DNA sequencing is the process of determining the
nucleic acid sequence – the order of
nucleotides in
DNA. It includes any method or technology that is used to determine the order of the four bases:
adenine
Adenine () ( symbol A or Ade) is a nucleobase (a purine derivative). It is one of the four nucleobases in the nucleic acid of DNA that are represented by the letters G–C–A–T. The three others are guanine, cytosine and thymine. Its deriv ...
,
guanine
Guanine () ( symbol G or Gua) is one of the four main nucleobases found in the nucleic acids DNA and RNA, the others being adenine, cytosine, and thymine ( uracil in RNA). In DNA, guanine is paired with cytosine. The guanine nucleoside is ...
,
cytosine
Cytosine () ( symbol C or Cyt) is one of the four nucleobases found in DNA and RNA, along with adenine, guanine, and thymine ( uracil in RNA). It is a pyrimidine derivative, with a heterocyclic aromatic ring and two substituents attached ( ...
, and
thymine
Thymine () ( symbol T or Thy) is one of the four nucleobases in the nucleic acid of DNA that are represented by the letters G–C–A–T. The others are adenine, guanine, and cytosine. Thymine is also known as 5-methyluracil, a pyrimidin ...
. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery.
Knowledge of DNA sequences has become indispensable for basic biological research,
DNA Genographic Projects and in numerous applied fields such as
medical diagnosis
Medical diagnosis (abbreviated Dx, Dx, or Ds) is the process of determining which disease or condition explains a person's symptoms and signs. It is most often referred to as diagnosis with the medical context being implicit. The information r ...
,
biotechnology
Biotechnology is the integration of natural sciences and engineering sciences in order to achieve the application of organisms, cells, parts thereof and molecular analogues for products and services. The term ''biotechnology'' was first used ...
,
forensic biology
Forensic biology is the application of biology to associate a person(s), whether suspect or victim, to a location, an item (or collection of items), another person (victim or suspect, respectively). It can be utilized to further investigations for ...
,
virology
Virology is the scientific study of biological viruses. It is a subfield of microbiology that focuses on their detection, structure, classification and evolution, their methods of infection and exploitation of host cells for reproduction, the ...
and biological
systematics
Biological systematics is the study of the diversification of living forms, both past and present, and the relationships among living things through time. Relationships are visualized as evolutionary trees (synonyms: cladograms, phylogenetic t ...
. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers, characterize antibody repertoire,
and can be used to guide patient treatment. Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged.
The rapid speed of sequencing attained with modern DNA sequencing technology has been instrumental in the sequencing of complete DNA sequences, or
genomes, of numerous types and species of life, including the
human genome and other complete DNA sequences of many animal, plant, and microbial species.
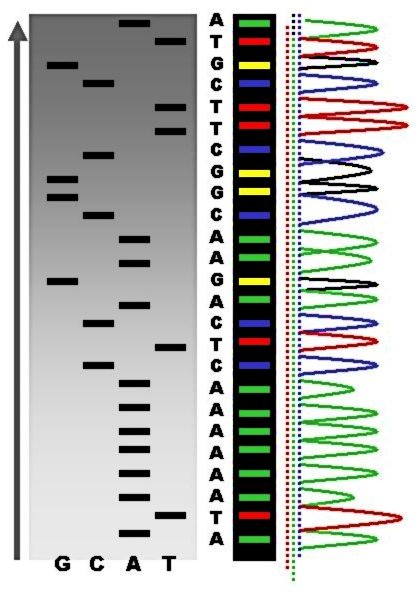
The first DNA sequences were obtained in the early 1970s by academic researchers using laborious methods based on
two-dimensional chromatography
Two-dimensional chromatography is a type of chromatographic technique in which the injected sample is separated by passing through two different separation stages. Two different chromatographic columns are connected in sequence, and the effluent fr ...
. Following the development of
fluorescence
Fluorescence is the emission of light by a substance that has absorbed light or other electromagnetic radiation. It is a form of luminescence. In most cases, the emitted light has a longer wavelength, and therefore a lower photon energy, tha ...
-based sequencing methods with a
DNA sequencer,
DNA sequencing has become easier and orders of magnitude faster.
Applications
DNA sequencing may be used to determine the sequence of individual
gene
In biology, the word gene (from , ; "...Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a b ...
s, larger genetic regions (i.e. clusters of genes or
operons), full chromosomes, or
entire genomes of any organism. DNA sequencing is also the most efficient way to indirectly sequence
RNA or
protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, res ...
s (via their
open reading frame
In molecular biology, open reading frames (ORFs) are defined as spans of DNA sequence between the start and stop codons. Usually, this is considered within a studied region of a prokaryotic DNA sequence, where only one of the six possible readi ...
s). In fact, DNA sequencing has become a key technology in many areas of biology and other sciences such as medicine,
forensics, and
anthropology
Anthropology is the scientific study of humanity, concerned with human behavior, human biology, cultures, societies, and linguistics, in both the present and past, including past human species. Social anthropology studies patterns of be ...
.
Molecular biology
Sequencing is used in
molecular biology
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions. The study of chemical and phys ...
to study genomes and the proteins they encode. Information obtained using sequencing allows researchers to identify changes in genes and noncoding DNA (including regulatory sequences), associations with diseases and phenotypes, and identify potential drug targets.
Evolutionary biology
Since DNA is an informative macromolecule in terms of transmission from one generation to another, DNA sequencing is used in
evolutionary biology
Evolutionary biology is the subfield of biology that studies the evolutionary processes (natural selection, common descent, speciation) that produced the diversity of life on Earth. It is also defined as the study of the history of life ...
to study how different organisms are related and how they evolved. In February 2021, scientists reported, for the first time, the sequencing of
DNA from
animal remains, a
mammoth
A mammoth is any species of the extinct elephantid genus ''Mammuthus'', one of the many genera that make up the order of trunked mammals called proboscideans. The various species of mammoth were commonly equipped with long, curved tusks an ...
in this instance, over a million years old, the oldest DNA sequenced to date.
Metagenomics
The field of
metagenomics involves identification of organisms present in a body of water,
sewage, dirt, debris filtered from the air, or swab samples from organisms. Knowing which organisms are present in a particular environment is critical to research in
ecology
Ecology () is the study of the relationships between living organisms, including humans, and their physical environment. Ecology considers organisms at the individual, population, community, ecosystem, and biosphere level. Ecology overl ...
,
epidemiology
Epidemiology is the study and analysis of the distribution (who, when, and where), patterns and determinants of health and disease conditions in a defined population.
It is a cornerstone of public health, and shapes policy decisions and evi ...
,
microbiology
Microbiology () is the scientific study of microorganisms, those being unicellular (single cell), multicellular (cell colony), or acellular (lacking cells). Microbiology encompasses numerous sub-disciplines including virology, bacteriology, ...
, and other fields. Sequencing enables researchers to determine which types of microbes may be present in a
microbiome
A microbiome () is the community of microorganisms that can usually be found living together in any given habitat. It was defined more precisely in 1988 by Whipps ''et al.'' as "a characteristic microbial community occupying a reasonably wel ...
, for example.
Virology
As most viruses are too small to be seen by a light microscope, sequencing is one of the main tools in virology to identify and study the virus.
Viral genomes can be based in DNA or RNA. RNA viruses are more time-sensitive for genome sequencing, as they degrade faster in clinical samples.
Traditional
Sanger sequencing
Sanger sequencing is a method of DNA sequencing that involves electrophoresis and is based on the random incorporation of chain-terminating dideoxynucleotides by DNA polymerase during in vitro DNA replication. After first being developed by Fred ...
and next-generation sequencing are used to sequence viruses in basic and clinical research, as well as for the diagnosis of emerging viral infections,
molecular epidemiology of viral pathogens, and drug-resistance testing. There are more than 2.3 million unique viral sequences in
GenBank.
Recently, NGS has surpassed traditional Sanger as the most popular approach for generating viral genomes.
During the
1990 avian influenza outbreak, viral sequencing determined that the influenza sub-type originated through
reassortment between
quail and poultry. This led to legislation in
Hong Kong
Hong Kong ( (US) or (UK); , ), officially the Hong Kong Special Administrative Region of the People's Republic of China (abbr. Hong Kong SAR or HKSAR), is a List of cities in China, city and Special administrative regions of China, special ...
that prohibited selling live quail and poultry together at market. Viral sequencing can also be used to estimate when a viral outbreak began by using a
molecular clock
The molecular clock is a figurative term for a technique that uses the mutation rate of biomolecules to deduce the time in prehistory when two or more life forms diverged. The biomolecular data used for such calculations are usually nucleo ...
technique.
Medicine
Medical technicians may sequence genes (or, theoretically, full genomes) from patients to determine if there is risk of genetic diseases. This is a form of
genetic testing, though some genetic tests may not involve DNA sequencing.
DNA sequencing is also being increasingly used to diagnose and treat rare diseases. As more and more genes are identified that cause rare genetic diseases, molecular diagnoses for patients becomes more mainstream. DNA sequencing allows clinicians to identify genetic diseases, improve disease management, provide reproductive counseling, and more effective therapies.
Also, DNA sequencing may be useful for determining a specific bacteria, to allow for more
precise antibiotics treatments, hereby reducing the risk of creating
antimicrobial resistance in bacteria populations.
Forensic investigation
DNA sequencing may be used along with
DNA profiling methods for
forensic identification and
paternity testing. DNA testing has evolved tremendously in the last few decades to ultimately link a DNA print to what is under investigation. The DNA patterns in fingerprint, saliva, hair follicles, etc. uniquely separate each living organism from another. Testing DNA is a technique which can detect specific genomes in a DNA strand to produce a unique and individualized pattern.
The four canonical bases
The canonical structure of DNA has four bases:
thymine
Thymine () ( symbol T or Thy) is one of the four nucleobases in the nucleic acid of DNA that are represented by the letters G–C–A–T. The others are adenine, guanine, and cytosine. Thymine is also known as 5-methyluracil, a pyrimidin ...
(T),
adenine
Adenine () ( symbol A or Ade) is a nucleobase (a purine derivative). It is one of the four nucleobases in the nucleic acid of DNA that are represented by the letters G–C–A–T. The three others are guanine, cytosine and thymine. Its deriv ...
(A),
cytosine
Cytosine () ( symbol C or Cyt) is one of the four nucleobases found in DNA and RNA, along with adenine, guanine, and thymine ( uracil in RNA). It is a pyrimidine derivative, with a heterocyclic aromatic ring and two substituents attached ( ...
(C), and
guanine
Guanine () ( symbol G or Gua) is one of the four main nucleobases found in the nucleic acids DNA and RNA, the others being adenine, cytosine, and thymine ( uracil in RNA). In DNA, guanine is paired with cytosine. The guanine nucleoside is ...
(G). DNA sequencing is the determination of the physical order of these bases in a molecule of DNA. However, there are many other bases that may be present in a molecule. In some viruses (specifically,
bacteriophage
A bacteriophage (), also known informally as a ''phage'' (), is a duplodnaviria virus that infects and replicates within bacteria and archaea. The term was derived from "bacteria" and the Greek φαγεῖν ('), meaning "to devour". Bac ...
), cytosine may be replaced by hydroxy methyl or hydroxy methyl glucose cytosine. In mammalian DNA, variant bases with
methyl
In organic chemistry, a methyl group is an alkyl derived from methane, containing one carbon atom bonded to three hydrogen atoms, having chemical formula . In formulas, the group is often abbreviated as Me. This hydrocarbon group occurs in ...
groups or phosphosulfate may be found. Depending on the sequencing technique, a particular modification, e.g., the 5mC (
5 methyl cytosine) common in humans, may or may not be detected.
History
Discovery of DNA structure and function
Deoxyribonucleic acid (
DNA) was first discovered and isolated by
Friedrich Miescher in 1869, but it remained under-studied for many decades because proteins, rather than DNA, were thought to hold the genetic blueprint to life. This situation changed after 1944 as a result of some experiments by
Oswald Avery,
Colin MacLeod, and
Maclyn McCarty
Maclyn McCarty (June 9, 1911 – January 2, 2005) was an American geneticist, a research scientist described in 2005 as "the last surviving member of a Manhattan scientific team that overturned medical dogma in the 1940's and became the first to ...
demonstrating that purified DNA could change one strain of bacteria into another. This was the first time that DNA was shown capable of transforming the properties of cells.
In 1953,
James Watson and
Francis Crick
Francis Harry Compton Crick (8 June 1916 – 28 July 2004) was an English molecular biologist, biophysicist, and neuroscientist. He, James Watson, Rosalind Franklin, and Maurice Wilkins played crucial roles in deciphering the helical stru ...
put forward their
double-helix model of DNA, based on
crystallized X-ray structures being studied by
Rosalind Franklin. According to the model, DNA is composed of two strands of nucleotides coiled around each other, linked together by hydrogen bonds and running in opposite directions. Each strand is composed of four complementary nucleotides – adenine (A), cytosine (C), guanine (G) and thymine (T) – with an A on one strand always paired with T on the other, and C always paired with G. They proposed that such a structure allowed each strand to be used to reconstruct the other, an idea central to the passing on of hereditary information between generations.

The foundation for sequencing proteins was first laid by the work of
Frederick Sanger who by 1955 had completed the sequence of all the amino acids in
insulin
Insulin (, from Latin ''insula'', 'island') is a peptide hormone produced by beta cells of the pancreatic islets encoded in humans by the ''INS'' gene. It is considered to be the main anabolic hormone of the body. It regulates the metabolism ...
, a small protein secreted by the pancreas. This provided the first conclusive evidence that proteins were chemical entities with a specific molecular pattern rather than a random mixture of material suspended in fluid. Sanger's success in sequencing insulin spurred on x-ray crystallographers, including Watson and Crick, who by now were trying to understand how DNA directed the formation of proteins within a cell. Soon after attending a series of lectures given by Frederick Sanger in October 1954, Crick began developing a theory which argued that the arrangement of nucleotides in DNA determined the sequence of amino acids in proteins, which in turn helped determine the function of a protein. He published this theory in 1958.
RNA sequencing
RNA sequencing
RNA-Seq (named as an abbreviation of RNA sequencing) is a sequencing technique which uses next-generation sequencing (NGS) to reveal the presence and quantity of RNA in a biological sample at a given moment, analyzing the continuously changing c ...
was one of the earliest forms of nucleotide sequencing. The major landmark of RNA sequencing is the sequence of the first complete gene and the complete genome of
Bacteriophage MS2
Bacteriophage MS2 (''Emesvirus zinderi''), commonly called MS2, is an icosahedral, positive-sense single-stranded RNA virus that infects the bacterium ''Escherichia coli'' and other members of the Enterobacteriaceae. MS2 is a member of a family ...
, identified and published by
Walter Fiers and his coworkers at the
University of Ghent (
Ghent
Ghent ( nl, Gent ; french: Gand ; traditional English: Gaunt) is a city and a municipality in the Flemish Region of Belgium. It is the capital and largest city of the East Flanders province, and the third largest in the country, exceeded i ...
,
Belgium
Belgium, ; french: Belgique ; german: Belgien officially the Kingdom of Belgium, is a country in Northwestern Europe. The country is bordered by the Netherlands to the north, Germany to the east, Luxembourg to the southeast, France to ...
), in 1972 and 1976. Traditional RNA sequencing methods require the creation of a
cDNA molecule which must be sequenced.
Early DNA sequencing methods
The first method for determining DNA sequences involved a location-specific primer extension strategy established by
Ray Wu
Ray Jui Wu (, 14 August 1928 – 10 February 2008) was a Chinese-born American geneticist. A pioneer of plant genetic engineering, Wu was Liberty Hyde Bailey Professor of Molecular Genetics and Biology at Cornell University.
Biography
Wu was th ...
at
Cornell University
Cornell University is a private statutory land-grant research university based in Ithaca, New York. It is a member of the Ivy League. Founded in 1865 by Ezra Cornell and Andrew Dickson White, Cornell was founded with the intention to tea ...
in 1970. DNA polymerase catalysis and specific nucleotide labeling, both of which figure prominently in current sequencing schemes, were used to sequence the cohesive ends of lambda phage DNA.
Between 1970 and 1973, Wu, R Padmanabhan and colleagues demonstrated that this method can be employed to determine any DNA sequence using synthetic location-specific primers.
then adopted this primer-extension strategy to develop more rapid DNA sequencing methods at the
MRC Centre,
Cambridge
Cambridge ( ) is a university city and the county town in Cambridgeshire, England. It is located on the River Cam approximately north of London. As of the 2021 United Kingdom census, the population of Cambridge was 145,700. Cambridge bec ...
, UK and published a method for "DNA sequencing with chain-terminating inhibitors" in 1977.
and
Allan Maxam
Allan Maxam (born October 28, 1942) is one of the pioneers of molecular genetics. He was one of the contributors to develop a DNA sequencing method at Harvard University, while working as a student in the laboratory of Walter Gilbert.
Walter Gi ...
at
Harvard
Harvard University is a private Ivy League research university in Cambridge, Massachusetts. Founded in 1636 as Harvard College and named for its first benefactor, the Puritan clergyman John Harvard, it is the oldest institution of higher le ...
also developed sequencing methods, including one for "DNA sequencing by chemical degradation".
[ In 1973, Gilbert and Maxam reported the sequence of 24 basepairs using a method known as wandering-spot analysis. Advancements in sequencing were aided by the concurrent development of ]recombinant DNA
Recombinant DNA (rDNA) molecules are DNA molecules formed by laboratory methods of genetic recombination (such as molecular cloning) that bring together genetic material from multiple sources, creating sequences that would not otherwise be f ...
technology, allowing DNA samples to be isolated from sources other than viruses.
Sequencing of full genomes
The first full DNA genome to be sequenced was that of bacteriophage φX174 in 1977. Medical Research Council scientists deciphered the complete DNA sequence of the Epstein-Barr virus in 1984, finding it contained 172,282 nucleotides. Completion of the sequence marked a significant turning point in DNA sequencing because it was achieved with no prior genetic profile knowledge of the virus.
A non-radioactive method for transferring the DNA molecules of sequencing reaction mixtures onto an immobilizing matrix during electrophoresis was developed by Herbert Pohl and co-workers in the early 1980s. Followed by the commercialization of the DNA sequencer "Direct-Blotting-Electrophoresis-System GATC 1500" by GATC Biotech, which was intensively used in the framework of the EU genome-sequencing programme, the complete DNA sequence of the yeast ''Saccharomyces cerevisiae
''Saccharomyces cerevisiae'' () (brewer's yeast or baker's yeast) is a species of yeast (single-celled fungus microorganisms). The species has been instrumental in winemaking, baking, and brewing since ancient times. It is believed to have b ...
'' chromosome II.Leroy E. Hood
Leroy "Lee" Edward Hood (born October 10, 1938) is an American biologist who has served on the faculties at the California Institute of Technology (Caltech) and the University of Washington.
Hood has developed ground-breaking scientific instrum ...
's laboratory at the California Institute of Technology announced the first semi-automated DNA sequencing machine in 1986. This was followed by Applied Biosystems
Applied Biosystems is one of various brands under the Life Technologies brand of Thermo Fisher Scientific corporation. The brand is focused on integrated systems for genetic analysis, which include computerized machines and the consumables used w ...
' marketing of the first fully automated sequencing machine, the ABI 370, in 1987 and by Dupont's Genesis 2000 which used a novel fluorescent labeling technique enabling all four dideoxynucleotides to be identified in a single lane. By 1990, the U.S. National Institutes of Health
The National Institutes of Health, commonly referred to as NIH (with each letter pronounced individually), is the primary agency of the United States government responsible for biomedical and public health research. It was founded in the lat ...
(NIH) had begun large-scale sequencing trials on '' Mycoplasma capricolum'', ''Escherichia coli
''Escherichia coli'' (),Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. also known as ''E. coli'' (), is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus '' Esc ...
'', ''Caenorhabditis elegans
''Caenorhabditis elegans'' () is a free-living transparent nematode about 1 mm in length that lives in temperate soil environments. It is the type species of its genus. The name is a blend of the Greek ''caeno-'' (recent), ''rhabditis'' (r ...
'', and ''Saccharomyces cerevisiae
''Saccharomyces cerevisiae'' () (brewer's yeast or baker's yeast) is a species of yeast (single-celled fungus microorganisms). The species has been instrumental in winemaking, baking, and brewing since ancient times. It is believed to have b ...
'' at a cost of US$0.75 per base. Meanwhile, sequencing of human cDNA sequences called expressed sequence tags began in Craig Venter
John Craig Venter (born October 14, 1946) is an American biotechnologist and businessman. He is known for leading one of the first draft sequences of the human genome and assembled the first team to transfect a cell with a synthetic chromosome. ...
's lab, an attempt to capture the coding fraction of the human genome.Haemophilus influenzae
''Haemophilus influenzae'' (formerly called Pfeiffer's bacillus or ''Bacillus influenzae'') is a Gram-negative, non-motile, coccobacillary, facultatively anaerobic, capnophilic pathogenic bacterium of the family Pasteurellaceae. The bact ...
''. The circular chromosome contains 1,830,137 bases and its publication in the journal Science marked the first published use of whole-genome shotgun sequencing, eliminating the need for initial mapping efforts.
By 2001, shotgun sequencing methods had been used to produce a draft sequence of the human genome.
High-throughput sequencing (HTS) methods
 Several new methods for DNA sequencing were developed in the mid to late 1990s and were implemented in commercial DNA sequencers by 2000. Together these were called the "next-generation" or "second-generation" sequencing (NGS) methods, in order to distinguish them from the earlier methods, including
Several new methods for DNA sequencing were developed in the mid to late 1990s and were implemented in commercial DNA sequencers by 2000. Together these were called the "next-generation" or "second-generation" sequencing (NGS) methods, in order to distinguish them from the earlier methods, including Sanger sequencing
Sanger sequencing is a method of DNA sequencing that involves electrophoresis and is based on the random incorporation of chain-terminating dideoxynucleotides by DNA polymerase during in vitro DNA replication. After first being developed by Fred ...
. In contrast to the first generation of sequencing, NGS technology is typically characterized by being highly scalable, allowing the entire genome to be sequenced at once. Usually, this is accomplished by fragmenting the genome into small pieces, randomly sampling for a fragment, and sequencing it using one of a variety of technologies, such as those described below. An entire genome is possible because multiple fragments are sequenced at once (giving it the name "massively parallel" sequencing) in an automated process.
NGS technology has tremendously empowered researchers to look for insights into health, anthropologists to investigate human origins, and is catalyzing the " Personalized Medicine" movement. However, it has also opened the door to more room for error. There are many software tools to carry out the computational analysis of NGS data, often compiled at online platforms such as CSI NGS Portal, each with its own algorithm. Even the parameters within one software package can change the outcome of the analysis. In addition, the large quantities of data produced by DNA sequencing have also required development of new methods and programs for sequence analysis. Several efforts to develop standards in the NGS field have been attempted to address these challenges, most of which have been small-scale efforts arising from individual labs. Most recently, a large, organized, FDA-funded effort has culminated in the BioCompute standard.
On 26 October 1990, Roger Tsien, Pepi Ross, Margaret Fahnestock and Allan J Johnston filed a patent describing stepwise ("base-by-base") sequencing with removable 3' blockers on DNA arrays (blots and single DNA molecules).Stockholm
Stockholm () is the capital and largest city of Sweden as well as the largest urban area in Scandinavia. Approximately 980,000 people live in the municipality, with 1.6 million in the urban area, and 2.4 million in the metropo ...
published their method of pyrosequencing
Pyrosequencing is a method of DNA sequencing (determining the order of nucleotides in DNA) based on the "sequencing by synthesis" principle, in which the sequencing is performed by detecting the nucleotide incorporated by a DNA polymerase. Pyrosequ ...
.polymerase chain reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) ...
(PCR) arraying methods described in this patent, coupled to Roger Tsien et al.'s "base-by-base" sequencing method, is now implemented in Illumina's Hi-Seq genome sequencers.
In 1998, Phil Green and Brent Ewing of the University of Washington described their phred quality score for sequencer data analysis, a landmark analysis technique that gained widespread adoption, and which is still the most common metric for assessing the accuracy of a sequencing platform.
Lynx Therapeutics published and marketed massively parallel signature sequencing
Massive parallel signature sequencing (MPSS) is a procedure that is used to identify and quantify mRNA transcripts, resulting in data similar to serial analysis of gene expression (SAGE), although it employs a series of biochemical and sequencing ...
(MPSS), in 2000. This method incorporated a parallelized, adapter/ligation-mediated, bead-based sequencing technology and served as the first commercially available "next-generation" sequencing method, though no DNA sequencers were sold to independent laboratories.
Basic methods
Maxam-Gilbert sequencing
Allan Maxam
Allan Maxam (born October 28, 1942) is one of the pioneers of molecular genetics. He was one of the contributors to develop a DNA sequencing method at Harvard University, while working as a student in the laboratory of Walter Gilbert.
Walter Gi ...
and Walter Gilbert published a DNA sequencing method in 1977 based on chemical modification of DNA and subsequent cleavage at specific bases.
Chain-termination methods
The chain-termination method developed by Frederick Sanger and coworkers in 1977 soon became the method of choice, owing to its relative ease and reliability.genomics
Genomics is an interdisciplinary field of biology focusing on the structure, function, evolution, mapping, and editing of genomes. A genome is an organism's complete set of DNA, including all of its genes as well as its hierarchical, three-dim ...
. However, later in the decade, radically different approaches reached the market, bringing the cost per genome down from $100 million in 2001 to $10,000 in 2011.
Large-scale sequencing and ''de novo'' sequencing
 Large-scale sequencing often aims at sequencing very long DNA pieces, such as whole
Large-scale sequencing often aims at sequencing very long DNA pieces, such as whole chromosome
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins ar ...
s, although large-scale sequencing can also be used to generate very large numbers of short sequences, such as found in phage display. For longer targets such as chromosomes, common approaches consist of cutting (with restriction enzyme
A restriction enzyme, restriction endonuclease, REase, ENase or'' restrictase '' is an enzyme that cleaves DNA into fragments at or near specific recognition sites within molecules known as restriction sites. Restriction enzymes are one class ...
s) or shearing (with mechanical forces) large DNA fragments into shorter DNA fragments. The fragmented DNA may then be cloned
Cloning is the process of producing individual organisms with identical or virtually identical DNA, either by natural or artificial means. In nature, some organisms produce clones through asexual reproduction. In the field of biotechnology, ...
into a DNA vector and amplified in a bacterial host such as ''Escherichia coli
''Escherichia coli'' (),Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. also known as ''E. coli'' (), is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus '' Esc ...
''. Short DNA fragments purified from individual bacterial colonies are individually sequenced and assembled electronically into one long, contiguous sequence. Studies have shown that adding a size selection step to collect DNA fragments of uniform size can improve sequencing efficiency and accuracy of the genome assembly. In these studies, automated sizing has proven to be more reproducible and precise than manual gel sizing.primer walking
Primer walking is a technique used to clone a gene (e.g., disease gene) from its known closest markers (e.g., known gene). As a result, it is employed in cloning and sequencing efforts in plants, fungi, and mammals with minor alterations. This te ...
. The different strategies have different tradeoffs in speed and accuracy; shotgun methods are often used for sequencing large genomes, but its assembly is complex and difficult, particularly with sequence repeats often causing gaps in genome assembly.
Most sequencing approaches use an ''in vitro'' cloning step to amplify individual DNA molecules, because their molecular detection methods are not sensitive enough for single molecule sequencing. Emulsion PCRpolymerase chain reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) ...
(PCR) then coats each bead with clonal copies of the DNA molecule followed by immobilization for later sequencing. Emulsion PCR is used in the methods developed by Marguilis et al. (commercialized by 454 Life Sciences), Shendure and Porreca et al. (also known as "polony sequencing
Polony sequencing is an inexpensive but highly accurate multiplex sequencing technique that can be used to “read” millions of immobilized DNA sequences in parallel. This technique was first developed by Dr. George Church's group at Harvard ...
") and SOLiD sequencing, (developed by Agencourt, later Applied Biosystems
Applied Biosystems is one of various brands under the Life Technologies brand of Thermo Fisher Scientific corporation. The brand is focused on integrated systems for genetic analysis, which include computerized machines and the consumables used w ...
, now Life Technologies).
Shotgun sequencing
Shotgun sequencing is a sequencing method designed for analysis of DNA sequences longer than 1000 base pairs, up to and including entire chromosomes. This method requires the target DNA to be broken into random fragments. After sequencing individual fragments using the chain termination method, the sequences can be reassembled on the basis of their overlapping regions.
High-throughput methods
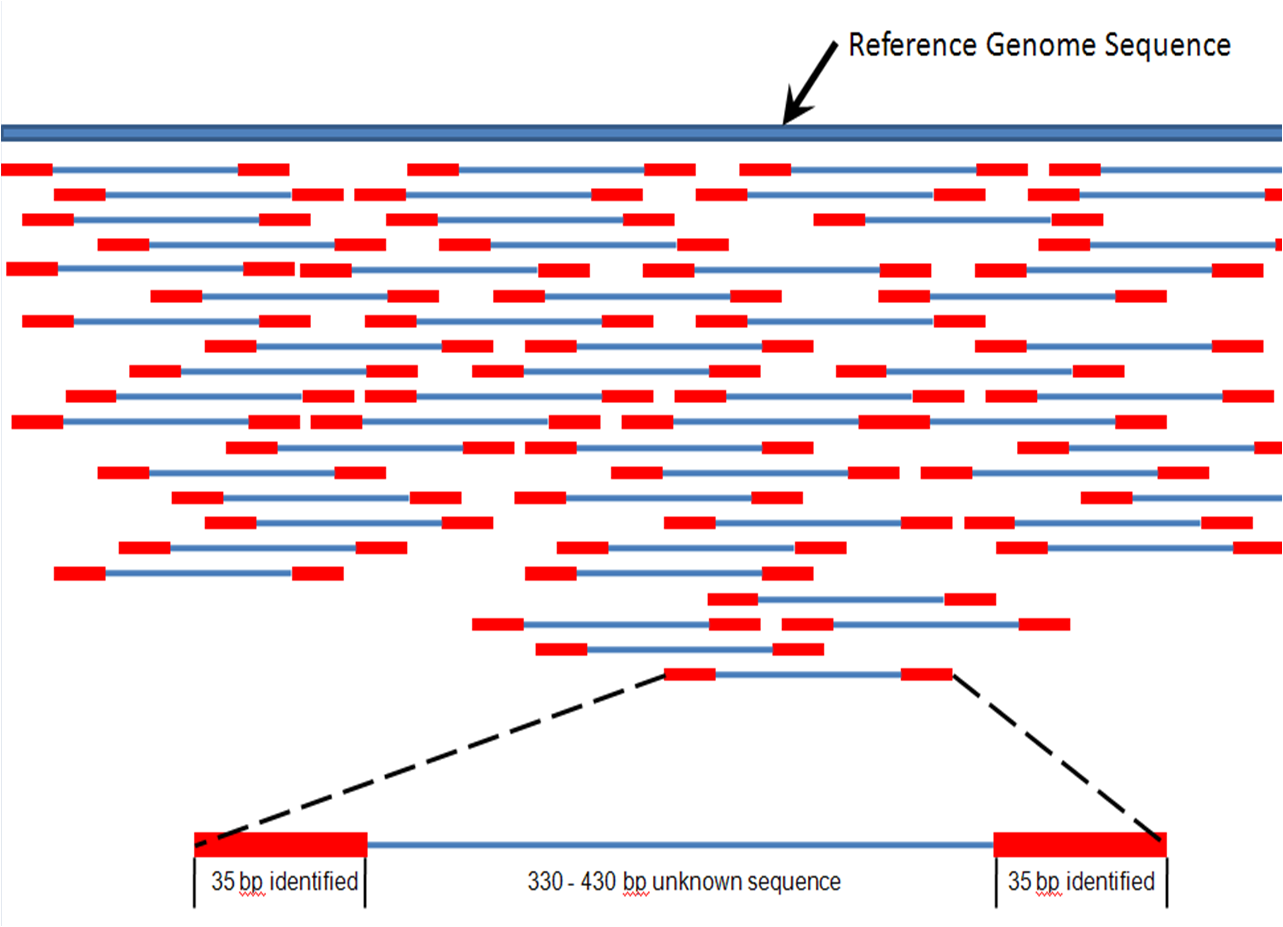 High-throughput sequencing, which includes next-generation "short-read" and third-generation "long-read" sequencing methods,
High-throughput sequencing, which includes next-generation "short-read" and third-generation "long-read" sequencing methods,["Next-generation" remains in broad use as of 2019. For instance, ] applies to exome sequencing
Exome sequencing, also known as whole exome sequencing (WES), is a genomic technique for sequencing all of the protein-coding regions of genes in a genome (known as the exome). It consists of two steps: the first step is to select only the sub ...
, genome sequencing, genome resequencing, transcriptome profiling ( RNA-Seq), DNA-protein interactions ( ChIP-sequencing), and epigenome characterization.
Long-read sequencing methods
Single molecule real time (SMRT) sequencing
SMRT sequencing is based on the sequencing by synthesis approach. The DNA is synthesized in zero-mode wave-guides (ZMWs) – small well-like containers with the capturing tools located at the bottom of the well. The sequencing is performed with use of unmodified polymerase (attached to the ZMW bottom) and fluorescently labelled nucleotides flowing freely in the solution. The wells are constructed in a way that only the fluorescence occurring by the bottom of the well is detected. The fluorescent label is detached from the nucleotide upon its incorporation into the DNA strand, leaving an unmodified DNA strand. According to Pacific Biosciences
Pacific Biosciences of California, Inc. (aka PacBio) is an American biotechnology company founded in 2004 that develops and manufactures systems for gene sequencing and some novel real time biological observation. PacBio describes its platform ...
(PacBio), the SMRT technology developer, this methodology allows detection of nucleotide modifications (such as cytosine methylation). This happens through the observation of polymerase kinetics. This approach allows reads of 20,000 nucleotides or more, with average read lengths of 5 kilobases.
Nanopore DNA sequencing
The DNA passing through the nanopore changes its ion current. This change is dependent on the shape, size and length of the DNA sequence. Each type of the nucleotide blocks the ion flow through the pore for a different period of time. The method does not require modified nucleotides and is performed in real time. Nanopore sequencing is referred to as " third-generation" or "long-read" sequencing, along with SMRT sequencing.
Early industrial research into this method was based on a technique called 'exonuclease sequencing', where the readout of electrical signals occurred as nucleotides passed by alpha(α)-hemolysin pores covalently bound with cyclodextrin. However the subsequent commercial method, 'strand sequencing', sequenced DNA bases in an intact strand.
Two main areas of nanopore sequencing in development are solid state nanopore sequencing, and protein based nanopore sequencing. Protein nanopore sequencing utilizes membrane protein complexes such as α-hemolysin, MspA ('' Mycobacterium smegmatis'' Porin A) or CssG, which show great promise given their ability to distinguish between individual and groups of nucleotides.
Short-read sequencing methods
Massively parallel signature sequencing (MPSS)
The first of the high-throughput sequencing technologies, massively parallel signature sequencing
Massive parallel signature sequencing (MPSS) is a procedure that is used to identify and quantify mRNA transcripts, resulting in data similar to serial analysis of gene expression (SAGE), although it employs a series of biochemical and sequencing ...
(or MPSS), was developed in the 1990s at Lynx Therapeutics, a company founded in 1992 by Sydney Brenner and Sam Eletr
Sam, SAM or variants may refer to:
Places
* Sam, Benin
* Sam, Boulkiemdé, Burkina Faso
* Sam, Bourzanga, Burkina Faso
* Sam, Kongoussi, Burkina Faso
* Sam, Iran
* Sam, Teton County, Idaho, United States, a populated place
People and fictional ...
. MPSS was a bead-based method that used a complex approach of adapter ligation followed by adapter decoding, reading the sequence in increments of four nucleotides. This method made it susceptible to sequence-specific bias or loss of specific sequences. Because the technology was so complex, MPSS was only performed 'in-house' by Lynx Therapeutics and no DNA sequencing machines were sold to independent laboratories. Lynx Therapeutics merged with Solexa (later acquired by Illumina) in 2004, leading to the development of sequencing-by-synthesis, a simpler approach acquired from Manteia Predictive Medicine, which rendered MPSS obsolete. However, the essential properties of the MPSS output were typical of later high-throughput data types, including hundreds of thousands of short DNA sequences. In the case of MPSS, these were typically used for sequencing cDNA for measurements of gene expression levels.
Polony sequencing
The polony sequencing
Polony sequencing is an inexpensive but highly accurate multiplex sequencing technique that can be used to “read” millions of immobilized DNA sequences in parallel. This technique was first developed by Dr. George Church's group at Harvard ...
method, developed in the laboratory of George M. Church
George McDonald Church (born August 28, 1954) is an American geneticist, molecular engineer, chemist, and a serial entrepreneur who is widely regarded as the "Founding Father of Genomics", and a pioneer in personal genomics and synthetic b ...
at Harvard, was among the first high-throughput sequencing systems and was used to sequence a full '' E. coli'' genome in 2005.Applied Biosystems
Applied Biosystems is one of various brands under the Life Technologies brand of Thermo Fisher Scientific corporation. The brand is focused on integrated systems for genetic analysis, which include computerized machines and the consumables used w ...
SOLiD platform. Applied Biosystems was later acquired by Life Technologies, now part of Thermo Fisher Scientific.
454 pyrosequencing
A parallelized version of pyrosequencing
Pyrosequencing is a method of DNA sequencing (determining the order of nucleotides in DNA) based on the "sequencing by synthesis" principle, in which the sequencing is performed by detecting the nucleotide incorporated by a DNA polymerase. Pyrosequ ...
was developed by 454 Life Sciences, which has since been acquired by Roche Diagnostics. The method amplifies DNA inside water droplets in an oil solution (emulsion PCR), with each droplet containing a single DNA template attached to a single primer-coated bead that then forms a clonal colony. The sequencing machine contains many picoliter-volume wells each containing a single bead and sequencing enzymes. Pyrosequencing uses luciferase to generate light for detection of the individual nucleotides added to the nascent DNA, and the combined data are used to generate sequence reads.
Illumina (Solexa) sequencing
Solexa, now part of Illumina, was founded by Shankar Balasubramanian and David Klenerman in 1998, and developed a sequencing method based on reversible dye-terminators technology, and engineered polymerases.
 In this method, DNA molecules and primers are first attached on a slide or flow cell and amplified with polymerase so that local clonal DNA colonies, later coined "DNA clusters", are formed. To determine the sequence, four types of reversible terminator bases (RT-bases) are added and non-incorporated nucleotides are washed away. A camera takes images of the fluorescently labeled nucleotides. Then the dye, along with the terminal 3' blocker, is chemically removed from the DNA, allowing for the next cycle to begin. Unlike pyrosequencing, the DNA chains are extended one nucleotide at a time and image acquisition can be performed at a delayed moment, allowing for very large arrays of DNA colonies to be captured by sequential images taken from a single camera.
In this method, DNA molecules and primers are first attached on a slide or flow cell and amplified with polymerase so that local clonal DNA colonies, later coined "DNA clusters", are formed. To determine the sequence, four types of reversible terminator bases (RT-bases) are added and non-incorporated nucleotides are washed away. A camera takes images of the fluorescently labeled nucleotides. Then the dye, along with the terminal 3' blocker, is chemically removed from the DNA, allowing for the next cycle to begin. Unlike pyrosequencing, the DNA chains are extended one nucleotide at a time and image acquisition can be performed at a delayed moment, allowing for very large arrays of DNA colonies to be captured by sequential images taken from a single camera.
 Decoupling the enzymatic reaction and the image capture allows for optimal throughput and theoretically unlimited sequencing capacity. With an optimal configuration, the ultimately reachable instrument throughput is thus dictated solely by the analog-to-digital conversion rate of the camera, multiplied by the number of cameras and divided by the number of pixels per DNA colony required for visualizing them optimally (approximately 10 pixels/colony). In 2012, with cameras operating at more than 10 MHz A/D conversion rates and available optics, fluidics and enzymatics, throughput can be multiples of 1 million nucleotides/second, corresponding roughly to 1 human genome equivalent at 1x
Decoupling the enzymatic reaction and the image capture allows for optimal throughput and theoretically unlimited sequencing capacity. With an optimal configuration, the ultimately reachable instrument throughput is thus dictated solely by the analog-to-digital conversion rate of the camera, multiplied by the number of cameras and divided by the number of pixels per DNA colony required for visualizing them optimally (approximately 10 pixels/colony). In 2012, with cameras operating at more than 10 MHz A/D conversion rates and available optics, fluidics and enzymatics, throughput can be multiples of 1 million nucleotides/second, corresponding roughly to 1 human genome equivalent at 1x coverage
Coverage may refer to:
Filmmaking
* Coverage (lens), the size of the image a lens can produce
* Camera coverage, the amount of footage shot and different camera setups used in filming a scene
* Script coverage, a short summary of a script, writ ...
per hour per instrument, and 1 human genome re-sequenced (at approx. 30x) per day per instrument (equipped with a single camera).
Combinatorial probe anchor synthesis (cPAS)
This method is an upgraded modification to combinatorial probe anchor ligation technology (cPAL) described by Complete Genomics
Complete Genomics is a life sciences company that has developed and commercialized a DNA sequencing platform for human genome sequencing and analysis. This solution combines the company's proprietary human genome sequencing technology with its in ...
 The patterned array of positively charged spots is fabricated through photolithography and etching techniques followed by chemical modification to generate a sequencing flow cell. Each spot on the flow cell is approximately 250 nm in diameter, are separated by 700 nm (centre to centre) and allows easy attachment of a single negatively charged DNB to the flow cell and thus reducing under or over-clustering on the flow cell.
The patterned array of positively charged spots is fabricated through photolithography and etching techniques followed by chemical modification to generate a sequencing flow cell. Each spot on the flow cell is approximately 250 nm in diameter, are separated by 700 nm (centre to centre) and allows easy attachment of a single negatively charged DNB to the flow cell and thus reducing under or over-clustering on the flow cell.
SOLiD sequencing

Applied Biosystems
Applied Biosystems is one of various brands under the Life Technologies brand of Thermo Fisher Scientific corporation. The brand is focused on integrated systems for genetic analysis, which include computerized machines and the consumables used w ...
' (now a Life Technologies brand) SOLiD technology employs sequencing by ligation Sequencing by ligation is a DNA sequencing method that uses the enzyme DNA ligase to identify the nucleotide present at a given position in a DNA sequence. Unlike most currently popular DNA sequencing methods, this method does not use a DNA polymera ...
. Here, a pool of all possible oligonucleotides of a fixed length are labeled according to the sequenced position. Oligonucleotides are annealed and ligated; the preferential ligation by DNA ligase
DNA ligase is a specific type of enzyme, a ligase, () that facilitates the joining of DNA strands together by catalyzing the formation of a phosphodiester bond. It plays a role in repairing single-strand breaks in duplex DNA in living orga ...
for matching sequences results in a signal informative of the nucleotide at that position. Each base in the template is sequenced twice, and the resulting data are decoded according to the 2 base encoding
2 Base Encoding, also called SOLiD (sequencing by oligonucleotide ligation and detection), is a next-generation sequencing technology developed by Applied Biosystems and has been commercially available since 2008. These technologies generate hundr ...
scheme used in this method. Before sequencing, the DNA is amplified by emulsion PCR. The resulting beads, each containing single copies of the same DNA molecule, are deposited on a glass slide.sequencing by ligation Sequencing by ligation is a DNA sequencing method that uses the enzyme DNA ligase to identify the nucleotide present at a given position in a DNA sequence. Unlike most currently popular DNA sequencing methods, this method does not use a DNA polymera ...
method has been reported to have some issue sequencing palindromic sequences.
Ion Torrent semiconductor sequencing
Ion Torrent Systems Inc. (now owned by Life Technologies) developed a system based on using standard sequencing chemistry, but with a novel, semiconductor-based detection system. This method of sequencing is based on the detection of hydrogen ion
A hydrogen ion is created when a hydrogen atom loses or gains an electron. A positively charged hydrogen ion (or proton) can readily combine with other particles and therefore is only seen isolated when it is in a gaseous state or a nearly particle ...
s that are released during the polymerisation of DNA, as opposed to the optical methods used in other sequencing systems. A microwell containing a template DNA strand to be sequenced is flooded with a single type of nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecu ...
. If the introduced nucleotide is complementary to the leading template nucleotide it is incorporated into the growing complementary strand. This causes the release of a hydrogen ion that triggers a hypersensitive ion sensor, which indicates that a reaction has occurred. If homopolymer repeats are present in the template sequence, multiple nucleotides will be incorporated in a single cycle. This leads to a corresponding number of released hydrogens and a proportionally higher electronic signal.
DNA nanoball sequencing
DNA nanoball sequencing is a type of high throughput sequencing technology used to determine the entire genomic sequence of an organism. The company Complete Genomics
Complete Genomics is a life sciences company that has developed and commercialized a DNA sequencing platform for human genome sequencing and analysis. This solution combines the company's proprietary human genome sequencing technology with its in ...
uses this technology to sequence samples submitted by independent researchers. The method uses rolling circle replication to amplify small fragments of genomic DNA into DNA nanoballs. Unchained sequencing by ligation is then used to determine the nucleotide sequence.reference genome
A reference genome (also known as a reference assembly) is a digital nucleic acid sequence database, assembled by scientists as a representative example of the set of genes in one idealized individual organism of a species. As they are assemble ...
difficult.
Heliscope single molecule sequencing
Heliscope sequencing is a method of single-molecule sequencing developed by Helicos Biosciences
Helicos BioSciences Corporation was a publicly traded life science company headquartered in Cambridge, Massachusetts focused on genetic analysis technologies for the research, drug discovery and diagnostic markets. The firm's Helicos Genetic Anal ...
. It uses DNA fragments with added poly-A tail adapters which are attached to the flow cell surface. The next steps involve extension-based sequencing with cyclic washes of the flow cell with fluorescently labeled nucleotides (one nucleotide type at a time, as with the Sanger method). The reads are performed by the Heliscope sequencer. The reads are short, averaging 35 bp. What made this technology especially novel was that it was the first of its class to sequence non-amplified DNA, thus preventing any read errors associated with amplification steps. In 2009 a human genome was sequenced using the Heliscope, however in 2012 the company went bankrupt.
Microfluidic Systems
There are two main microfluidic systems that are used to sequence DNA; droplet based microfluidics and digital microfluidics
Digital microfluidics (DMF) is a platform for lab-on-a-chip systems that is based upon the manipulation of microdroplets. Droplets are dispensed, moved, stored, mixed, reacted, or analyzed on a platform with a set of insulated electrodes. Digital ...
. Microfluidic devices solve many of the current limitations of current sequencing arrays.
Abate et al. studied the use of droplet-based microfluidic devices for DNA sequencing.pyrosequencing
Pyrosequencing is a method of DNA sequencing (determining the order of nucleotides in DNA) based on the "sequencing by synthesis" principle, in which the sequencing is performed by detecting the nucleotide incorporated by a DNA polymerase. Pyrosequ ...
.
Methods in development
DNA sequencing methods currently under development include reading the sequence as a DNA strand transits through nanopores (a method that is now commercial but subsequent generations such as solid-state nanopores are still in development),transmission electron microscopy
Transmission electron microscopy (TEM) is a microscopy technique in which a beam of electrons is transmitted through a specimen to form an image. The specimen is most often an ultrathin section less than 100 nm thick or a suspension on a ...
that are used to identify the positions of individual nucleotides within long DNA fragments (>5,000 bp) by nucleotide labeling with heavier elements (e.g., halogens) for visual detection and recording.
Third generation technologies aim to increase throughput and decrease the time to result and cost by eliminating the need for excessive reagents and harnessing the processivity of DNA polymerase.
Tunnelling currents DNA sequencing
Another approach uses measurements of the electrical tunnelling currents across single-strand DNA as it moves through a channel. Depending on its electronic structure, each base affects the tunnelling current differently, allowing differentiation between different bases.
The use of tunnelling currents has the potential to sequence orders of magnitude faster than ionic current methods and the sequencing of several DNA oligomers and micro-RNA has already been achieved.
Sequencing by hybridization
'' Sequencing by hybridization'' is a non-enzymatic method that uses a DNA microarray. A single pool of DNA whose sequence is to be determined is fluorescently labeled and hybridized to an array containing known sequences. Strong hybridization signals from a given spot on the array identifies its sequence in the DNA being sequenced.
This method of sequencing utilizes binding characteristics of a library of short single stranded DNA molecules (oligonucleotides), also called DNA probes, to reconstruct a target DNA sequence. Non-specific hybrids are removed by washing and the target DNA is eluted.
Sequencing with mass spectrometry
Mass spectrometry may be used to determine DNA sequences. Matrix-assisted laser desorption ionization time-of-flight mass spectrometry, or MALDI-TOF MS, has specifically been investigated as an alternative method to gel electrophoresis for visualizing DNA fragments. With this method, DNA fragments generated by chain-termination sequencing reactions are compared by mass rather than by size. The mass of each nucleotide is different from the others and this difference is detectable by mass spectrometry. Single-nucleotide mutations in a fragment can be more easily detected with MS than by gel electrophoresis alone. MALDI-TOF MS can more easily detect differences between RNA fragments, so researchers may indirectly sequence DNA with MS-based methods by converting it to RNA first.
The higher resolution of DNA fragments permitted by MS-based methods is of special interest to researchers in forensic science, as they may wish to find single-nucleotide polymorphisms in human DNA samples to identify individuals. These samples may be highly degraded so forensic researchers often prefer mitochondrial DNA
Mitochondrial DNA (mtDNA or mDNA) is the DNA located in mitochondria, cellular organelles within eukaryotic cells that convert chemical energy from food into a form that cells can use, such as adenosine triphosphate (ATP). Mitochondrial D ...
for its higher stability and applications for lineage studies. MS-based sequencing methods have been used to compare the sequences of human mitochondrial DNA from samples in a Federal Bureau of Investigation
The Federal Bureau of Investigation (FBI) is the domestic intelligence and security service of the United States and its principal federal law enforcement agency. Operating under the jurisdiction of the United States Department of Justice ...
database and from bones found in mass graves of World War I soldiers.
Early chain-termination and TOF MS methods demonstrated read lengths of up to 100 base pairs. Researchers have been unable to exceed this average read size; like chain-termination sequencing alone, MS-based DNA sequencing may not be suitable for large ''de novo'' sequencing projects. Even so, a recent study did use the short sequence reads and mass spectroscopy to compare single-nucleotide polymorphisms in pathogenic '' Streptococcus'' strains.
Microfluidic Sanger sequencing
In microfluidic Sanger sequencing
Sanger sequencing is a method of DNA sequencing that involves electrophoresis and is based on the random incorporation of chain-terminating dideoxynucleotides by DNA polymerase during in vitro DNA replication. After first being developed by Fred ...
the entire thermocycling amplification of DNA fragments as well as their separation by electrophoresis is done on a single glass wafer (approximately 10 cm in diameter) thus reducing the reagent usage as well as cost. In some instances researchers have shown that they can increase the throughput of conventional sequencing through the use of microchips. Research will still need to be done in order to make this use of technology effective.
Microscopy-based techniques
This approach directly visualizes the sequence of DNA molecules using electron microscopy. The first identification of DNA base pairs within intact DNA molecules by enzymatically incorporating modified bases, which contain atoms of increased atomic number, direct visualization and identification of individually labeled bases within a synthetic 3,272 base-pair DNA molecule and a 7,249 base-pair viral genome has been demonstrated.
RNAP sequencing
This method is based on use of RNA polymerase (RNAP), which is attached to a polystyrene
Polystyrene (PS) is a synthetic polymer made from monomers of the Aromatic hydrocarbon, aromatic hydrocarbon styrene. Polystyrene can be solid or foamed. General-purpose polystyrene is clear, hard, and brittle. It is an inexpensive resin pe ...
bead. One end of DNA to be sequenced is attached to another bead, with both beads being placed in optical traps. RNAP motion during transcription brings the beads in closer and their relative distance changes, which can then be recorded at a single nucleotide resolution. The sequence is deduced based on the four readouts with lowered concentrations of each of the four nucleotide types, similarly to the Sanger method. A comparison is made between regions and sequence information is deduced by comparing the known sequence regions to the unknown sequence regions.
''In vitro'' virus high-throughput sequencing
A method has been developed to analyze full sets of protein interactions
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respond ...
using a combination of 454 pyrosequencing and an ''in vitro'' virus mRNA display method. Specifically, this method covalently links proteins of interest to the mRNAs encoding them, then detects the mRNA pieces using reverse transcription PCRs. The mRNA may then be amplified and sequenced. The combined method was titled IVV-HiTSeq and can be performed under cell-free conditions, though its results may not be representative of ''in vivo'' conditions.
Sample preparation
The success of any DNA sequencing protocol relies upon the DNA or RNA sample extraction and preparation from the biological material of interest.
* A successful DNA extraction will yield a DNA sample with long, non-degraded strands.
* A successful RNA extraction will yield a RNA sample that should be converted to complementary DNA (cDNA) using reverse transcriptase—a DNA polymerase that synthesizes a complementary DNA based on existing strands of RNA in a PCR-like manner. Complementary DNA can then be processed the same way as genomic DNA.
After DNA or RNA extraction, samples may require further preparation depending on the sequencing method. For Sanger sequencing, either cloning procedures or PCR are required prior to sequencing. In the case of next-generation sequencing methods, library preparation is required before processing. Assessing the quality and quantity of nucleic acids both after extraction and after library preparation identifies degraded, fragmented, and low-purity samples and yields high-quality sequencing data.
The high-throughput nature of current DNA/RNA sequencing technologies has posed a challenge for sample preparation method to scale-up. Several liquid handling instruments are being used for the preparation of higher numbers of samples with a lower total hands-on time:
Development initiatives
 In October 2006, the X Prize Foundation established an initiative to promote the development of
In October 2006, the X Prize Foundation established an initiative to promote the development of full genome sequencing
Whole genome sequencing (WGS), also known as full genome sequencing, complete genome sequencing, or entire genome sequencing, is the process of determining the entirety, or nearly the entirety, of the DNA sequence of an organism's genome at a ...
technologies, called the Archon X Prize, intending to award $10 million to "the first Team that can build a device and use it to sequence 100 human genomes within 10 days or less, with an accuracy of no more than one error in every 100,000 bases sequenced, with sequences accurately covering at least 98% of the genome, and at a recurring cost of no more than $10,000 (US) per genome."
Each year the National Human Genome Research Institute, or NHGRI, promotes grants for new research and developments in genomics
Genomics is an interdisciplinary field of biology focusing on the structure, function, evolution, mapping, and editing of genomes. A genome is an organism's complete set of DNA, including all of its genes as well as its hierarchical, three-dim ...
. 2010 grants and 2011 candidates include continuing work in microfluidic, polony and base-heavy sequencing methodologies.
Computational challenges
The sequencing technologies described here produce raw data that needs to be assembled into longer sequences such as complete genomes ( sequence assembly). There are many computational challenges to achieve this, such as the evaluation of the raw sequence data which is done by programs and algorithms such as Phred and Phrap Phrap is a widely used program for DNA sequence assembly. It is part of the Phred-Phrap- Consed package.
History
Phrap was originally developed by Prof. Phil Green for the assembly of cosmids in large-scale cosmid shotgun sequencing within th ...
. Other challenges have to deal with repetitive sequences that often prevent complete genome assemblies because they occur in many places of the genome. As a consequence, many sequences may not be assigned to particular chromosome
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins ar ...
s. The production of raw sequence data is only the beginning of its detailed bioinformatical analysis.
Read trimming
Sometimes, the raw reads produced by the sequencer are correct and precise only in a fraction of their length. Using the entire read may introduce artifacts in the downstream analyses like genome assembly, SNP calling, or gene expression estimation. Two classes of trimming programs have been introduced, based on the window-based or the running-sum classes of algorithms. This is a partial list of the trimming algorithms currently available, specifying the algorithm class they belong to:
Ethical issues
Human genetics have been included within the field of bioethics
Bioethics is both a field of study and professional practice, interested in ethical issues related to health (primarily focused on the human, but also increasingly includes animal ethics), including those emerging from advances in biology, me ...
since the early 1970sMoore v. Regents of the University of California
''Moore v. Regents of the University of California'' was a landmark Supreme Court of California decision. Filed on July 9, 1990, it dealt with the issue of property rights to one's own cells taken in samples by doctors or researchers.
In 1976, Jo ...
'' (1990) ruled that individuals have no property rights to discarded cells or any profits made using these cells (for instance, as a patented cell line). However, individuals have a right to informed consent regarding removal and use of cells. Regarding the data produced through DNA sequencing, ''Moore'' gives the individual no rights to the information derived from their DNA.[Statement of Administration policy](_blank)
Executive Office of the President, Office of Management and Budget, 27 April 2007Presidential Commission for the Study of Bioethical Issues
The Presidential Commission for the Study of Bioethical Issues (the Bioethics Commission) was created by on November 24, 2009. Executive Order 13521 - ''Establishing the Presidential Commission for the Study of Bioethical Issues'', November ...
reported that existing privacy legislation for DNA sequencing data such as GINA and the Health Insurance Portability and Accountability Act were insufficient, noting that whole-genome sequencing data was particularly sensitive, as it could be used to identify not only the individual from which the data was created, but also their relatives.
/ref>
Ethical issues have also been raised by the increasing use of genetic variation screening, both in newborns, and in adults by companies such as 23andMe.anxiety
Anxiety is an emotion which is characterized by an unpleasant state of inner turmoil and includes feelings of dread over anticipated events. Anxiety is different than fear in that the former is defined as the anticipation of a future threat wh ...
in individuals who have been found to have an increased risk of disease.Time
Time is the continued sequence of existence and event (philosophy), events that occurs in an apparently irreversible process, irreversible succession from the past, through the present, into the future. It is a component quantity of various me ...
'', doctors screening an ill baby for genetic variants chose not to inform the parents of an unrelated variant linked to dementia
Dementia is a disorder which manifests as a set of related symptoms, which usually surfaces when the brain is damaged by injury or disease. The symptoms involve progressive impairments in memory, thinking, and behavior, which negatively affe ...
due to the harm it would cause to the parents.The New England Journal of Medicine
''The New England Journal of Medicine'' (''NEJM'') is a weekly medical journal published by the Massachusetts Medical Society. It is among the most prestigious peer-reviewed medical journals as well as the oldest continuously published one.
H ...
'' has shown that individuals undergoing disease risk profiling did not show increased levels of anxiety.
See also
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
Notes
References
External links
* A wikibook on next generation sequencing
{{DEFAULTSORT:Dna Sequencing
Biotechnology
DNA
Genetic mapping
Molecular biology
Molecular biology techniques
1970 introductions
1970 in biology
1970 in biotechnology
1970 in science
1998 in technology
 The first DNA sequences were obtained in the early 1970s by academic researchers using laborious methods based on
The first DNA sequences were obtained in the early 1970s by academic researchers using laborious methods based on  The foundation for sequencing proteins was first laid by the work of Frederick Sanger who by 1955 had completed the sequence of all the amino acids in
The foundation for sequencing proteins was first laid by the work of Frederick Sanger who by 1955 had completed the sequence of all the amino acids in  Several new methods for DNA sequencing were developed in the mid to late 1990s and were implemented in commercial DNA sequencers by 2000. Together these were called the "next-generation" or "second-generation" sequencing (NGS) methods, in order to distinguish them from the earlier methods, including
Several new methods for DNA sequencing were developed in the mid to late 1990s and were implemented in commercial DNA sequencers by 2000. Together these were called the "next-generation" or "second-generation" sequencing (NGS) methods, in order to distinguish them from the earlier methods, including  Large-scale sequencing often aims at sequencing very long DNA pieces, such as whole
Large-scale sequencing often aims at sequencing very long DNA pieces, such as whole  High-throughput sequencing, which includes next-generation "short-read" and third-generation "long-read" sequencing methods,"Next-generation" remains in broad use as of 2019. For instance, applies to
High-throughput sequencing, which includes next-generation "short-read" and third-generation "long-read" sequencing methods,"Next-generation" remains in broad use as of 2019. For instance, applies to 
 In this method, DNA molecules and primers are first attached on a slide or flow cell and amplified with polymerase so that local clonal DNA colonies, later coined "DNA clusters", are formed. To determine the sequence, four types of reversible terminator bases (RT-bases) are added and non-incorporated nucleotides are washed away. A camera takes images of the fluorescently labeled nucleotides. Then the dye, along with the terminal 3' blocker, is chemically removed from the DNA, allowing for the next cycle to begin. Unlike pyrosequencing, the DNA chains are extended one nucleotide at a time and image acquisition can be performed at a delayed moment, allowing for very large arrays of DNA colonies to be captured by sequential images taken from a single camera.
In this method, DNA molecules and primers are first attached on a slide or flow cell and amplified with polymerase so that local clonal DNA colonies, later coined "DNA clusters", are formed. To determine the sequence, four types of reversible terminator bases (RT-bases) are added and non-incorporated nucleotides are washed away. A camera takes images of the fluorescently labeled nucleotides. Then the dye, along with the terminal 3' blocker, is chemically removed from the DNA, allowing for the next cycle to begin. Unlike pyrosequencing, the DNA chains are extended one nucleotide at a time and image acquisition can be performed at a delayed moment, allowing for very large arrays of DNA colonies to be captured by sequential images taken from a single camera.
 Decoupling the enzymatic reaction and the image capture allows for optimal throughput and theoretically unlimited sequencing capacity. With an optimal configuration, the ultimately reachable instrument throughput is thus dictated solely by the analog-to-digital conversion rate of the camera, multiplied by the number of cameras and divided by the number of pixels per DNA colony required for visualizing them optimally (approximately 10 pixels/colony). In 2012, with cameras operating at more than 10 MHz A/D conversion rates and available optics, fluidics and enzymatics, throughput can be multiples of 1 million nucleotides/second, corresponding roughly to 1 human genome equivalent at 1x
Decoupling the enzymatic reaction and the image capture allows for optimal throughput and theoretically unlimited sequencing capacity. With an optimal configuration, the ultimately reachable instrument throughput is thus dictated solely by the analog-to-digital conversion rate of the camera, multiplied by the number of cameras and divided by the number of pixels per DNA colony required for visualizing them optimally (approximately 10 pixels/colony). In 2012, with cameras operating at more than 10 MHz A/D conversion rates and available optics, fluidics and enzymatics, throughput can be multiples of 1 million nucleotides/second, corresponding roughly to 1 human genome equivalent at 1x  The patterned array of positively charged spots is fabricated through photolithography and etching techniques followed by chemical modification to generate a sequencing flow cell. Each spot on the flow cell is approximately 250 nm in diameter, are separated by 700 nm (centre to centre) and allows easy attachment of a single negatively charged DNB to the flow cell and thus reducing under or over-clustering on the flow cell.
Sequencing is then performed by addition of an oligonucleotide probe that attaches in combination to specific sites within the DNB. The probe acts as an anchor that then allows one of four single reversibly inactivated, labelled nucleotides to bind after flowing across the flow cell. Unbound nucleotides are washed away before laser excitation of the attached labels then emit fluorescence and signal is captured by cameras that is converted to a digital output for base calling. The attached base has its terminator and label chemically cleaved at completion of the cycle. The cycle is repeated with another flow of free, labelled nucleotides across the flow cell to allow the next nucleotide to bind and have its signal captured. This process is completed a number of times (usually 50 to 300 times) to determine the sequence of the inserted piece of DNA at a rate of approximately 40 million nucleotides per second as of 2018.
The patterned array of positively charged spots is fabricated through photolithography and etching techniques followed by chemical modification to generate a sequencing flow cell. Each spot on the flow cell is approximately 250 nm in diameter, are separated by 700 nm (centre to centre) and allows easy attachment of a single negatively charged DNB to the flow cell and thus reducing under or over-clustering on the flow cell.
Sequencing is then performed by addition of an oligonucleotide probe that attaches in combination to specific sites within the DNB. The probe acts as an anchor that then allows one of four single reversibly inactivated, labelled nucleotides to bind after flowing across the flow cell. Unbound nucleotides are washed away before laser excitation of the attached labels then emit fluorescence and signal is captured by cameras that is converted to a digital output for base calling. The attached base has its terminator and label chemically cleaved at completion of the cycle. The cycle is repeated with another flow of free, labelled nucleotides across the flow cell to allow the next nucleotide to bind and have its signal captured. This process is completed a number of times (usually 50 to 300 times) to determine the sequence of the inserted piece of DNA at a rate of approximately 40 million nucleotides per second as of 2018.