|
Recombinant Gene
Recombinant DNA (rDNA) molecules are DNA molecules formed by laboratory methods of genetic recombination (such as molecular cloning) that bring together genetic material from multiple sources, creating sequences that would not otherwise be found in the genome. Recombinant DNA is the general name for a piece of DNA that has been created by combining at least two fragments from two different sources. Recombinant DNA is possible because DNA molecules from all organisms share the same chemical structure, and differ only in the nucleotide sequence within that identical overall structure. Recombinant DNA molecules are sometimes called chimeric DNA, because they can be made of material from two different species, like the mythical chimera. R-DNA technology uses palindromic sequences and leads to the production of sticky and blunt ends. The DNA sequences used in the construction of recombinant DNA molecules can originate from any species. For example, plant DNA may be joined to bacte ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Recombinant Formation Of Plasmids
Recombinant may refer to: * Recombinant organism – an organism that contains a different combination of alleles from either of its parents. * Recombinant DNA – a form of artificial DNA sequence * Recombinant protein - artificially produced (and often purified) protein * Recombinant virus – a virus formed by recombining genetic material * VRLA – a valve regulated lead acid (VRLA) battery that is also referred to as a recombinant battery *''InSoc Recombinant'' – an album by synthpop band Information Society * Recombinant Inc., - an interactive music technology company co-founded by David Cope See also *Recombination (other) {{disambiguation ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Expression
Gene expression is the process by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, protein or non-coding RNA, and ultimately affect a phenotype, as the final effect. These products are often proteins, but in non-protein-coding genes such as transfer RNA (tRNA) and small nuclear RNA (snRNA), the product is a functional non-coding RNA. Gene expression is summarized in the central dogma of molecular biology first formulated by Francis Crick in 1958, further developed in his 1970 article, and expanded by the subsequent discoveries of reverse transcription and RNA replication. The process of gene expression is used by all known life— eukaryotes (including multicellular organisms), prokaryotes (bacteria and archaea), and utilized by viruses—to generate the macromolecular machinery for life. In genetics, gene expression is the most fundamental level at which the genotype gives rise to the phenot ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Biotechnology
Biotechnology is the integration of natural sciences and engineering sciences in order to achieve the application of organisms, cells, parts thereof and molecular analogues for products and services. The term ''biotechnology'' was first used by Károly Ereky in 1919, meaning the production of products from raw materials with the aid of living organisms. Definition The concept of biotechnology encompasses a wide range of procedures for modifying living organisms according to human purposes, going back to domestication of animals, cultivation of the plants, and "improvements" to these through breeding programs that employ artificial selection and hybridization. Modern usage also includes genetic engineering as well as cell and tissue culture technologies. The American Chemical Society defines biotechnology as the application of biological organisms, systems, or processes by various industries to learning about the science of life and the improvement of the value of materi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Knockout
