|
Minor Allele Frequency
Minor allele frequency (MAF) is the frequency at which the ''second most common'' allele occurs in a given population. They play a surprising role in heritability since MAF variants which occur only once, known as "singletons", drive an enormous amount of selection. Single nucleotide polymorphisms (SNPs) with a minor allele frequency of 0.05 (5%) or greater were targeted by the HapMap project. MAF is widely used in population genetics studies because it provides information to differentiate between common and rare variants in the population. As an example, a 2015 study sequenced the whole genomes of Sardinian individuals. The authors classified the variants found in the study in three classes according to their MAF. It was observed that rare variants (MAF 0.05) in this population. Interpreting MAF data 1. Introduce the reference of a SNP of interest, as an example: rs429358, in a database (dbSNP or other). 2. Find MAF/MinorAlleleCount link. MAF/MinorAlleleCount: C=0.1506/ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Allele
An allele (, ; ; modern formation from Greek ἄλλος ''állos'', "other") is a variation of the same sequence of nucleotides at the same place on a long DNA molecule, as described in leading textbooks on genetics and evolution. ::"The chromosomal or genomic location of a gene or any other genetic element is called a locus (plural: loci) and alternative DNA sequences at a locus are called alleles." The simplest alleles are single nucleotide polymorphisms (SNP). but they can also be insertions and deletions of up to several thousand base pairs. Popular definitions of 'allele' typically refer only to different alleles within genes. For example, the ABO blood grouping is controlled by the ABO gene, which has six common alleles (variants). In population genetics, nearly every living human's phenotype for the ABO gene is some combination of just these six alleles. Most alleles observed result in little or no change in the function of the gene product it codes for. However, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Heritability
Heritability is a statistic used in the fields of breeding and genetics that estimates the degree of ''variation'' in a phenotypic trait in a population that is due to genetic variation between individuals in that population. The concept of heritability can be expressed in the form of the following question: "What is the proportion of the variation in a given trait within a population that is ''not'' explained by the environment or random chance?" Other causes of measured variation in a trait are characterized as environmental factors, including observational error. In human studies of heritability these are often apportioned into factors from "shared environment" and "non-shared environment" based on whether they tend to result in persons brought up in the same household being more or less similar to persons who were not. Heritability is estimated by comparing individual phenotypic variation among related individuals in a population, by examining the association between ind ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Single Nucleotide Polymorphism
In genetics, a single-nucleotide polymorphism (SNP ; plural SNPs ) is a germline substitution of a single nucleotide at a specific position in the genome. Although certain definitions require the substitution to be present in a sufficiently large fraction of the population (e.g. 1% or more), many publications do not apply such a frequency threshold. For example, at a specific base position in the human genome, the G nucleotide may appear in most individuals, but in a minority of individuals, the position is occupied by an A. This means that there is a SNP at this specific position, and the two possible nucleotide variations – G or A – are said to be the alleles for this specific position. SNPs pinpoint differences in our susceptibility to a wide range of diseases, for example age-related macular degeneration (a common SNP in the CFH gene is associated with increased risk of the disease) or nonalcoholic fatty liver disease (a SNP in the PNPLA3 gene is associated with incr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
HapMap
The International HapMap Project was an organization that aimed to develop a haplotype map (HapMap) of the human genome, to describe the common patterns of human genetic variation. HapMap is used to find genetic variants affecting health, disease and responses to drugs and environmental factors. The information produced by the project is made freely available for research. The International HapMap Project is a collaboration among researchers at academic centers, non-profit biomedical research groups and private companies in Canada, China (including Hong Kong), Japan, Nigeria, the United Kingdom, and the United States. It officially started with a meeting on October 27 to 29, 2002, and was expected to take about three years. It comprises two phases; the complete data obtained in Phase I were published on 27 October 2005. The analysis of the Phase II dataset was published in October 2007. The Phase III dataset was released in spring 2009 and the publication presenting the final resul ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Population Genetics
Population genetics is a subfield of genetics that deals with genetic differences within and between populations, and is a part of evolutionary biology. Studies in this branch of biology examine such phenomena as adaptation, speciation, and population structure. Population genetics was a vital ingredient in the emergence of the modern evolutionary synthesis. Its primary founders were Sewall Wright, J. B. S. Haldane and Ronald Fisher, who also laid the foundations for the related discipline of quantitative genetics. Traditionally a highly mathematical discipline, modern population genetics encompasses theoretical, laboratory, and field work. Population genetic models are used both for statistical inference from DNA sequence data and for proof/disproof of concept. What sets population genetics apart from newer, more phenotypic approaches to modelling evolution, such as evolutionary game theory and adaptive dynamics, is its emphasis on such genetic phenomena as dominance, epi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding genes, other functional regions of the genome such as regulatory sequences (see non-coding DNA), and often a substantial fraction of 'junk' DNA with no evident function. Almost all eukaryotes have mitochondria and a small mitochondrial genome. Algae and plants also contain chloroplasts with a chloroplast genome. The study of the genome is called genomics. The genomes of many organisms have been sequenced and various regions have been annotated. The International Human Genome Project reported the sequence of the genome for ''Homo sapiens'' in 200The Human Genome Project although the initial "finished" sequence was missing 8% of the genome consisting mostly of repetitive sequences. With advancements in technology that could handle sequenci ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sardinian People
The Sardinians, or Sards ( sc, Sardos or ; Italian and Sassarese: ''Sardi''; Gallurese: ''Saldi''), are a Romance language-speaking ethnic group native to Sardinia, from which the western Mediterranean island and autonomous region of Italy derives its name. Etymology Not much can be gathered from the classical literature about the origins of the Sardinian people. The ethnonym "S(a)rd" belongs to the Pre-Indo-European linguistic substratum, and whilst they might have derived from the Iberians, the accounts of the old authors differ greatly in this respect. The oldest written attestation of the ethnonym is on the Nora stone, where the word ''Šrdn'' (''Shardan'') bears witness to its original existence by the time the Phoenician merchants first arrived to the Sardinian shores. According to Timaeus, one of Plato's dialogues, Sardinia and its people as well, the "Sardonioi" or "Sardianoi" (''Σαρδονιοί'' or ''Σαρδιανοί''), might have been named after "Sardò" ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Rs429358
Apolipoprotein E (APOE) is a protein involved in the metabolism of fats in the body of mammals. A subtype is implicated in Alzheimer's disease and cardiovascular disease. APOE belongs to a family of fat-binding proteins called apolipoproteins. In the circulation, it is present as part of several classes of lipoprotein particles, including chylomicron remnants, VLDL, Intermediate-density lipoprotein, IDL, and some High-density lipoprotein, HDL. APOE interacts significantly with the LDL receptor, low-density lipoprotein receptor (LDLR), which is essential for the normal processing (catabolism) of triglyceride-rich lipoproteins. In peripheral tissues, APOE is primarily produced by the liver and macrophages, and mediates cholesterol metabolism. In the central nervous system, APOE is mainly produced by astrocytes and transports cholesterol to neurons via APOE receptors, which are members of the low density lipoprotein receptor gene family. APOE is the principal cholesterol carrier in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
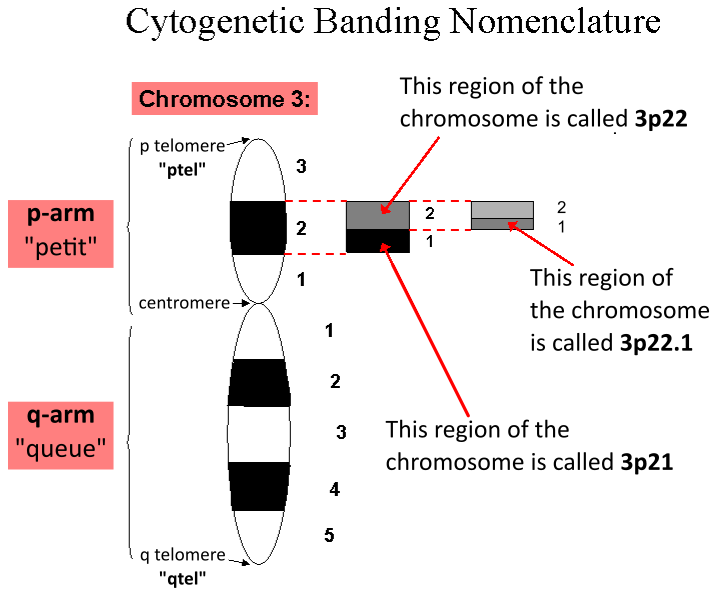
Locus (genetics)
In genetics, a locus (plural loci) is a specific, fixed position on a chromosome where a particular gene or genetic marker is located. Each chromosome carries many genes, with each gene occupying a different position or locus; in humans, the total number of protein-coding genes in a complete haploid set of 23 chromosomes is estimated at 19,000–20,000. Genes may possess multiple variants known as alleles, and an allele may also be said to reside at a particular locus. Diploid and polyploid cells whose chromosomes have the same allele at a given locus are called homozygous with respect to that locus, while those that have different alleles at a given locus are called heterozygous. The ordered list of loci known for a particular genome is called a gene map. Gene mapping is the process of determining the specific locus or loci responsible for producing a particular phenotype or biological trait. Association mapping, also known as "linkage disequilibrium mapping", is a method of ma ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Allele Frequency
Allele frequency, or gene frequency, is the relative frequency of an allele (variant of a gene) at a particular locus in a population, expressed as a fraction or percentage. Specifically, it is the fraction of all chromosomes in the population that carry that allele over the total population or sample size. Microevolution is the change in allele frequencies that occurs over time within a population. Given the following: # A particular locus on a chromosome and a given allele at that locus # A population of ''N'' individuals with ploidy ''n'', i.e. an individual carries ''n'' copies of each chromosome in their somatic cells (e.g. two chromosomes in the cells of diploid species) # The allele exists in ''i'' chromosomes in the population then the allele frequency is the fraction of all the occurrences ''i'' of that allele and the total number of chromosome copies across the population, ''i''/(''nN''). The allele frequency is distinct from the genotype frequency, although they are ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |