s, secondly, their activation into reactive forms using energy from ATP, and thirdly, the assembly of these precursors into complex molecules such as protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, res ...
s, polysaccharides, lipids and nucleic acids.
Anabolism in organisms can be different according to the source of constructed molecules in their cells. Autotroph
An autotroph or primary producer is an organism that produces complex organic compounds (such as carbohydrates, fats, and proteins) using carbon from simple substances such as carbon dioxide,Morris, J. et al. (2019). "Biology: How Life Wo ...
s such as plants can construct the complex organic molecules in their cells such as polysaccharides and proteins from simple molecules like carbon dioxide
Carbon dioxide ( chemical formula ) is a chemical compound made up of molecules that each have one carbon atom covalently double bonded to two oxygen atoms. It is found in the gas state at room temperature. In the air, carbon dioxide is trans ...
and water. Heterotrophs, on the other hand, require a source of more complex substances, such as monosaccharides and amino acids, to produce these complex molecules. Organisms can be further classified by ultimate source of their energy: photoautotrophs and photoheterotrophs obtain energy from light, whereas chemoautotrophs and chemoheterotrophs obtain energy from oxidation reactions.
Carbon fixation
 Photosynthesis is the synthesis of carbohydrates from sunlight and
Photosynthesis is the synthesis of carbohydrates from sunlight and carbon dioxide
Carbon dioxide ( chemical formula ) is a chemical compound made up of molecules that each have one carbon atom covalently double bonded to two oxygen atoms. It is found in the gas state at room temperature. In the air, carbon dioxide is trans ...
(CO2). In plants, cyanobacteria and algae, oxygenic photosynthesis splits water, with oxygen produced as a waste product. This process uses the ATP and NADPH produced by the photosynthetic reaction centre
A photosynthetic reaction center is a complex of several proteins, pigments and other co-factors that together execute the primary energy conversion reactions of photosynthesis. Molecular excitations, either originating directly from sunlight or t ...
s, as described above, to convert CO2 into glycerate 3-phosphate
3-Phosphoglyceric acid (3PG, 3-PGA, or PGA) is the conjugate acid of 3-phosphoglycerate or glycerate 3-phosphate (GP or G3P). This glycerate is a biochemically significant metabolic intermediate in both glycolysis and the Calvin-Benson cycle. The ...
, which can then be converted into glucose. This carbon-fixation reaction is carried out by the enzyme RuBisCO
Ribulose-1,5-bisphosphate carboxylase-oxygenase, commonly known by the abbreviations RuBisCo, rubisco, RuBPCase, or RuBPco, is an enzyme () involved in the first major step of carbon fixation, a process by which atmospheric carbon dioxide is con ...
as part of the Calvin – Benson cycle. Three types of photosynthesis occur in plants, C3 carbon fixation
carbon fixation is the most common of three metabolic pathways for carbon fixation in photosynthesis, along with C4 carbon fixation, and Crassulacean acid metabolism, CAM. This process converts carbon dioxide and ribulose bisphosphate (RuBP, a ...
, C4 carbon fixation and CAM photosynthesis. These differ by the route that carbon dioxide takes to the Calvin cycle, with C3 plants fixing CO2 directly, while C4 and CAM photosynthesis incorporate the CO2 into other compounds first, as adaptations to deal with intense sunlight and dry conditions.
In photosynthetic prokaryotes the mechanisms of carbon fixation are more diverse. Here, carbon dioxide can be fixed by the Calvin – Benson cycle, a reversed citric acid cycle, or the carboxylation of acetyl-CoA. Prokaryotic chemoautotrophs also fix CO2 through the Calvin–Benson cycle, but use energy from inorganic compounds to drive the reaction.
Carbohydrates and glycans
In carbohydrate anabolism, simple organic acids can be converted into monosaccharides such asglucose
Glucose is a simple sugar with the molecular formula . Glucose is overall the most abundant monosaccharide, a subcategory of carbohydrates. Glucose is mainly made by plants and most algae during photosynthesis from water and carbon dioxide, u ...
and then used to assemble polysaccharides such as starch. The generation of glucose
Glucose is a simple sugar with the molecular formula . Glucose is overall the most abundant monosaccharide, a subcategory of carbohydrates. Glucose is mainly made by plants and most algae during photosynthesis from water and carbon dioxide, u ...
from compounds like pyruvate, lactate, glycerol, glycerate 3-phosphate
3-Phosphoglyceric acid (3PG, 3-PGA, or PGA) is the conjugate acid of 3-phosphoglycerate or glycerate 3-phosphate (GP or G3P). This glycerate is a biochemically significant metabolic intermediate in both glycolysis and the Calvin-Benson cycle. The ...
and amino acids is called gluconeogenesis. Gluconeogenesis converts pyruvate to glucose-6-phosphate through a series of intermediates, many of which are shared with glycolysis. However, this pathway is not simply glycolysis run in reverse, as several steps are catalyzed by non-glycolytic enzymes. This is important as it allows the formation and breakdown of glucose to be regulated separately, and prevents both pathways from running simultaneously in a futile cycle
A futile cycle, also known as a substrate cycle, occurs when two metabolic pathways run simultaneously in opposite directions and have no overall effect other than to dissipate energy in the form of heat. The reason this cycle was called "futile" c ...
.
Although fat is a common way of storing energy, in vertebrate
Vertebrates () comprise all animal taxa within the subphylum Vertebrata () (chordates with backbones), including all mammals, birds, reptiles, amphibians, and fish. Vertebrates represent the overwhelming majority of the phylum Chordata, with c ...
s such as humans the fatty acids in these stores cannot be converted to glucose through gluconeogenesis as these organisms cannot convert acetyl-CoA into pyruvate; plants do, but animals do not, have the necessary enzymatic machinery. As a result, after long-term starvation, vertebrates need to produce ketone bodies from fatty acids to replace glucose in tissues such as the brain that cannot metabolize fatty acids. In other organisms such as plants and bacteria, this metabolic problem is solved using the glyoxylate cycle
The glyoxylate cycle, a variation of the tricarboxylic acid cycle, is an anabolic pathway occurring in plants, bacteria, protists, and fungi. The glyoxylate cycle centers on the conversion of acetyl-CoA to succinate for the synthesis of carbohydrat ...
, which bypasses the decarboxylation step in the citric acid cycle and allows the transformation of acetyl-CoA to oxaloacetate
Oxaloacetic acid (also known as oxalacetic acid or OAA) is a crystalline organic compound with the chemical formula HO2CC(O)CH2CO2H. Oxaloacetic acid, in the form of its conjugate base oxaloacetate, is a metabolic intermediate in many processes ...
, where it can be used for the production of glucose. Other than fat, glucose is stored in most tissues, as an energy resource available within the tissue through glycogenesis which was usually being used to maintained glucose level in blood.
Polysaccharides and glycans are made by the sequential addition of monosaccharides by glycosyltransferase
Glycosyltransferases (GTFs, Gtfs) are enzymes (EC 2.4) that establish natural glycosidic linkages. They catalyze the transfer of saccharide moieties from an activated nucleotide sugar (also known as the "glycosyl donor") to a nucleophilic glyco ...
from a reactive sugar-phosphate donor such as uridine diphosphate glucose (UDP-Glc) to an acceptor hydroxyl group on the growing polysaccharide. As any of the hydroxyl groups on the ring of the substrate can be acceptors, the polysaccharides produced can have straight or branched structures. The polysaccharides produced can have structural or metabolic functions themselves, or be transferred to lipids and proteins by enzymes called oligosaccharyltransferase
Oligosaccharyltransferase or OST () is a membrane protein complex that transfers a 14-sugar oligosaccharide from dolichol to nascent protein. It is a type of glycosyltransferase. The sugar Glc3Man9GlcNAc2 (where Glc=Glucose, Man=Mannose, and GlcN ...
s.
Fatty acids, isoprenoids and sterol
isopentenyl pyrophosphate
Isopentenyl pyrophosphate (IPP, isopentenyl diphosphate, or IDP) is an isoprenoid precursor. IPP is an intermediate in the classical, HMG-CoA reductase pathway (commonly called the mevalonate pathway) and in the ''non-mevalonate'' MEP pathway of i ...
and dimethylallyl pyrophosphate. These precursors can be made in different ways. In animals and archaea, the mevalonate pathway
The mevalonate pathway, also known as the isoprenoid pathway or HMG-CoA reductase pathway is an essential metabolic pathway present in eukaryotes, archaea, and some bacteria. The pathway produces two five-carbon building blocks called isopentenyl ...
produces these compounds from acetyl-CoA, while in plants and bacteria the non-mevalonate pathway uses pyruvate and glyceraldehyde 3-phosphate
Glyceraldehyde 3-phosphate, also known as triose phosphate or 3-phosphoglyceraldehyde and abbreviated as G3P, GA3P, GADP, GAP, TP, GALP or PGAL, is a metabolite that occurs as an intermediate in several central pathways of all organisms.Nelson, D ...
as substrates. One important reaction that uses these activated isoprene donors is sterol biosynthesis. Here, the isoprene units are joined to make squalene
Squalene is an organic compound. It is a triterpenoid with the formula C30H50. It is a colourless oil, although impure samples appear yellow. It was originally obtained from shark liver oil (hence its name, as ''Squalus'' is a genus of sharks). A ...
and then folded up and formed into a set of rings to make lanosterol
Lanosterol is a tetracyclic triterpenoid and is the compound from which all animal and fungal steroids are derived. By contrast plant steroids are produced via cycloartenol.
Role in biosynthesis of other steroids
Elaboration of lanosterol under en ...
. Lanosterol can then be converted into other sterols such as cholesterol and ergosterol.
Proteins
Organisms vary in their ability to synthesize the 20 common amino acids. Most bacteria and plants can synthesize all twenty, but mammals can only synthesize eleven nonessential amino acids, so nineessential amino acid
An essential amino acid, or indispensable amino acid, is an amino acid that cannot be synthesized from scratch by the organism fast enough to supply its demand, and must therefore come from the diet. Of the 21 amino acids common to all life form ...
s must be obtained from food. Some simple parasites, such as the bacteria ''Mycoplasma pneumoniae
''Mycoplasma pneumoniae'' is a very small bacterium in the class Mollicutes. It is a human pathogen that causes the disease mycoplasma pneumonia, a form of atypical bacterial pneumonia related to cold agglutinin disease. ''M. pneumoniae'' is c ...
'', lack all amino acid synthesis and take their amino acids directly from their hosts. All amino acids are synthesized from intermediates in glycolysis, the citric acid cycle, or the pentose phosphate pathway. Nitrogen is provided by glutamate and glutamine
Glutamine (symbol Gln or Q) is an α-amino acid that is used in the biosynthesis of proteins. Its side chain is similar to that of glutamic acid, except the carboxylic acid group is replaced by an amide. It is classified as a charge-neutral ...
. Nonessensial amino acid synthesis depends on the formation of the appropriate alpha-keto acid, which is then transaminated to form an amino acid.
Amino acids are made into proteins by being joined in a chain of peptide bonds. Each different protein has a unique sequence of amino acid residues: this is its primary structure. Just as the letters of the alphabet can be combined to form an almost endless variety of words, amino acids can be linked in varying sequences to form a huge variety of proteins. Proteins are made from amino acids that have been activated by attachment to a transfer RNA molecule through an ester bond. This aminoacyl-tRNA
Aminoacyl-tRNA (also aa-tRNA or charged tRNA) is tRNA to which its cognate amino acid is chemically bonded (charged). The aa-tRNA, along with particular elongation factors, deliver the amino acid to the ribosome for incorporation into the polypept ...
precursor is produced in an ATP-dependent reaction carried out by an aminoacyl tRNA synthetase
An aminoacyl-tRNA synthetase (aaRS or ARS), also called tRNA-ligase, is an enzyme that attaches the appropriate amino acid onto its corresponding tRNA. It does so by catalyzing the transesterification of a specific cognate amino acid or its pre ...
. This aminoacyl-tRNA is then a substrate for the ribosome, which joins the amino acid onto the elongating protein chain, using the sequence information in a messenger RNA.
Nucleotide synthesis and salvage
Nucleotides are made from amino acids, carbon dioxide and formic acid in pathways that require large amounts of metabolic energy. Consequently, most organisms have efficient systems to salvage preformed nucleotides. Purines are synthesized asnucleoside
Nucleosides are glycosylamines that can be thought of as nucleotides without a phosphate group. A nucleoside consists simply of a nucleobase (also termed a nitrogenous base) and a five-carbon sugar (ribose or 2'-deoxyribose) whereas a nucleoti ...
s (bases attached to ribose). Both adenine and guanine are made from the precursor nucleoside inosine
Inosine is a nucleoside that is formed when hypoxanthine is attached to a ribose ring (also known as a ribofuranose) via a β-N9- glycosidic bond. It was discovered in 1965 in analysis of RNA transferase.
Inosine is commonly found in tRNAs and is ...
monophosphate, which is synthesized using atoms from the amino acids glycine
Glycine (symbol Gly or G; ) is an amino acid that has a single hydrogen atom as its side chain. It is the simplest stable amino acid ( carbamic acid is unstable), with the chemical formula NH2‐ CH2‐ COOH. Glycine is one of the proteinog ...
, glutamine
Glutamine (symbol Gln or Q) is an α-amino acid that is used in the biosynthesis of proteins. Its side chain is similar to that of glutamic acid, except the carboxylic acid group is replaced by an amide. It is classified as a charge-neutral ...
, and aspartic acid, as well as formate transferred from the coenzyme tetrahydrofolate
Tetrahydrofolic acid (THFA), or tetrahydrofolate, is a folic acid derivative.
Metabolism
Human synthesis
Tetrahydrofolic acid is produced from dihydrofolic acid by dihydrofolate reductase. This reaction is inhibited by methotrexate.
It is co ...
. Pyrimidines, on the other hand, are synthesized from the base orotate
Orotic acid is a pyrimidinedione and a carboxylic acid. Historically, it was believed to be part of the vitamin B complex and was called vitamin B13, but it is now known that it is not a vitamin.
The compound is synthesized in the body via a m ...
, which is formed from glutamine and aspartate.
Xenobiotics and redox metabolism
All organisms are constantly exposed to compounds that they cannot use as foods and that would be harmful if they accumulated in cells, as they have no metabolic function. These potentially damaging compounds are called xenobiotics. Xenobiotics such as synthetic drugs, natural poisons and antibiotics are detoxified by a set of xenobiotic-metabolizing enzymes. In humans, these include cytochrome P450 oxidases, UDP-glucuronosyltransferases, and glutathione ''S''-transferases. This system of enzymes acts in three stages to firstly oxidize the xenobiotic (phase I) and then conjugate water-soluble groups onto the molecule (phase II). The modified water-soluble xenobiotic can then be pumped out of cells and in multicellular organisms may be further metabolized before being excreted (phase III). In ecology, these reactions are particularly important in microbial biodegradation of pollutants and thebioremediation
Bioremediation broadly refers to any process wherein a biological system (typically bacteria, microalgae, fungi, and plants), living or dead, is employed for removing environmental pollutants from air, water, soil, flue gasses, industrial effluent ...
of contaminated land and oil spills. Many of these microbial reactions are shared with multicellular organisms, but due to the incredible diversity of types of microbes these organisms are able to deal with a far wider range of xenobiotics than multicellular organisms, and can degrade even persistent organic pollutant
Persistent organic pollutants (POPs), sometimes known as "forever chemicals", are organic compounds that are resistant to environmental degradation through chemical, biological, and photolytic processes. They are toxic chemicals that adversel ...
s such as organochloride
An organochloride, organochlorine compound, chlorocarbon, or chlorinated hydrocarbon is an organic compound containing at least one covalently bonded atom of chlorine. The chloroalkane class (alkanes with one or more hydrogens substituted by chlo ...
compounds.
A related problem for aerobic organism
Aerobic means "requiring air," in which "air" usually means oxygen.
Aerobic may also refer to
* Aerobic exercise, prolonged exercise of moderate intensity
* Aerobics
Aerobics is a form of physical exercise that combines rhythmic aerobic exe ...
s is oxidative stress
Oxidative stress reflects an imbalance between the systemic manifestation of reactive oxygen species and a biological system's ability to readily detoxify the reactive intermediates or to repair the resulting damage. Disturbances in the normal ...
. Here, processes including oxidative phosphorylation and the formation of disulfide bond
In biochemistry, a disulfide (or disulphide in British English) refers to a functional group with the structure . The linkage is also called an SS-bond or sometimes a disulfide bridge and is usually derived by the coupling of two thiol groups. In ...
s during protein folding produce reactive oxygen species
In chemistry, reactive oxygen species (ROS) are highly reactive chemicals formed from diatomic oxygen (). Examples of ROS include peroxides, superoxide, hydroxyl radical, singlet oxygen, and alpha-oxygen.
The reduction of molecular oxygen () p ...
such as hydrogen peroxide. These damaging oxidants are removed by antioxidant metabolites such as glutathione
Glutathione (GSH, ) is an antioxidant in plants, animals, fungi, and some bacteria and archaea. Glutathione is capable of preventing damage to important cellular components caused by sources such as reactive oxygen species, free radicals, pe ...
and enzymes such as catalases and peroxidase
Peroxidases or peroxide reductases ( EC numberbr>1.11.1.x are a large group of enzymes which play a role in various biological processes. They are named after the fact that they commonly break up peroxides.
Functionality
Peroxidases typically ca ...
s.
Thermodynamics of living organisms
Living organisms must obey thelaws of thermodynamics
The laws of thermodynamics are a set of scientific laws which define a group of physical quantities, such as temperature, energy, and entropy, that characterize thermodynamic systems in thermodynamic equilibrium. The laws also use various paramet ...
, which describe the transfer of heat and work
Work may refer to:
* Work (human activity), intentional activity people perform to support themselves, others, or the community
** Manual labour, physical work done by humans
** House work, housework, or homemaking
** Working animal, an animal t ...
. The second law of thermodynamics states that in any isolated system, the amount of entropy (disorder) cannot decrease. Although living organisms' amazing complexity appears to contradict this law, life is possible as all organisms are open systems that exchange matter and energy with their surroundings. Living systems are not in equilibrium, but instead are dissipative system
A dissipative system is a thermodynamically open system which is operating out of, and often far from, thermodynamic equilibrium in an environment with which it exchanges energy and matter. A tornado may be thought of as a dissipative system. Dis ...
s that maintain their state of high complexity by causing a larger increase in the entropy of their environments. The metabolism of a cell achieves this by coupling the spontaneous process In thermodynamics, a spontaneous process is a process which occurs without any external input to the system. A more technical definition is the time-evolution of a system in which it releases free energy and it moves to a lower, more thermodynamic ...
es of catabolism to the non-spontaneous processes of anabolism. In thermodynamic terms, metabolism maintains order by creating disorder.
Regulation and control
As the environments of most organisms are constantly changing, the reactions of metabolism must be finelyregulated
Regulation is the management of complex systems according to a set of rules and trends. In systems theory, these types of rules exist in various fields of biology and society, but the term has slightly different meanings according to context. Fo ...
to maintain a constant set of conditions within cells, a condition called homeostasis
In biology, homeostasis (British also homoeostasis) (/hɒmɪə(ʊ)ˈsteɪsɪs/) is the state of steady internal, physical, and chemical conditions maintained by living systems. This is the condition of optimal functioning for the organism and ...
. Metabolic regulation also allows organisms to respond to signals and interact actively with their environments. Two closely linked concepts are important for understanding how metabolic pathways are controlled. Firstly, the ''regulation'' of an enzyme in a pathway is how its activity is increased and decreased in response to signals. Secondly, the ''control'' exerted by this enzyme is the effect that these changes in its activity have on the overall rate of the pathway (the flux through the pathway). For example, an enzyme may show large changes in activity (''i.e.'' it is highly regulated) but if these changes have little effect on the flux of a metabolic pathway, then this enzyme is not involved in the control of the pathway.
growth factor
A growth factor is a naturally occurring substance capable of stimulating cell proliferation, wound healing, and occasionally cellular differentiation. Usually it is a secreted protein or a steroid hormone. Growth factors are important for regul ...
s and are detected by specific receptors
Receptor may refer to:
*Sensory receptor, in physiology, any structure which, on receiving environmental stimuli, produces an informative nerve impulse
*Receptor (biochemistry), in biochemistry, a protein molecule that receives and responds to a n ...
on the cell surface. These signals are then transmitted inside the cell by second messenger system
Second messengers are intracellular signaling molecules released by the cell in response to exposure to extracellular signaling molecules—the first messengers. (Intercellular signals, a non-local form or cell signaling, encompassing both first me ...
s that often involved the phosphorylation of proteins.
A very well understood example of extrinsic control is the regulation of glucose metabolism by the hormone insulin. Insulin is produced in response to rises in blood glucose levels. Binding of the hormone to insulin receptors on cells then activates a cascade of protein kinase
A protein kinase is a kinase which selectively modifies other proteins by covalently adding phosphates to them (phosphorylation) as opposed to kinases which modify lipids, carbohydrates, or other molecules. Phosphorylation usually results in a fu ...
s that cause the cells to take up glucose and convert it into storage molecules such as fatty acids and glycogen. The metabolism of glycogen is controlled by activity of phosphorylase
In biochemistry, phosphorylases are enzymes that catalyze the addition of a phosphate group from an inorganic phosphate (phosphate+hydrogen) to an acceptor.
:A-B + P A + P-B
They include allosteric enzymes that catalyze the production of gluco ...
, the enzyme that breaks down glycogen, and glycogen synthase, the enzyme that makes it. These enzymes are regulated in a reciprocal fashion, with phosphorylation inhibiting glycogen synthase, but activating phosphorylase. Insulin causes glycogen synthesis by activating protein phosphatases and producing a decrease in the phosphorylation of these enzymes.
Evolution
last universal common ancestor
The last universal common ancestor (LUCA) is the most recent population from which all organisms now living on Earth share common descent—the most recent common ancestor of all current life on Earth. This includes all cellular organisms; th ...
. This universal ancestral cell was prokaryotic
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
and probably a methanogen that had extensive amino acid, nucleotide, carbohydrate and lipid metabolism. The retention of these ancient pathways during later evolution
Evolution is change in the heritable characteristics of biological populations over successive generations. These characteristics are the expressions of genes, which are passed on from parent to offspring during reproduction. Variation ...
may be the result of these reactions having been an optimal solution to their particular metabolic problems, with pathways such as glycolysis and the citric acid cycle producing their end products highly efficiently and in a minimal number of steps. The first pathways of enzyme-based metabolism may have been parts of purine nucleotide metabolism, while previous metabolic pathways were a part of the ancient RNA world
The RNA world is a hypothetical stage in the evolutionary history of life on Earth, in which self-replicating RNA molecules proliferated before the evolution of DNA and proteins. The term also refers to the hypothesis that posits the existen ...
.
Many models have been proposed to describe the mechanisms by which novel metabolic pathways evolve. These include the sequential addition of novel enzymes to a short ancestral pathway, the duplication and then divergence of entire pathways as well as the recruitment of pre-existing enzymes and their assembly into a novel reaction pathway. The relative importance of these mechanisms is unclear, but genomic studies have shown that enzymes in a pathway are likely to have a shared ancestry, suggesting that many pathways have evolved in a step-by-step fashion with novel functions created from pre-existing steps in the pathway. An alternative model comes from studies that trace the evolution of proteins' structures in metabolic networks, this has suggested that enzymes are pervasively recruited, borrowing enzymes to perform similar functions in different metabolic pathways (evident in the MANET database
The Molecular Ancestry Network (MANET) database is a bioinformatics database that maps evolutionary relationships of protein architectures directly onto biological networks. It was originally developed by Hee Shin Kim, Jay E. Mittenthal and Gustav ...
) These recruitment processes result in an evolutionary enzymatic mosaic. A third possibility is that some parts of metabolism might exist as "modules" that can be reused in different pathways and perform similar functions on different molecules.
As well as the evolution of new metabolic pathways, evolution can also cause the loss of metabolic functions. For example, in some parasites metabolic processes that are not essential for survival are lost and preformed amino acids, nucleotides and carbohydrates may instead be scavenged from the host. Similar reduced metabolic capabilities are seen in endosymbiotic
An ''endosymbiont'' or ''endobiont'' is any organism that lives within the body or cells of another organism most often, though not always, in a mutualistic relationship.
(The term endosymbiosis is from the Greek: ἔνδον ''endon'' "within ...
organisms.
Investigation and manipulation
 Classically, metabolism is studied by a reductionist approach that focuses on a single metabolic pathway. Particularly valuable is the use of radioactive tracers at the whole-organism, tissue and cellular levels, which define the paths from precursors to final products by identifying radioactively labelled intermediates and products. The enzymes that catalyze these chemical reactions can then be purified and their kinetics and responses to inhibitors investigated. A parallel approach is to identify the small molecules in a cell or tissue; the complete set of these molecules is called the
Classically, metabolism is studied by a reductionist approach that focuses on a single metabolic pathway. Particularly valuable is the use of radioactive tracers at the whole-organism, tissue and cellular levels, which define the paths from precursors to final products by identifying radioactively labelled intermediates and products. The enzymes that catalyze these chemical reactions can then be purified and their kinetics and responses to inhibitors investigated. A parallel approach is to identify the small molecules in a cell or tissue; the complete set of these molecules is called the metabolome
The metabolome refers to the complete set of Small molecule, small-molecule chemicals found within a biological sample. The biological sample can be a Cell (biology), cell, a cellular organelle, an Organ (anatomy), organ, a Tissue (biology), tiss ...
. Overall, these studies give a good view of the structure and function of simple metabolic pathways, but are inadequate when applied to more complex systems such as the metabolism of a complete cell.
An idea of the complexity of the metabolic network
A metabolic network is the complete set of metabolic and physical processes that determine the physiological and biochemical properties of a cell. As such, these networks comprise the chemical reactions of metabolism, the metabolic pathways, as w ...
s in cells that contain thousands of different enzymes is given by the figure showing the interactions between just 43 proteins and 40 metabolites to the right: the sequences of genomes provide lists containing anything up to 26.500 genes. However, it is now possible to use this genomic data to reconstruct complete networks of biochemical reactions and produce more holistic
Holism () is the idea that various systems (e.g. physical, biological, social) should be viewed as wholes, not merely as a collection of parts. The term "holism" was coined by Jan Smuts in his 1926 book '' Holism and Evolution''."holism, n." OED On ...
mathematical models that may explain and predict their behavior. These models are especially powerful when used to integrate the pathway and metabolite data obtained through classical methods with data on gene expression from proteomic
Proteomics is the large-scale study of proteins. Proteins are vital parts of living organisms, with many functions such as the formation of structural fibers of muscle tissue, enzymatic digestion of food, or synthesis and replication of DNA. In ...
and DNA microarray studies. Using these techniques, a model of human metabolism has now been produced, which will guide future drug discovery and biochemical research. These models are now used in network analysis, to classify human diseases into groups that share common proteins or metabolites.
Bacterial metabolic networks are a striking example of bow-tie
The bow tie is a type of necktie. A modern bow tie is tied using a common shoelace knot, which is also called the bow knot for that reason. It consists of a ribbon of fabric tied around the collar of a shirt in a symmetrical manner so that t ...
organization, an architecture able to input a wide range of nutrients and produce a large variety of products and complex macromolecules using a relatively few intermediate common currencies.
A major technological application of this information is metabolic engineering. Here, organisms such as yeast, plants or bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one Cell (biology), biological cell. They constitute a large domain (biology), domain of prokaryotic microorganisms. Typically a few micrometr ...
are genetically modified to make them more useful in biotechnology and aid the production of drugs such as antibiotics or industrial chemicals such as 1,3-propanediol and shikimic acid
Shikimic acid, more commonly known as its anionic form shikimate, is a cyclohexene, a cyclitol and a cyclohexanecarboxylic acid. It is an important biochemical metabolite in plants and microorganisms. Its name comes from the Japanese flower ''shik ...
. These genetic modifications usually aim to reduce the amount of energy used to produce the product, increase yields and reduce the production of wastes.
History
The term ''metabolism'' is derived from French "métabolisme" orAncient Greek
Ancient Greek includes the forms of the Greek language used in ancient Greece and the ancient world from around 1500 BC to 300 BC. It is often roughly divided into the following periods: Mycenaean Greek (), Dark Ages (), the Archaic p ...
μεταβολή – "Metabole" for "a change" which derived from μεταβάλλ –"Metaballein" means "To change"

Greek philosophy
Aristotle
Aristotle (; grc-gre, Ἀριστοτέλης ''Aristotélēs'', ; 384–322 BC) was a Greek philosopher and polymath during the Classical period in Ancient Greece. Taught by Plato, he was the founder of the Peripatetic school of ph ...
's '' The Parts of Animals'' sets out enough details of his views on metabolism for an open flow model to be made. He believed that at each stage of the process, materials from food were transformed, with heat being released as the classical element of fire, and residual materials being excreted as urine, bile, or faeces.
Ibn al-Nafis described metabolism in his 1260 AD work titled Al-Risalah al-Kamiliyyah fil Siera al-Nabawiyyah (The Treatise of Kamil on the Prophet's Biography) which included the following phrase "Both the body and its parts are in a continuous state of dissolution and nourishment, so they are inevitably undergoing permanent change."
Application of the scientific method and Modern metabolic theories
The history of the scientific study of metabolism spans several centuries and has moved from examining whole animals in early studies, to examining individual metabolic reactions in modern biochemistry. The first controlled experiments in human metabolism were published by Santorio Santorio in 1614 in his book ''Ars de statica medicina''. He described how he weighed himself before and after eating,sleep
Sleep is a sedentary state of mind and body. It is characterized by altered consciousness, relatively inhibited sensory activity, reduced muscle activity and reduced interactions with surroundings. It is distinguished from wakefulness by a de ...
, working, sex, fasting, drinking, and excreting. He found that most of the food he took in was lost through what he called " insensible perspiration".
 In these early studies, the mechanisms of these metabolic processes had not been identified and a vital force was thought to animate living tissue. In the 19th century, when studying the fermentation of sugar to alcohol by yeast, Louis Pasteur concluded that fermentation was catalyzed by substances within the yeast cells he called "ferments". He wrote that "alcoholic fermentation is an act correlated with the life and organization of the yeast cells, not with the death or putrefaction of the cells." This discovery, along with the publication by Friedrich Wöhler in 1828 of a paper on the chemical synthesis of urea, and is notable for being the first organic compound prepared from wholly inorganic precursors. This proved that the organic compounds and chemical reactions found in cells were no different in principle than any other part of chemistry.
It was the discovery of
In these early studies, the mechanisms of these metabolic processes had not been identified and a vital force was thought to animate living tissue. In the 19th century, when studying the fermentation of sugar to alcohol by yeast, Louis Pasteur concluded that fermentation was catalyzed by substances within the yeast cells he called "ferments". He wrote that "alcoholic fermentation is an act correlated with the life and organization of the yeast cells, not with the death or putrefaction of the cells." This discovery, along with the publication by Friedrich Wöhler in 1828 of a paper on the chemical synthesis of urea, and is notable for being the first organic compound prepared from wholly inorganic precursors. This proved that the organic compounds and chemical reactions found in cells were no different in principle than any other part of chemistry.
It was the discovery of enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products ...
s at the beginning of the 20th century by Eduard Buchner
Eduard Buchner (; 20 May 1860 – 13 August 1917) was a German chemist and zymologist, awarded the 1907 Nobel Prize in Chemistry for his work on fermentation.
Biography
Early years
Buchner was born in Munich to a physician and Doctor Extraor ...
that separated the study of the chemical reactions of metabolism from the biological study of cells, and marked the beginnings of biochemistry
Biochemistry or biological chemistry is the study of chemical processes within and relating to living organisms. A sub-discipline of both chemistry and biology, biochemistry may be divided into three fields: structural biology, enzymology and ...
. The mass of biochemical knowledge grew rapidly throughout the early 20th century. One of the most prolific of these modern biochemists was Hans Krebs who made huge contributions to the study of metabolism. He discovered the urea cycle and later, working with Hans Kornberg
Sir Hans Leo Kornberg, FRS (14 January 1928 – 16 December 2019) was a British-American biochemist. He was Sir William Dunn Professor of Biochemistry in the University of Cambridge from 1975 to 1995, and Master of Christ's College, Cambridge ...
, the citric acid cycle and the glyoxylate cycle.
See also
* * * * * * , a "metabolism first" theory of theorigin of life
In biology, abiogenesis (from a- 'not' + Greek bios 'life' + genesis 'origin') or the origin of life is the natural process by which life has arisen from non-living matter, such as simple organic compounds. The prevailing scientific hypothes ...
*
* Microphysiometry Microphysiometry is the ''in vitro'' measurement of the functions and activities of life or of living matter (as organs, tissues, or cells) and of the physical and chemical phenomena involved on a very small (micrometer) scale. The term microphysiom ...
*
*
*
*
*
*
*
*
* Oncometabolism
*
*
References
Further reading
Introductory * * * Advanced * * * * * * *External links
General informationThe Biochemistry of Metabolism
Sparknotes SAT biochemistry
Overview of biochemistry. School level.
Undergraduate-level guide to molecular biology. Human metabolism
Guide to human metabolic pathways. School level.
THE Medical Biochemistry Page
Comprehensive resource on human metabolism. Databases
Flow Chart of Metabolic Pathways
at ExPASy
IUBMB-Nicholson Metabolic Pathways Chart
SuperCYP: Database for Drug-Cytochrome-Metabolism
Metabolic pathways
* {{Authority control, state=collapsed Underwater diving physiology