A gene knockout (abbreviation: KO) is a genetic technique in which one of an organism's genes is made inoperative ("knocked out" of the organism). However, KO can also refer to the gene that is knocked out or the organism that carries the gene knockout. Knockout organisms or simply knockouts are used to study gene function, usually by investigating the effect of gene loss. Researchers draw inferences from the difference between the knockout organism and normal individuals. The KO technique is essentially the opposite of a gene knock-in. Knocking out two genes simultaneously in an organism is known as a double knockout (DKO). Similarly the terms triple knockout (TKO) and quadruple knockouts (QKO) are used to describe three or four knocked out genes, respectively. However, one needs to distinguish between heterozygous and homozygous KOs. In the former, only one of two gene copies ( alleles) is knocked out, in the latter both are knocked out. Methods Knockouts are accomplished thr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
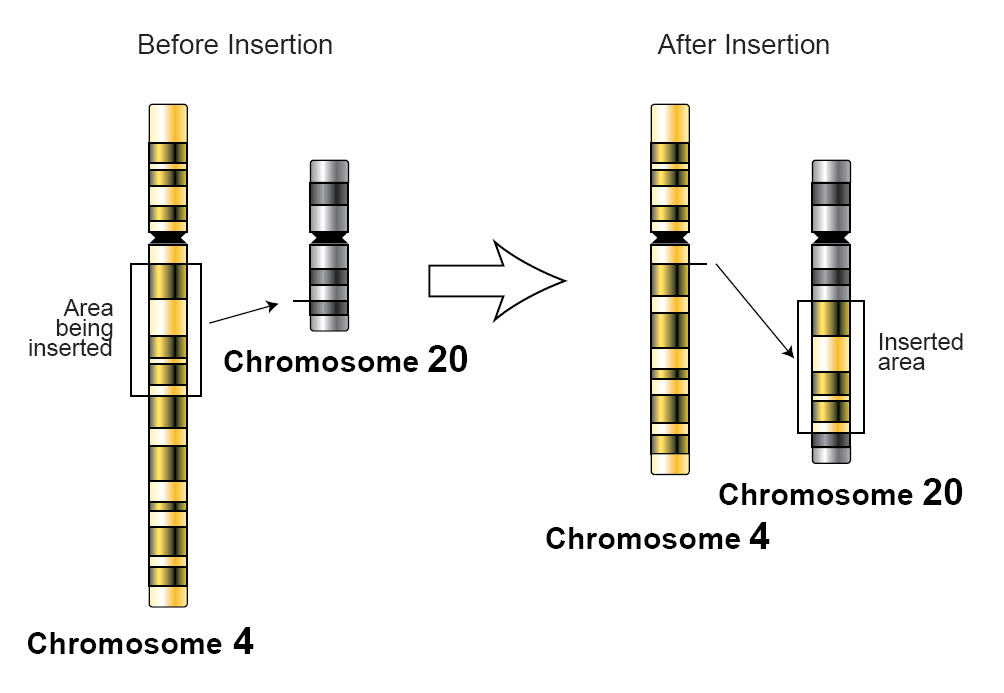
Insertion (genetics)
In genetics, an insertion (also called an insertion mutation) is the addition of one or more nucleotide base pairs into a DNA sequence. This can often happen in microsatellite regions due to the DNA polymerase slipping. Insertions can be anywhere in size from one base pair incorrectly inserted into a DNA sequence to a section of one chromosome inserted into another. The mechanism of the smallest single base insertion mutations is believed to be through base-pair separation between the template and primer strands followed by non-neighbor base stacking, which can occur locally within the DNA polymerase active site. On a chromosome level, an ''insertion'' refers to the insertion of a larger sequence into a chromosome. This can happen due to unequal crossover during meiosis. N region addition is the addition of non-coded nucleotides during recombination by terminal deoxynucleotidyl transferase. P nucleotide insertion is the insertion of palindromic sequences encoded by the ends o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Expression (biotechnology)
Protein production is the biotechnological process of generating a specific protein. It is typically achieved by the manipulation of gene expression in an organism such that it expresses large amounts of a recombinant gene. This includes the transcription of the recombinant DNA to messenger RNA (mRNA), the translation of mRNA into polypeptide chains, which are ultimately folded into functional proteins and may be targeted to specific subcellular or extracellular locations. Protein production systems (also known as expression systems) are used in the life sciences, biotechnology, and medicine. Molecular biology research uses numerous proteins and enzymes, many of which are from expression systems; particularly DNA polymerase for PCR, reverse transcriptase for RNA analysis, restriction endonucleases for cloning, and to make proteins that are screened in drug discovery as biological targets or as potential drugs themselves. There are also significant applications for expression ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
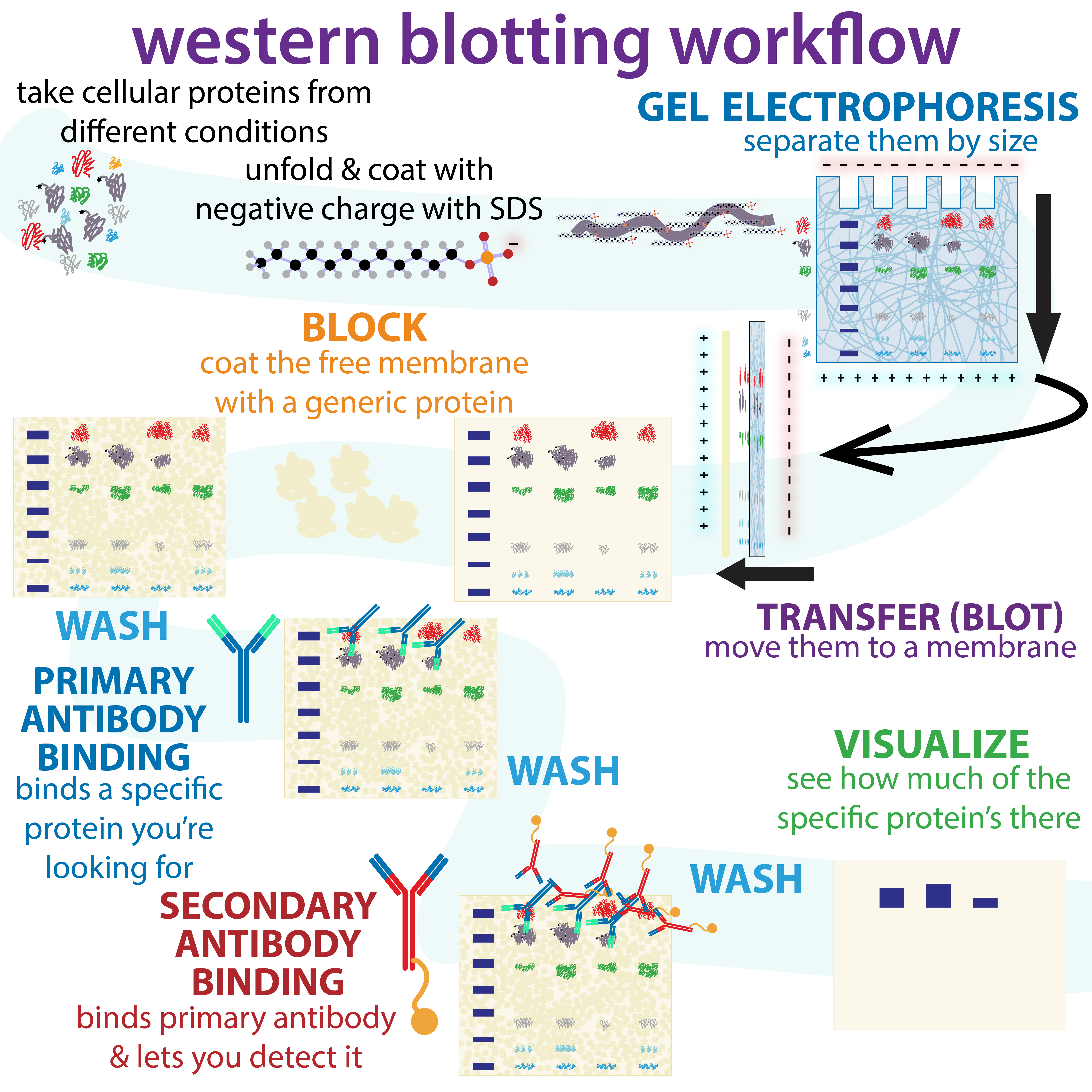
Western Blot
The western blot (sometimes called the protein immunoblot), or western blotting, is a widely used analytical technique in molecular biology and immunogenetics to detect specific proteins in a sample of tissue homogenate or extract. Besides detecting the proteins, this technique is also utilized to visualize, distinguish, and quantify the different proteins in a complicated protein combination. Western blot technique uses three elements to achieve its task of separating a specific protein from a complex: separation by size, transfer of protein to a solid support, and marking target protein using a primary and secondary antibody to visualize. A synthetic or animal-derived antibody (known as the primary antibody) is created that recognizes and binds to a specific target protein. The electrophoresis membrane is washed in a solution containing the primary antibody, before excess antibody is washed off. A secondary antibody is added which recognizes and binds to the primary antibod ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RT-PCR
Reverse transcription polymerase chain reaction (RT-PCR) is a laboratory technique combining reverse transcription of RNA into DNA (in this context called complementary DNA or cDNA) and amplification of specific DNA targets using polymerase chain reaction (PCR). It is primarily used to measure the amount of a specific RNA. This is achieved by monitoring the amplification reaction using fluorescence, a technique called real-time PCR or quantitative PCR (qPCR). Combined RT-PCR and qPCR are routinely used for analysis of gene expression and quantification of viral RNA in research and clinical settings. The close association between RT-PCR and qPCR has led to metonymic use of the term qPCR to mean RT-PCR. Such use may be confusing, as RT-PCR can be used without qPCR, for example to enable molecular cloning, sequencing or simple detection of RNA. Conversely, qPCR may be used without RT-PCR, for example to quantify the copy number of a specific piece of DNA. Nomenclature The com ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phenotype
In genetics, the phenotype () is the set of observable characteristics or traits of an organism. The term covers the organism's morphology or physical form and structure, its developmental processes, its biochemical and physiological properties, its behavior, and the products of behavior. An organism's phenotype results from two basic factors: the expression of an organism's genetic code, or its genotype, and the influence of environmental factors. Both factors may interact, further affecting phenotype. When two or more clearly different phenotypes exist in the same population of a species, the species is called polymorphic. A well-documented example of polymorphism is Labrador Retriever coloring; while the coat color depends on many genes, it is clearly seen in the environment as yellow, black, and brown. Richard Dawkins in 1978 and then again in his 1982 book '' The Extended Phenotype'' suggested that one can regard bird nests and other built structures such as ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Terminator (genetics)
In genetics, a transcription terminator is a section of nucleic acid sequence that marks the end of a gene or operon in genomic DNA during transcription. This sequence mediates transcriptional termination by providing signals in the newly synthesized transcript RNA that trigger processes which release the transcript RNA from the transcriptional complex. These processes include the direct interaction of the mRNA secondary structure with the complex and/or the indirect activities of recruited termination factors. Release of the transcriptional complex frees RNA polymerase and related transcriptional machinery to begin transcription of new mRNAs. In prokaryotes Two classes of transcription terminators, Rho-dependent and Rho-independent, have been identified throughout prokaryotic genomes. These widely distributed sequences are responsible for triggering the end of transcription upon normal completion of gene or operon transcription, mediating early termination of transcripts a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Promoter (biology)
In genetics, a promoter is a sequence of DNA to which proteins bind to initiate transcription of a single RNA transcript from the DNA downstream of the promoter. The RNA transcript may encode a protein (mRNA), or can have a function in and of itself, such as tRNA or rRNA. Promoters are located near the transcription start sites of genes, upstream on the DNA (towards the 5' region of the sense strand). Promoters can be about 100–1000 base pairs long, the sequence of which is highly dependent on the gene and product of transcription, type or class of RNA polymerase recruited to the site, and species of organism. Promoters control gene expression in bacteria and eukaryotes. RNA polymerase must attach to DNA near a gene for transcription to occur. Promoter DNA sequences provide an enzyme binding site. The -10 sequence is TATAAT. -35 sequences are conserved on average, but not in most promoters. Artificial promoters with conserved -10 and -35 elements transcribe more slowly. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |