|
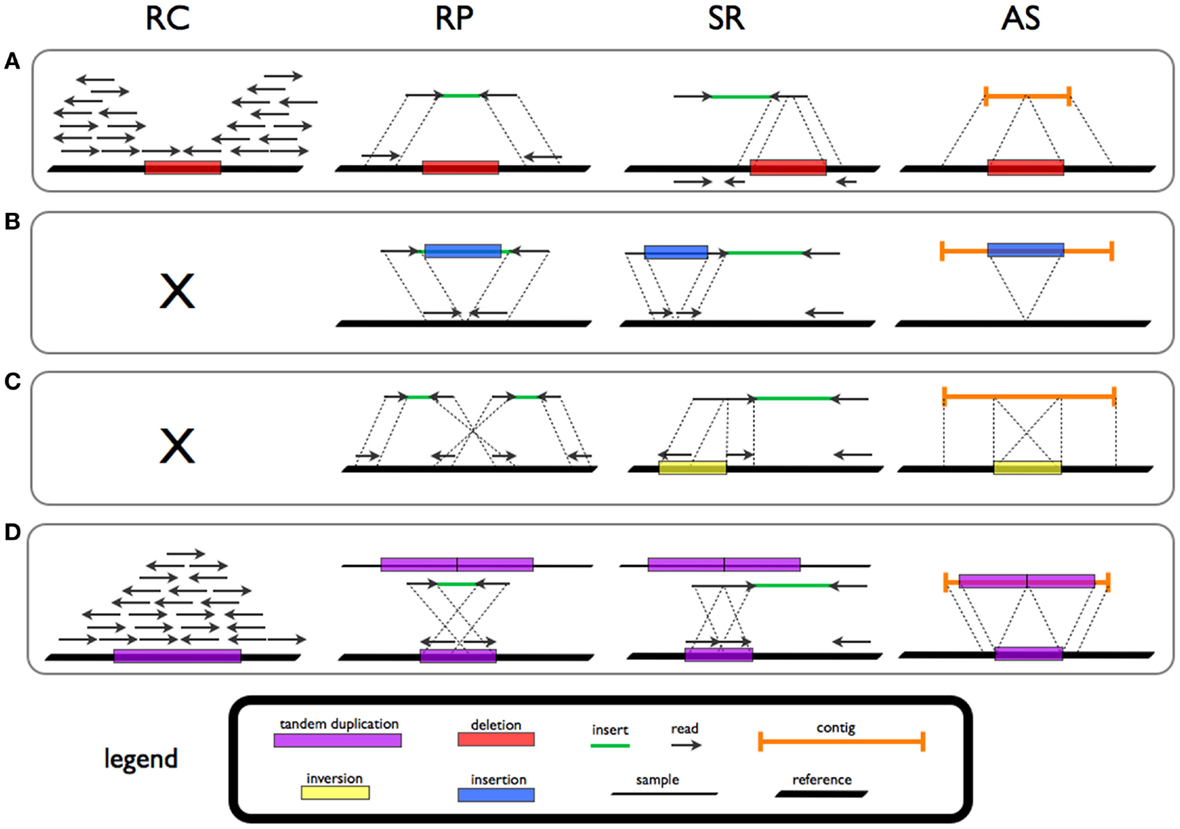
Structural Variation
Genomic structural variation is the variation in structure of an organism's chromosome. It consists of many kinds of variation in the genome of one species, and usually includes microscopic and submicroscopic types, such as deletions, duplications, copy-number variants, insertions, inversions and translocations. Originally, a structure variation affects a sequence length about 1kb to 3Mb, which is larger than SNPs and smaller than chromosome abnormality (though the definitions have some overlap). However, the operational range of structural variants has widened to include events > 50bp. The definition of structural variation does not imply anything about frequency or phenotypical effects. Many structural variants are associated with genetic diseases, however many are not. Recent research about SVs indicates that SVs are more difficult to detect than SNPs. Approximately 13% of the human genome is defined as structurally variant in the normal population, and there are at least ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
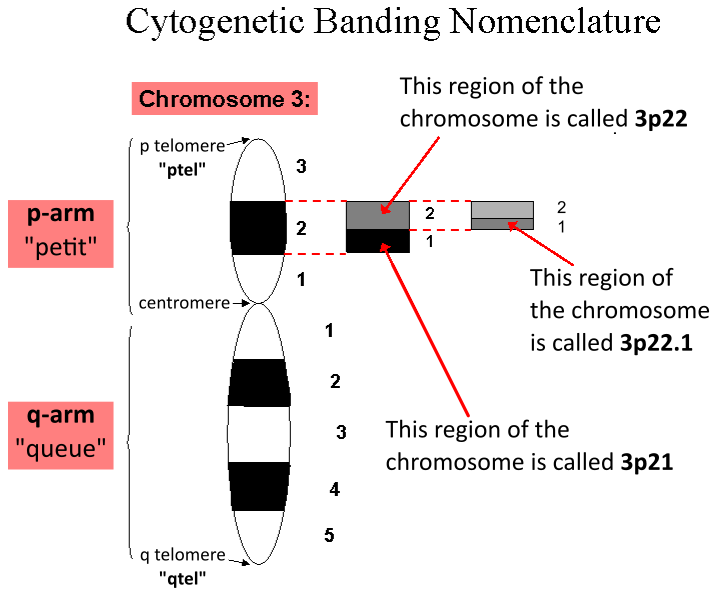
Chromosome
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins are the histones. These proteins, aided by chaperone proteins, bind to and condense the DNA molecule to maintain its integrity. These chromosomes display a complex three-dimensional structure, which plays a significant role in transcriptional regulation. Chromosomes are normally visible under a light microscope only during the metaphase of cell division (where all chromosomes are aligned in the center of the cell in their condensed form). Before this happens, each chromosome is duplicated (S phase), and both copies are joined by a centromere, resulting either in an X-shaped structure (pictured above), if the centromere is located equatorially, or a two-arm structure, if the centromere is located distally. The joined copies are now ca ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Locus (genetics)
In genetics, a locus (plural loci) is a specific, fixed position on a chromosome where a particular gene or genetic marker is located. Each chromosome carries many genes, with each gene occupying a different position or locus; in humans, the total number of protein-coding genes in a complete haploid set of 23 chromosomes is estimated at 19,000–20,000. Genes may possess multiple variants known as alleles, and an allele may also be said to reside at a particular locus. Diploid and polyploid cells whose chromosomes have the same allele at a given locus are called homozygous with respect to that locus, while those that have different alleles at a given locus are called heterozygous. The ordered list of loci known for a particular genome is called a gene map. Gene mapping is the process of determining the specific locus or loci responsible for producing a particular phenotype or biological trait. Association mapping, also known as "linkage disequilibrium mapping", is a method ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Unequal Crossing Over
Unequal crossing over is a type of gene duplication or deletion event that deletes a sequence in one strand and replaces it with a duplication from its sister chromatid in mitosis or from its homologous chromosome during meiosis. It is a type of chromosomal crossover between homologous sequences that are not paired precisely. Normally genes are responsible for occurrence of crossing over. It exchanges sequences of different links between chromosomes. Along with gene conversion, it is believed to be the main driver for the generation of gene duplications and is a source of mutation in the genome. Mechanisms During meiosis, the duplicated chromosomes ( chromatids) in eukaryotic organisms are attached to each other in the centromere region and are thus paired. The maternal and paternal chromosomes then align alongside each other. During this time, recombination can take place via crossing over of sections of the paternal and maternal chromatids and leads to reciprocal recombinatio ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |