G-quadruplex on:
[Wikipedia]
[Google]
[Amazon]
 In molecular biology, G-quadruplex secondary structures (G4) are formed in nucleic acids by sequences that are rich in
In molecular biology, G-quadruplex secondary structures (G4) are formed in nucleic acids by sequences that are rich in
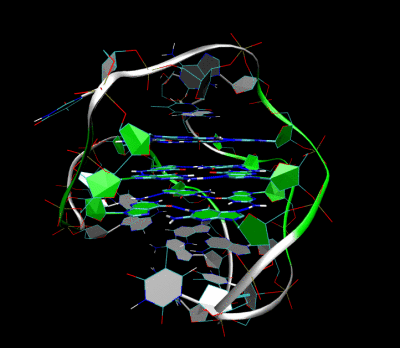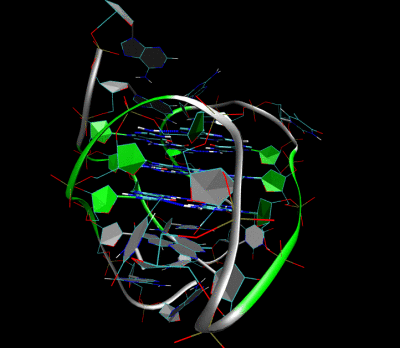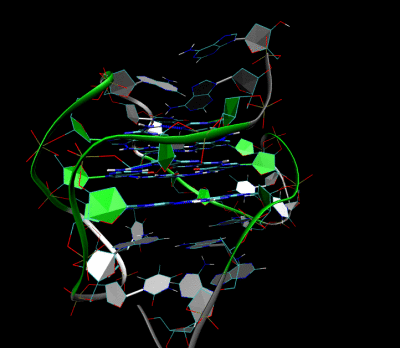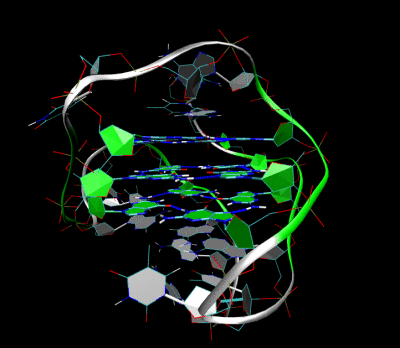
Nanopore and Aptamer Biosensor group
G-Quadruplex World
– a website to discuss publications and other information of interest to those working in the field of G-quadruplexes
Greglist
– a database listing potential G-quadruplex regulated genes
Database on Quadruplex information: QuadBase from IGIB
GRSDB
a database of G-quadruplexes near RNA processing sites.
GRS_UTRdb
- a database of G-quadruplexes in the UTRs.
G-quadruplex Resource Site
non-B Motif Search Tool at non-B DB
a web server to predict G-quadruplex forming motifs and other non-B DNA forming motifs from users' DNA sequences.
QGRS Mapper: a web-based application for predicting G-quadruplexes in nucleotide sequences and NCBI genes
from Bagga's group.
Quadfinder: Tool for Prediction and Analysis of G Quadruplex Motifs in DNA/RNA Sequences from Maiti's group, IGIB, Delhi, India
{dead link, date=October 2017 , bot=InternetArchiveBot , fix-attempted=yes
G4Hunter from Mergny's group but user need to run the code in R.
pqsfinder: an exhaustive and imperfection-tolerant search tool for potential quadruplex-forming sequences in R.
pqsfinder: online search tool using the latest R/Bioconductor package G-quadruplex, DNA
guanine
Guanine () ( symbol G or Gua) is one of the four main nucleobases found in the nucleic acids DNA and RNA, the others being adenine, cytosine, and thymine (uracil in RNA). In DNA, guanine is paired with cytosine. The guanine nucleoside is c ...
. They are helical in shape and contain guanine tetrads that can form from one, two or four strands. The unimolecular forms often occur naturally near the ends of the chromosomes, better known as the telomeric regions, and in transcriptional regulatory regions of multiple genes, both in microbes and across vertebrates including oncogenes in humans. Four guanine bases can associate through Hoogsteen hydrogen bonding to form a square planar structure called a guanine tetrad
In molecular biology, a guanine tetrad (also known as a G-tetrad or G-quartet) is a structure composed of four guanine bases in a square planar array. They most prominently contribute to the structure of G-quadruplexes, where their hydrogen bon ...
(G-tetrad or G-quartet), and two or more guanine tetrads (from G-tracts, continuous runs of guanine) can stack on top of each other to form a G-quadruplex.
The placement and bonding to form G-quadruplexes is not random and serve very unusual functional purposes. The quadruplex structure is further stabilized by the presence of a cation, especially potassium
Potassium is the chemical element with the symbol K (from Neo-Latin ''kalium'') and atomic number19. Potassium is a silvery-white metal that is soft enough to be cut with a knife with little force. Potassium metal reacts rapidly with atmosph ...
, which sits in a central channel between each pair of tetrads. They can be formed of DNA, RNA, LNA, and PNA, and may be intramolecular, bimolecular
In chemistry, molecularity is the number of molecules that come together to react in an elementary (single-step) reactionAtkins, P.; de Paula, J. Physical Chemistry. Oxford University Press, 2014 and is equal to the sum of stoichiometric coeffici ...
, or tetramolecular. Depending on the direction of the strands or parts of a strand that form the tetrads, structures may be described as parallel
Parallel is a geometric term of location which may refer to:
Computing
* Parallel algorithm
* Parallel computing
* Parallel metaheuristic
* Parallel (software), a UNIX utility for running programs in parallel
* Parallel Sysplex, a cluster of ...
or antiparallel. G-quadruplex structures can be computationally predicted from DNA or RNA sequence motifs, but their actual structures can be quite varied within and between the motifs, which can number over 100,000 per genome. Their activities in basic genetic processes are an active area of research in telomere, gene regulation, and functional genomics research.
History
The identification of structures with a highguanine
Guanine () ( symbol G or Gua) is one of the four main nucleobases found in the nucleic acids DNA and RNA, the others being adenine, cytosine, and thymine (uracil in RNA). In DNA, guanine is paired with cytosine. The guanine nucleoside is c ...
association became apparent in the early 1960s, through the identification of gel-like substances associated with guanines. More specifically, this research detailed the four-stranded DNA structures with a high association of guanines, which was later identified in eukaryotic telomeric
A telomere (; ) is a region of repetitive nucleotide sequences associated with specialized proteins at the ends of linear chromosomes. Although there are different architectures, telomeres, in a broad sense, are a widespread genetic feature mos ...
regions of DNA in the 1980s. The importance of discovering G-quadruplex structure was described through the statement, “If G-quadruplexes form so readily ''in vitro'', Nature will have found a way of using them ''in vivo''” - Aaron Klug, Nobel Prize Winner in Chemistry (1982). Interest in ''in vivo'' function of G-quadruplexes surged after large scale genome-wide analysis showed the prevalence of potential G-quadruplex (pG4)-forming sequences within gene promoters of human, chimpanzee, mouse, and rat - presented in the First International G-quadruplex Meeting held in April 2007 in Louisville, Kentucky. In 2006, the prevalence of G-quadruplexes within gene promoters of several bacterial genomes was reported predicting G-quadruplex-mediated gene regulation. With the abundance of G-quadruplexes ''in vivo'', these structures hold a biologically relevant role through interactions with the promoter regions of oncogenes and the telomeric regions of DNA strands. Current research consists of identifying the biological function of these G-Quadruplex structures for specific oncogenes and discovering effective therapeutic treatments for cancer based on interactions with G-quadruplexes.
Topology
The length of the nucleic acid sequences involved in tetrad formation determines how the quadruplex folds. Short sequences, consisting of only a single contiguous run of three or more guanine bases, require four individual strands to form a quadruplex. Such a quadruplex is described as tetramolecular, reflecting the requirement of four separate strands. The term G4 DNA was originally reserved for these tetramolecular structures that might play a role inmeiosis
Meiosis (; , since it is a reductional division) is a special type of cell division of germ cells in sexually-reproducing organisms that produces the gametes, such as sperm or egg cells. It involves two rounds of division that ultimately r ...
. However, as currently used in molecular biology, the term G4 can mean G-quadruplexes of any molecularity. Longer sequences, which contain two contiguous runs of three or more guanine bases, where the guanine regions are separated by one or more bases, only require two such sequences to provide enough guanine bases to form a quadruplex. These structures, formed from two separate G-rich strands, are termed bimolecular quadruplexes. Finally, sequences which contain four distinct runs of guanine bases can form stable quadruplex structures by themselves, and a quadruplex formed entirely from a single strand is called an intramolecular quadruplex.
Depending on how the individual runs of guanine bases are arranged in a bimolecular or intramolecular quadruplex, a quadruplex can adopt one of a number of topologies with varying loop configurations. If all strands of DNA proceed in the same direction, the quadruplex is termed parallel. For intramolecular quadruplexes, this means that any loop regions present must be of the propeller type, positioned to the sides of the quadruplex. If one or more of the runs of guanine bases has a 5’-3’ direction opposite to the other runs of guanine bases, the quadruplex is said to have adopted an antiparallel topology. The loops joining runs of guanine bases in intramolecular antiparallel quadruplexes are either diagonal, joining two diagonally opposite runs of guanine bases, or lateral (edgewise) type loops, joining two adjacent runs of guanine base pairs.
In quadruplexes formed from double-stranded DNA, possible interstrand topologies have also been discussed
.
Interstrand quadruplexes contain guanines that originate from both strands of dsDNA.
Structure and functional role in genome
Following sequencing of the humangenome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding g ...
, many guanine-rich sequences that had the potential to form quadruplexes were discovered. Depending on cell type and cell cycle, mediating factors such as DNA-binding proteins on chromatin
Chromatin is a complex of DNA and protein found in eukaryotic cells. The primary function is to package long DNA molecules into more compact, denser structures. This prevents the strands from becoming tangled and also plays important roles in r ...
, composed of DNA tightly wound around histone
In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei. They act as spools around which DNA winds to create structural units called nucleosomes. Nucleosomes in turn a ...
proteins, and other environmental conditions and stresses affect the dynamic formation of quadruplexes. For instance, quantitative assessments of the thermodynamics
Thermodynamics is a branch of physics that deals with heat, work, and temperature, and their relation to energy, entropy, and the physical properties of matter and radiation. The behavior of these quantities is governed by the four laws of th ...
of molecular crowding indicate that the antiparallel g-quadruplex is stabilized by molecular crowding. This effect seems to be mediated by alteration of the hydration of the DNA and its effect on Hoogsteen base pair
A Hoogsteen base pair is a variation of base-pairing in nucleic acids such as the A•T pair. In this manner, two nucleobases, one on each strand, can be held together by hydrogen bonds in the major groove. A Hoogsteen base pair applies the N7 pos ...
bonding. These quadruplexes seemed to readily occur at the ends of chromosome
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins are ...
. In addition, the propensity of g-quadruplex formation during transcription in RNA sequences with the potential to form mutually exclusive hairpin
A hairpin or hair pin is a long device used to hold a person's hair in place. It may be used simply to secure long hair out of the way for convenience or as part of an elaborate hairstyle or coiffure. The earliest evidence for dressing the hai ...
or G-quadruplex structures depends heavily on the position of the hairpin-forming sequence.
Because repair enzymes would naturally recognize ends of linear chromosomes as damaged DNA and would process them as such to harmful effect for the cell, clear signaling and tight regulation is needed at the ends of linear chromosomes. Telomeres
A telomere (; ) is a region of repetitive nucleotide sequences associated with specialized proteins at the ends of linear chromosomes. Although there are different architectures, telomeres, in a broad sense, are a widespread genetic feature mos ...
function to provide this signaling. Telomeres, rich in guanine and with a propensity to form g-quadruplexes, are located at the terminal ends of chromosomes and help maintain genome integrity by protecting these vulnerable terminal ends from instability.
These telomeric regions are characterized by long regions of double-stranded CCCTAA:TTAGGG repeats. The repeats end with a 3’ protrusion of between 10 and 50 single-stranded TTAGGG repeats. The heterodimeric complex ribonucleoprotein enzyme telomerase
Telomerase, also called terminal transferase, is a ribonucleoprotein that adds a species-dependent telomere repeat sequence to the 3' end of telomeres. A telomere is a region of repetitive sequences at each end of the chromosomes of most euka ...
adds TTAGGG repeats at the 3’ end of DNA strands. At these 3’ end protrusions, the G-rich overhang can form secondary structures such as G-quadruplexes if the overhang is longer than four TTAGGG repeats. The presence of these structures prevent telomere elongation by the telomerase complex.
Telomeric quadruplexes
Telomeric
A telomere (; ) is a region of repetitive nucleotide sequences associated with specialized proteins at the ends of linear chromosomes. Although there are different architectures, telomeres, in a broad sense, are a widespread genetic feature mos ...
repeats in a variety of organisms have been shown to form these quadruplex structures ''in vitro
''In vitro'' (meaning in glass, or ''in the glass'') studies are performed with microorganisms, cells, or biological molecules outside their normal biological context. Colloquially called " test-tube experiments", these studies in biology ...
'', and subsequently they have also been shown to form ''in vivo
Studies that are ''in vivo'' (Latin for "within the living"; often not italicized in English) are those in which the effects of various biological entities are tested on whole, living organisms or cells, usually animals, including humans, and ...
''. The human telomeric repeat (which is the same for all vertebrate
Vertebrates () comprise all animal taxa within the subphylum Vertebrata () (chordates with backbones), including all mammals, birds, reptiles, amphibians, and fish. Vertebrates represent the overwhelming majority of the phylum Chordata, with c ...
s) consists of many repeats of the sequenced (GGTTAG), and the quadruplexes formed by this structure can be in bead-like structures of 5 nm to 8 nm in size and have been well studied by NMR
Nuclear magnetic resonance (NMR) is a physical phenomenon in which nuclei in a strong constant magnetic field are perturbed by a weak oscillating magnetic field (in the near field) and respond by producing an electromagnetic signal with ...
, TEM and X-ray
An X-ray, or, much less commonly, X-radiation, is a penetrating form of high-energy electromagnetic radiation. Most X-rays have a wavelength ranging from 10 picometers to 10 nanometers, corresponding to frequencies in the range 30&nb ...
crystal structure
In crystallography, crystal structure is a description of the ordered arrangement of atoms, ions or molecules in a crystalline material. Ordered structures occur from the intrinsic nature of the constituent particles to form symmetric patterns ...
determination. The formation of these quadruplexes in telomeres has been shown to decrease the activity of the enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products ...
telomerase
Telomerase, also called terminal transferase, is a ribonucleoprotein that adds a species-dependent telomere repeat sequence to the 3' end of telomeres. A telomere is a region of repetitive sequences at each end of the chromosomes of most euka ...
, which is responsible for maintaining length of telomeres and is involved in around 85% of all cancer
Cancer is a group of diseases involving abnormal cell growth with the potential to invade or spread to other parts of the body. These contrast with benign tumors, which do not spread. Possible signs and symptoms include a lump, abnormal b ...
s. This is an active target of drug discovery, including telomestatin.
Non-telomeric quadruplexes
Quadruplexes are present in locations other than at thetelomere
A telomere (; ) is a region of repetitive nucleotide sequences associated with specialized proteins at the ends of linear chromosomes. Although there are different architectures, telomeres, in a broad sense, are a widespread genetic feature mos ...
. Analysis of human, chimpanzee, mouse and rat genomes showed enormous number of potential G-quadruplex (pG4)-forming sequences in non-telomeric regions. A large number of the non-telomeric G-quadruplexes were found within gene promoters, and were conserved across the species. Similarly, large number of G-quadruplexes were found in the E. coli and hundreds of other microbial genomes. Here also, like vertebrates G-quadruplexes were enriched within gene promoters. In addition, there was found more than one-billion-year conserved G-quadruplex locus in plants and algae, in gene encoding large subunit of RNA polymerase II. Although these studies predicted G-quadruplex-mediated gene regulation, it is unlikely that all pG4s would form in vivo. The proto-oncogene
An oncogene is a gene that has the potential to cause cancer. In tumor cells, these genes are often mutated, or expressed at high levels.
c-myc
''Myc'' is a family of regulator genes and proto-oncogenes that code for transcription factors. The ''Myc'' family consists of three related human genes: ''c-myc'' ( MYC), ''l-myc'' ( MYCL), and ''n-myc'' ( MYCN). ''c-myc'' (also sometimes re ...
forms a quadruplex in a nuclease hypersensitive region critical for gene activity. Other genes shown to form G-quadruplexes in their promoter regions include the chicken
The chicken (''Gallus gallus domesticus'') is a domesticated junglefowl species, with attributes of wild species such as the grey and the Ceylon junglefowl that are originally from Southeastern Asia. Rooster or cock is a term for an adu ...
β-globin gene
In biology, the word gene (from , ; "... Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a b ...
, human ubiquitin
Ubiquitin is a small (8.6 kDa) regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ''ubiquitously''. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 1980s. Fo ...
-ligase RFP2, and the proto-oncogenes
An oncogene is a gene that has the potential to cause cancer. In tumor cells, these genes are often mutated, or expressed at high levels.
c-kit
Proto-oncogene c-KIT is the gene encoding the receptor tyrosine kinase protein known as tyrosine-protein kinase KIT, CD117 (cluster of differentiation 117) or mast/stem cell growth factor receptor (SCFR). Multiple transcript variants encoding dif ...
, bcl-2
Bcl-2 (B-cell lymphoma 2), encoded in humans by the ''BCL2'' gene, is the founding member of the Bcl-2 family of regulator proteins that regulate cell death (apoptosis), by either inhibiting (anti-apoptotic) or inducing (pro-apoptotic) apoptosi ...
, VEGF
Vascular endothelial growth factor (VEGF, ), originally known as vascular permeability factor (VPF), is a signal protein produced by many cells that stimulates the formation of blood vessels. To be specific, VEGF is a sub-family of growth factors ...
, H-ras and N-ras.
Genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding g ...
-wide surveys based on a quadruplex folding rule have been performed, which have identified 376,000 Putative Quadruplex Sequences (PQS) in the human
Humans (''Homo sapiens'') are the most abundant and widespread species of primate, characterized by bipedalism and exceptional cognitive skills due to a large and complex brain. This has enabled the development of advanced tools, cultu ...
genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding g ...
, although not all of these probably form ''in vivo''. A similar studies have identified putative G-quadruplexes in prokaryotes, namely the bacterium ''E. coli''. There are several possible models for how quadruplexes could influence gene activity, either by upregulation
In the biological context of organisms' production of gene products, downregulation is the process by which a cell decreases the quantity of a cellular component, such as RNA or protein, in response to an external stimulus. The complementary proc ...
or downregulation
In the biological context of organisms' production of gene products, downregulation is the process by which a cell decreases the quantity of a cellular component, such as RNA or protein, in response to an external stimulus. The complementary pr ...
. One model is shown below, with G-quadruplex formation in or near a promoter blocking transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
of the gene, and hence de-activating it. In another model, quadruplex formed at the non-coding DNA strand helps to maintain an open conformation of the coding DNA strand and enhance an expression of the respective gene.
Function
It has been suggested that quadruplex formation plays a role in immunoglobulin heavy chain switching. As cells have evolved mechanisms for resolving (i.e., unwinding) quadruplexes that form. Quadruplex formation may be potentially damaging for a cell; the helicases WRN andBloom syndrome protein
Bloom syndrome protein is a protein that in humans is encoded by the ''BLM'' gene and is not expressed in Bloom syndrome.
The Bloom syndrome gene product is related to the RecQ subset of DExH box-containing DNA helicases and has both DNA-stimula ...
have a high affinity for resolving DNA G-quadruplexes. The DEAH/RHA helicase, DHX36, has also been identified as a key G-quadruplex resolvase. In 2009, a metastasis suppressor protein NM23H2 (also known as NME2) was found to directly interact with G-quadruplex in the promoter of the c-myc gene, and transcriptionally regulate c-myc. More recently, NM23H2 was reported to interact with G-quadruplex in the promoter of the human telomerase (hTERT) gene and regulate hTERT expression In 2019, the telomere-binding-factor-2 (TRF2 or TERF2) was shown to bind to thousands of non-telomeric G-quadruplexes in the human genome by TRF2 ChIP-seq. There are many studies that implicate quadruplexes in both positive and negative transcriptional regulation, including epigenetic regulation of genes like hTERT. Function of G-quadruplexes have also been reported in allowing programmed recombination of immunologlobin heavy genes and the pilin antigenic variation system of the pathogenic ''Neisseria
''Neisseria'' is a large genus of bacteria that colonize the mucosal surfaces of many animals. Of the 11 species that colonize humans, only two are pathogens, '' N. meningitidis'' and ''N. gonorrhoeae''.
''Neisseria'' species are Gram-negativ ...
''. The roles of quadruplex structure in translation control are not as well explored. The direct visualization of G-quadruplex structures in human cells as well as the co-crystal structure of an RNA helicase bound to a G-quadruplex have provided important confirmations of their relevance to cell biology. The potential positive and negative roles of quadruplexes in telomere replication and function remains controversial. T-loops and G-quadruplexes are described as the two tertiary DNA structures that protect telomere ends and regulate telomere length.
Genome Regulation through formation of G-quadruplex structures
Many of the genome regulatory processes have been linked to the formation of G-quadruplex structures, attributable to the huge role it plays in DNA repair of apurinic/apyrimidinic sites also known as AP sites. A new technique to map AP sites has been developed known as AP-seq which utilizes a biotin-labeled aldehyde-reactive probe (ARP) to tag certain regions of the genome where AP site damage occurrence has been significant. Another genome-wide mapping sequencing method known asChIP-sequencing
ChIP-sequencing, also known as ChIP-seq, is a method used to analyze protein interactions with DNA. ChIP-seq combines chromatin immunoprecipitation (ChIP) with massively parallel DNA sequencing to identify the binding sites of DNA-associated prot ...
, was utilized to map both; damage in AP sites, and the enzyme responsible for its repair, AP endonuclease 1 (APE1). Both of these genome-wide mapping sequencing methods, ChIP-sequencing
ChIP-sequencing, also known as ChIP-seq, is a method used to analyze protein interactions with DNA. ChIP-seq combines chromatin immunoprecipitation (ChIP) with massively parallel DNA sequencing to identify the binding sites of DNA-associated prot ...
and ARP, have indicated that AP site damage occurrence is nonrandom. AP site damage was also more prevalent in certain regions of the genome that contain specific active promoter and enhancer markers, some of which were linked to regions responsible for lung adenocarcinoma and colon cancer. AP site damage was found to be predominant in PQS regions of the genome, where formation of G-quadruplex structures is regulated and promoted by the DNA repair process, base excision repair
Base excision repair (BER) is a cellular mechanism, studied in the fields of biochemistry and genetics, that repairs damaged DNA throughout the cell cycle. It is responsible primarily for removing small, non-helix-distorting base lesions from t ...
(BER). Base excision repair processes in cells have been proved to be reduced with aging as its components in the mitochondria begin to decline, which can lead to the formation of many diseases such as Alzheimer's disease (AD). These G-quadruplex structures are said to be formed in the promoter regions of DNA through superhelicity, which favors the unwinding of the double helical structure of DNA and in turn loops the strands to form G-quadruplex structures in guanine rich regions. The BER pathway is signalled when it indicates an oxidative DNA base damage, where structures like, 8-Oxoguanine-DNA glycosylase 1 (OGG1), APE1 and G-quadruplex play a huge role in its repair. These enzymes participate in BER to repair certain DNA lesions such as 7,8-dihydro-8-oxoguanine (8-oxoG), which forms under oxidative stress to guanine bases.
Role of Endogenous Oxidized DNA Base Damage on G4 formation
Guanine (G) bases in G-quadruplex have the lowest redox potential causing it to be more susceptible to the formation of 8-oxoguanine (8-oxoG), an endogenous oxidized DNA base damage in the genome. Due toGuanine
Guanine () ( symbol G or Gua) is one of the four main nucleobases found in the nucleic acids DNA and RNA, the others being adenine, cytosine, and thymine (uracil in RNA). In DNA, guanine is paired with cytosine. The guanine nucleoside is c ...
having a lower electron reduction potential than the other nucleotides bases,8-oxo-2'-deoxyguanosine
8-Oxo-2'-deoxyguanosine (8-oxo-dG) is an oxidized derivative of deoxyguanosine. 8-Oxo-dG is one of the major products of DNA oxidation. Concentrations of 8-oxo-dG within a cell are a measurement of oxidative stress.
In DNA
Steady-state leve ...
(8-oxo-dG), is a known major product of DNA oxidation. Its concentration is used as a measurement of oxidative stress
Oxidative stress reflects an imbalance between the systemic manifestation of reactive oxygen species and a biological system's ability to readily detoxify the reactive intermediates or to repair the resulting damage. Disturbances in the normal ...
within a cell. When DNA undergoes oxidative damage, a possible structural change in guanine, after ionizing radiation, gives rise to an enol form, 8-OH-Gua. This oxidative product is formed through a tautomer
Tautomers () are structural isomers (constitutional isomers) of chemical compounds that readily interconvert.
The chemical reaction interconverting the two is called tautomerization. This conversion commonly results from the relocation of a hyd ...
ic shift from the original damage guanine, 8-oxo-Gua, and represents DNA damage that causes changes in the structure. This form allows for the base excision repair (BER) enzyme OGG1 to bind and remove the oxidative damage with the help of APE1, resulting in an AP site. Moreover, an AP site is a location in DNA that has neither a purine or a pyrimidine base due to DNA damage, they are the most prevalent type of endogenous DNA damage in cells. AP sites can be generated spontaneously or after the cleavage of modified bases, like 8-OH-Gua. The generation of an AP site enables the melting of the duplex DNA to unmask the PQS, adopting a G-quadruplex fold. With the use of genome-wide ChIP-sequencing
ChIP-sequencing, also known as ChIP-seq, is a method used to analyze protein interactions with DNA. ChIP-seq combines chromatin immunoprecipitation (ChIP) with massively parallel DNA sequencing to identify the binding sites of DNA-associated prot ...
analyses, cell-based assays, and in vitro biochemical analyses, a connection has been made between oxidized DNA base-derived AP sites, and the formation of the G-quadruplex.
DNA Oxidation Contribution to Diseases
Furthermore, the concentration of 8-oxo-dG is a known biomarker of oxidative stress within a cell, and excessive amount of oxidative stress has been linked to carcinogenesis and other diseases. When produced, 8-oxo-dG, has the ability to inactivate OGG1, thus preventing the repair of DNA damage caused by the oxidation of guanine. The possible inactivation allows for un-repaired DNA damages to gather in non-replicating cells, like muscle, and can cause aging as well. Moreover, oxidative DNA damage like 8-oxo-dG contributes to carcinogenesis
Carcinogenesis, also called oncogenesis or tumorigenesis, is the formation of a cancer, whereby normal cells are transformed into cancer cells. The process is characterized by changes at the cellular, genetic, and epigenetic levels and abno ...
through the modulation of gene expression, or the induction of mutations. On the condition that 8-oxo-dG is repaired by BER, parts of the repair protein is left behind which can lead to epigenetic alterations, or the modulation of gene expression. Upon insertion of 8-oxo-dG into thymidine kinase
Thymidine kinase is an enzyme, a phosphotransferase (a kinase): 2'-deoxythymidine kinase, ATP-thymidine 5'-phosphotransferase, EC 2.7.1.21. It can be found in most living cells. It is present in two forms in mammalian cells, TK1 and TK2. Cert ...
gene of humans, it was determine that if 8-oxo-dG was left unchecked and not repaired by BER, it can lead to frequent mutations and eventually carcinogenesis.
APE1 role in Gene Regulation
AP endonuclease 1 (APE1) is an enzyme responsible for the promotion and the formation of G-quadruplex structures. APE1 is mainly in charge of repairing damage caused to AP sites through the BER pathway. APE1 is considered to be very crucial as AP site damage is known to be the most recurring type of endogenous damage to DNA. The oxidation of certain purine bases, like guanine, forms oxidized nucleotides that impairs DNA function by mismatching nucleotides in the sequences. This is more common in PQS sequences which form oxidized structures, such as8-oxoguanine
8-Oxoguanine (8-hydroxyguanine, 8-oxo-Gua, or OH8Gua) is one of the most common DNA lesions resulting from reactive oxygen species modifying guanine, and can result in a mismatched pairing with adenine resulting in G to T and C to A substitutions ...
. Once the cell is aware of oxidative stress and damage, it recruits OGG1 to the site, whose main function is to initiate the BER pathway. OGG1 does so by cleaving the oxidized base and thus creating an AP site, primarily through the process of negative superhelicity. This AP site then signals cells to engage APE1 binding, which binds to the open duplex region. The binding of APE1 then plays an important role by stabilizing the formation of G-quadruplex structures in that region. This promotes formation of G-quadruplex structures by the folding of the stand. This looping process brings four bases in close proximity that will be held together by Hoogsteen base pairing. After this stage the APE1 gets acetylated by multiple lysine residues on the chromatin, forming acetylated APE1 (AcAPE1). AcAPE1 is very crucial to the BER pathway as it acts as a transcriptional coactivator or corepressor, functioning to load transcription factors (TF) into the site of damage allowing it to regulate the gene expression. AcAPE1 is also very important since it allows APE1 to bind for longer periods of time by delay of its dissociation from the sequence, allowing the repair process to be more efficient. Deacetylation of AcAPE1 is the driving force behind the loading of these TFs, where APE1 dissociates from the G-quadruplex structures. When a study downregulated the presence of APE1 and AcAPE1 in the cell, the formation of G-quadruplex structures was inhibited, which proves the importance of APE1 for the formation of these structures. However, not all G-quadruplex structures require APE1 for formation, in fact some of them formed greater G-quadruplex structures in its absence. Therefore, we can conclude that APE1 has two important roles in genome regulation- Stabilizing the formation of g-quadruplex structures and loading the transcriptional factors onto the AP site
Cancer
Telomeres
G-quadruplex forming sequences are prevalent ineukaryotic
Eukaryotes () are organisms whose Cell (biology), cells have a cell nucleus, nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the ...
cells, especially in telomeres, 5` untranslated strands, and translocation hot spots. G-quadruplexes can inhibit normal cell function, and in healthy cells, are easily and readily unwound by helicase
Helicases are a class of enzymes thought to be vital to all organisms. Their main function is to unpack an organism's genetic material. Helicases are motor proteins that move directionally along a nucleic acid phosphodiester backbone, separatin ...
. However, in cancer cells that have mutated helicase these complexes cannot be unwound and leads to potential damage of the cell. This causes replication of damaged and cancerous cells. For therapeutic advances, stabilizing the G-quadruplexes of cancerous cells can inhibit cell growth and replication leading to the cell's death.
Promoter Regions
Along with the association of G-quadruplexes intelomeric
A telomere (; ) is a region of repetitive nucleotide sequences associated with specialized proteins at the ends of linear chromosomes. Although there are different architectures, telomeres, in a broad sense, are a widespread genetic feature mos ...
regions of DNA, G-quadruplex structures have been identified in various human proto-oncogene
An oncogene is a gene that has the potential to cause cancer. In tumor cells, these genes are often mutated, or expressed at high levels.
promoter regions. The structures most present in the promoter regions of these oncogenes tend to be parallel-stranded G-quadruplex DNA structures. Some of these oncogenes include c-KIT, PDGF-A, ''c-Myc'' and VEGF, showing the importance of this secondary structure in cancer growth and development. While the formation of G-quadruplex structure vary to some extent for the different promoter regions of oncogenes, the consistent stabilization of these structures have been found in cancer development. Current therapeutic research actively focuses on targeting this stabilization of G-quadruplex structures to arrest unregulated cell growth and division.
One particular gene region, the c-myc pathway, plays an integral role in the regulation of a protein product, c-Myc. With this product, the c-Myc protein functions in the processes of apoptosis and cell growth or development and as a transcriptional control on human telomerase reverse transcriptase
Telomerase reverse transcriptase (abbreviated to TERT, or hTERT in humans) is a catalytic subunit of the enzyme telomerase, which, together with the telomerase RNA component (TERC), comprises the most important unit of the telomerase complex.
T ...
. Interaction of c-Myc promoter G-quadruplex with NM23H2 was shown to regulate c-Myc in cancer cells in 2009
Regulation of c-myc through Human telomerase reverse transcriptase (hTERT) is also directly regulated through promoter G-quadruplex by interaction with the transcription factor NM23H2 where epigenetic modifications were dependent on NM23H2-G-quadruplex association. Recently, hTERT epigenetic regulation reported to be mediated through interaction of hTERT promoter G-quadruplex with the telomeric factor TRF2.
Another gene pathway deals with the VEGF gene, Vascular Endothelial Growth Factor, which remains involved in the process of angiogenesis or the formation of new blood vessels. The formation of an intramolecular G-quadruplex structure has been shown through studies on the polypurine tract of the promoter region
In genetics, a promoter is a sequence of DNA to which proteins bind to initiate transcription of a single RNA transcript from the DNA downstream of the promoter. The RNA transcript may encode a protein (mRNA), or can have a function in and of ...
of the VEGF gene. Through recent research on the role of G-quadruplex function in vivo, the stabilization of G-quadruplex structures was shown to regulate VEGF gene transcription, with inhibition of transcription factors in this pathway. The intramolecular G-quadruplex structures are formed mostly through the abundant guanine sequence in the promoter region of this specific pathway. The cyclin-dependent cell cycle checkpoint kinase inhibitor-1 CDKN1A (also known as p21) gene harbours promoter G-quadruplex. Interaction of this G-quadruplex with TRF2 (also known as TERF2) resulted in epigenetic regulation of p21, which was tested using the G-quadruplex-binding ligand 360A.
Hypoxia inducible factor 1ɑ, HIF-1ɑ, remains involved in cancer signaling through its binding to Hypoxia Response Element, HRE, in the presence of hypoxia to begin the process of angiogenesis. Through recent research into this specific gene pathway, the polypurine and polypyrimidine region allows for the transcription of this specific gene and the formation of an intramolecular G-quadruplex structure. However, more research is necessary to determine whether the formation of G-quadruplex regulates the expression of this gene in a positive or negative manner.
The c-kit oncogene deals with a pathway that encodes an RTK, which was shown to have elevated expression levels in certain types of cancer. The rich guanine sequence of this promoter region has shown the ability to form a variety of quadruplexes. Current research on this pathway is focusing on discovering the biological function of this specific quadruplex formation on the c-kit pathway, while this quadruplex sequence has been noticed in various species.
The RET oncogene functions in the transcription of kinase which has been abundant in certain types of cancer. The guanine rich sequence in the promoter region for this pathway exudes a necessity for baseline transcription of this receptor tyrosine kinase. In certain types of cancers, the RET protein has shown increased expression levels. The research on this pathway suggested the formation of a G-quadruplex in the promoter region and an applicable target for therapeutic treatments.
Another oncogene pathway involving PDGF-A, platelet-derived growth factor, involves the process of wound healing and function as mitogenic growth factors for cells. High levels of expression of PDGF have been associated with increased cell growth and cancer. The presence of a guanine-rich sequence in the promoter region of PDGF-A has exhibited the ability to form intramolecular parallel G-quadruplex structures and remains suggested to play a role in transcriptional regulation of PDGF-A. However, research has also identified the presence of G-quadruplex structures within this region due to the interaction of TMPyP4 with this promoter sequence.
Therapeutics
Telomeres are generally made up of G-quadruplexes and remain important targets for therapeutic research and discoveries. These complexes have a high affinity for porphyrin rings which makes them effective anticancer agents. However, TMPyP4 has been limited for used due to its non-selectivity toward cancer cell telomeres and normal double stranded DNA (dsDNA). To address this issue analog of TMPyP4, it was synthesized known as 5Me which targets only G quadruplex DNA which inhibits cancer growth more effectively than TMPyP4. Ligand design and development remains an important field of research into therapeutic reagents due to the abundance of G-quadruplexes and their multiple conformational differences. One type of ligand involving a Quindoline derivative, SYUIQ-05, utilizes the stabilization of G-quadruplexes in promoter regions to inhibit the production of both the c-Myc protein product and the human telomerase reverse transcriptase (hTERT). This main pathway of targeting this region results in the lack of telomerase elongation, leading to arrested cell development. Further research remains necessary for the discovery of a single gene target to minimize unwanted reactivity with more efficient antitumor activity.Ligands which bind quadruplexes
One way of inducing or stabilizing G-quadruplex formation is to introduce a molecule which can bind to the G-quadruplex structure. A number ofligand
In coordination chemistry, a ligand is an ion or molecule ( functional group) that binds to a central metal atom to form a coordination complex. The bonding with the metal generally involves formal donation of one or more of the ligand's elec ...
s, which can be both small molecules and protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, res ...
s, can bind to the G-quadruplex. These ligands can be naturally occurring or synthetic. This has become an increasingly large field of research in genetics, biochemistry, and pharmacology.
Cationic porphyrin
Porphyrins ( ) are a group of heterocyclic macrocycle organic compounds, composed of four modified pyrrole subunits interconnected at their α carbon atoms via methine bridges (=CH−). The parent of porphyrin is porphine, a rare chemical com ...
s have been shown to bind intercalatively with G-quadruplexes, as well as the molecule telomestatin.
The binding of ligands to G-quadruplexes is vital for anti-cancer pursuits because G-quadruplexes are found typically at translocation hot spots. MM41, a ligand that binds selectively for a quadruplex on the BCL-2
Bcl-2 (B-cell lymphoma 2), encoded in humans by the ''BCL2'' gene, is the founding member of the Bcl-2 family of regulator proteins that regulate cell death (apoptosis), by either inhibiting (anti-apoptotic) or inducing (pro-apoptotic) apoptosi ...
promoter, is shaped with a central core and 4 side chains branching sterically out. The shape of the ligand is vital because it closely matches the quadruplex which has stacked quartets and the loops of nucleic acids holding it together. When bound, MM41's central chromophore is situated on top of the 3’ terminal G-quartet and the side chains of the ligand associate to the loops of the quadruplex. The quartet and the chromophore are bound with a π-π bond while the side chains and loops are not bound but are in close proximity. What makes this binding strong is the fluidity in the position of the loops to better associate with the ligand side chains.
TMPyP4, a cationic porphyrin, is a more well known G4 binding ligand that helps to repress ''c-Myc.'' The way in which TMPyP4 binds to G4's is similar to MM41, with the ring stacking onto the external G-quartet and side chains associating to the loops of G4's.
When designing ligands to be bound to G-quadruplexes, the ligands have a higher affinity for parallel folded G-quadruplexes. It has been found that ligands with smaller side chains bind better to the quadruplex because smaller ligands have more concentrated electron density
In quantum chemistry, electron density or electronic density is the measure of the probability of an electron being present at an infinitesimal element of space surrounding any given point. It is a scalar quantity depending upon three spatial va ...
. Furthermore, the hydrogen bonds of ligands with smaller side chains are shorter and therefore stronger. Ligands with mobile side chains, ones that are able to rotate around its center chromophore, associate more strongly to G-quadruplexes because conformation of the G4 loops and the ligand side chains can align.
Quadruplex prediction techniques
Identifying and predicting sequences which have the capacity to form quadruplexes is an important tool in further understanding their role. Generally, a simple pattern match is used for searching for possible intrastrand quadruplex forming sequences: d(G3+N1-7G3+N1-7G3+N1-7G3+), where N is anynucleotide base
Nucleobases, also known as ''nitrogenous bases'' or often simply ''bases'', are nitrogen-containing biological compounds that form nucleosides, which, in turn, are components of nucleotides, with all of these monomers constituting the basic b ...
(including guanine
Guanine () ( symbol G or Gua) is one of the four main nucleobases found in the nucleic acids DNA and RNA, the others being adenine, cytosine, and thymine (uracil in RNA). In DNA, guanine is paired with cytosine. The guanine nucleoside is c ...
). This rule has been widely used in on-line algorithm
In mathematics and computer science, an algorithm () is a finite sequence of rigorous instructions, typically used to solve a class of specific problems or to perform a computation. Algorithms are used as specifications for performing ...
s. Although the rule effectively identifies sites of G-quadruplex formation, it also identifies a subset of the imperfect homopurine mirror repeats capable of triplex formation and C-strand i-motif formation. Moreover, these sequences also have the capacity to form slipped and foldback structures that are implicit intermediates in the formation of both quadruplex and triplex DNA structures. In one study, it was found that the observed number per base pair (i.e. the frequency) of these motifs has increased rapidly in the eumetazoa for which complete genomic sequences are available. This suggests that the sequences may be under positive selection enabled by the evolution of systems capable of suppressing non-B structure formation.
Methods for studying G-quadruplexes
A number of experimental methods have been developed to identify G-quadruplexes. These methods can be broadly defined into two classes: biophysical and biochemical methods.Biochemical methods
Biochemical techniques were employed to interrogate G-quadruplex formation in a longer sequence context. In the DNA polymerase stop assay, the formation of a G-quadruplex in a DNA template can act as a roadblock and cause polymerase stalling, which halts the primer extension. The dimethyl sulfate (DMS) followed by the piperidine cleavage assay is based on the fact that the formation of a G-quadruplex will prohibit the N7 guanine methylation caused by DMS, leading to a protection pattern observed at the DNA G-quadruplex region after piperidine cleavage.Biophysical methods
The topology of the G-quadruplex structure can be determined by monitoring the positive or negative circular dichroism (CD) signals at specific wavelengths. Parallel G-quadruplexes have negative and positive CD signals at 240 and 262 nm, respectively, whereas antiparallel G-quadruplexes place these signals at 262 and 295 nm, respectively. To verify G-quadruplex formation, one should also perform the CD experiments under non-G-quadruplex stabilizing (Li+) and G-quadruplex stabilizing conditions (such as K+ or with G-quadruplex ligands), and scan toward the far-UV region (180–230 nm). Likewise, the thermostability of the G-quadruplex structure can be identified by observing the UV signal at 295 nm. Upon G-quadruplex melting, the UV absorbance at 295 nm decreases, leading to a hypochromic shift that is a distinctive feature of G-quadruplex structure. Another approach for detection of G-quadruplexes includesnanopore
A nanopore is a pore of nanometer size. It may, for example, be created by a pore-forming protein or as a hole in synthetic materials such as silicon or graphene.
When a nanopore is present in an electrically insulating membrane, it can be used a ...
-based methods. Firstly, it was shown that biological nanopores can detect G-quadruplexes based on size exclusion and specific interaction of G-quadruplex and protein nanocavity. The novel approach combines solid-state nanopores and DNA nanotechnology
DNA nanotechnology is the design and manufacture of artificial nucleic acid structures for technological uses. In this field, nucleic acids are used as non-biological engineering materials for nanotechnology rather than as the carriers of geneti ...
for label-free detection of G-quadruplexes, for their mapping on dsDNA, and for monitoring G-quadruplex formation.
Role in neurological disorders
G-quadruplexes have been implicated in neurological disorders through two main mechanisms. The first is through expansions of G-repeats within genes that lead to the formation of G-quadruplex structures that directly cause disease, as is the case with the C9orf72 gene and amyotrophic lateral sclerosis (ALS) or frontotemporal dementia (FTD). The second mechanism is through mutations that affect the expression of G-quadruplex binding proteins, as seen in the fragile X mental retardation gene 1 (FMR1) gene and Fragile X Syndrome. The C9orf72 gene codes for the proteinC9orf72
C9orf72 (chromosome 9 open reading frame 72) is a protein which in humans is encoded by the gene ''C9orf72''.
The human ''C9orf72'' gene is located on the short (p) arm of chromosome 9 open reading frame 72, from base pair 27,546,546 to base pa ...
which is found throughout the brain in neuron
A neuron, neurone, or nerve cell is an electrically excitable cell that communicates with other cells via specialized connections called synapses. The neuron is the main component of nervous tissue in all animals except sponges and placozoa. ...
al cytoplasm
In cell biology, the cytoplasm is all of the material within a eukaryotic cell, enclosed by the cell membrane, except for the cell nucleus. The material inside the nucleus and contained within the nuclear membrane is termed the nucleoplasm. ...
and at presynaptic
In the nervous system, a synapse is a structure that permits a neuron (or nerve cell) to pass an electrical or chemical signal to another neuron or to the target effector cell.
Synapses are essential to the transmission of nervous impulses from ...
terminals. Mutations of the C9orf72 gene have been linked to the development of FTD and ALS. These two diseases have a causal relationship to GGGGCC (G4C2) repeats within the 1st intron of C9orf72 gene. Normal individuals typically have around 2 to 8 G4C2 repeats, but individuals with FTD or ALS have from 500 to several thousand G4C2 repeats. The transcribed RNA of these repeats have been shown to form stable G-quadruplexes, with evidence showing that the G4C2 repeats in DNA have the ability to form mixed parallel-antiparallel G-quadruplex structures as well. These RNA transcripts containing G4C2 repeats were shown to bind and separate a wide variety of proteins, including nucleolin
Nucleolin is a protein that in humans is encoded by the ''NCL'' gene.
Gene
The human NCL gene is located on chromosome 2 and consists of 14 exons with 13 introns and spans approximately 11kb. The intron 11 of the NCL gene encodes a small nucl ...
. Nucleolin is involved in the synthesis and maturation of ribosomes within the nucleus, and separation of nucleolin by the mutated RNA transcripts impairs nucleolar function and ribosomal RNA synthesis.
Fragile X mental retardation protein (FMRP) is a widely expressed protein coded by the FMR1 gene that binds to G-quadruplex secondary structures in neurons and is involved in synaptic plasticity
In neuroscience, synaptic plasticity is the ability of synapses to strengthen or weaken over time, in response to increases or decreases in their activity. Since memories are postulated to be represented by vastly interconnected neural circuits ...
. FMRP acts as a negative regulator of translation, and its binding stabilizes G-quadruplex structures in mRNA transcripts, inhibiting ribosome elongation of mRNA in the neuron's dendrite
Dendrites (from Greek δένδρον ''déndron'', "tree"), also dendrons, are branched protoplasmic extensions of a nerve cell that propagate the electrochemical stimulation received from other neural cells to the cell body, or soma, of the ...
and controlling the timing of the transcript's expression. Mutations of this gene can cause the development of Fragile X Syndrome, autism, and other neurological disorders. Specifically, Fragile X Syndrome is caused by an increase from 50 to over 200 CGG repeats within exon 13 of the FMR1 gene. This repeat expansion promotes DNA methylation and other epigenetic heterochromatin modifications of FMR1 that prevent the transcription of the gene, leading to pathological low levels of FMRP.
Therapeutic approaches
Antisense-mediated interventions and small-molecule ligands are common strategies used to target neurological diseases linked to G-quadruplex expansion repeats. Therefore, these techniques are especially advantageous for targeting neurological diseases that have a gain-of-function mechanism, which is when the altered gene product has a new function or new expression of a gene; this has been detected in theC9orf72
C9orf72 (chromosome 9 open reading frame 72) is a protein which in humans is encoded by the gene ''C9orf72''.
The human ''C9orf72'' gene is located on the short (p) arm of chromosome 9 open reading frame 72, from base pair 27,546,546 to base pa ...
(chromosome 9 open reading frame 72).
Antisense therapy
Antisense therapy is a form of treatment that uses antisense oligonucleotides (ASOs) to target messenger RNA (mRNA). ASOs are capable of altering mRNA expression through a variety of mechanisms, including ribonuclease H mediated decay of the pre- ...
is the process by which synthesized strands of nucleic acids are used to bind directly and specifically to the mRNA produced by a certain gene, which will inactivate it. Antisense oligonucleotides (ASOs) are commonly used to target C9orf72 RNA of the G-quadruplex GGGGCC expansion repeat region, which has lowered the toxicity in cellular models of C9orf72. ASOs have previously been used to restore normal phenotypes in other neurological diseases that have gain-of-function mechanisms, the only difference is that it was used in the absence of G-quadruplex expansion repeat regions.
Another commonly used technique is the utilization of small-molecule ligands. These can be used to target G-quadruplex regions that cause neurological disorders. Approximately 1,000 various G-quadruplex ligands exist in which they are able to interact via their aromatic rings
In chemistry, aromaticity is a chemical property of cyclic ( ring-shaped), ''typically'' planar (flat) molecular structures with pi bonds in resonance (those containing delocalized electrons) that gives increased stability compared to saturat ...
; this allows the small-molecule ligands to stack on the planar terminal tetrads within the G-quadruplex regions. A disadvantage of using small-molecule ligands as a therapeutic technique is that specificity is difficult to manage due to the variability of G-quadruplexes in their primary sequences, orientation, thermodynamic stability, and nucleic acid strand stoichiometry. As of now, no single small-molecule ligand has been able to be perfectly specific for a single G-quadruplex sequence. However, a cationic porphyrin known as TMPyP4 is able to bind to the C9orf72 GGGGCC repeat region, which causes the G-quadruplex repeat region to unfold and lose its interactions with proteins causing it to lose its functionality. Small-molecule ligands, composed primarily of lead, can target GGGGCC repeat regions as well and ultimately decreased both repeat-associated non-ATG translation and RNA foci in neuron cells derived from patients with Amyotrophic lateral sclerosis (ALS). This provides evidence that small-molecule ligands are an effective and efficient process to target GGGGCC regions, and that specificity for small-molecule ligand binding is a feasible goal for the scientific community.
Metal complexes have a number of features that make them particularly suitable as G4 DNA binders and therefore as potential drugs. While the metal plays largely a structural role in most G4 binders, there are also examples where it interacts directly with G4s by electrostatic interactions or direct coordination with nucleobases.
References
Further reading
* * * * * * * * * *External links
Nanopore and Aptamer Biosensor group
Quadruplex websites
G-Quadruplex World
– a website to discuss publications and other information of interest to those working in the field of G-quadruplexes
Greglist
– a database listing potential G-quadruplex regulated genes
Database on Quadruplex information: QuadBase from IGIB
GRSDB
a database of G-quadruplexes near RNA processing sites.
GRS_UTRdb
- a database of G-quadruplexes in the UTRs.
G-quadruplex Resource Site
non-B Motif Search Tool at non-B DB
a web server to predict G-quadruplex forming motifs and other non-B DNA forming motifs from users' DNA sequences.
Tools to predict G-quadruplex motifs
QGRS Mapper: a web-based application for predicting G-quadruplexes in nucleotide sequences and NCBI genes
from Bagga's group.
Quadfinder: Tool for Prediction and Analysis of G Quadruplex Motifs in DNA/RNA Sequences from Maiti's group, IGIB, Delhi, India
{dead link, date=October 2017 , bot=InternetArchiveBot , fix-attempted=yes
G4Hunter from Mergny's group but user need to run the code in R.
pqsfinder: an exhaustive and imperfection-tolerant search tool for potential quadruplex-forming sequences in R.
pqsfinder: online search tool using the latest R/Bioconductor package G-quadruplex, DNA