|
Ligand (biochemistry)
In biochemistry and pharmacology, a ligand is a substance that forms a complex with a biomolecule to serve a biological purpose. The etymology stems from ''ligare'', which means 'to bind'. In protein-ligand binding, the ligand is usually a molecule which produces a signal by binding to a site on a target protein. The binding typically results in a change of conformational isomerism (conformation) of the target protein. In DNA-ligand binding studies, the ligand can be a small molecule, ion, or protein which binds to the DNA double helix. The relationship between ligand and binding partner is a function of charge, hydrophobicity, and molecular structure. Binding occurs by intermolecular forces, such as ionic bonds, hydrogen bonds and Van der Waals forces. The association or docking is actually reversible through dissociation. Measurably irreversible covalent bonding between a ligand and target molecule is atypical in biological systems. In contrast to the definition of lig ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Myoglobin And Heme
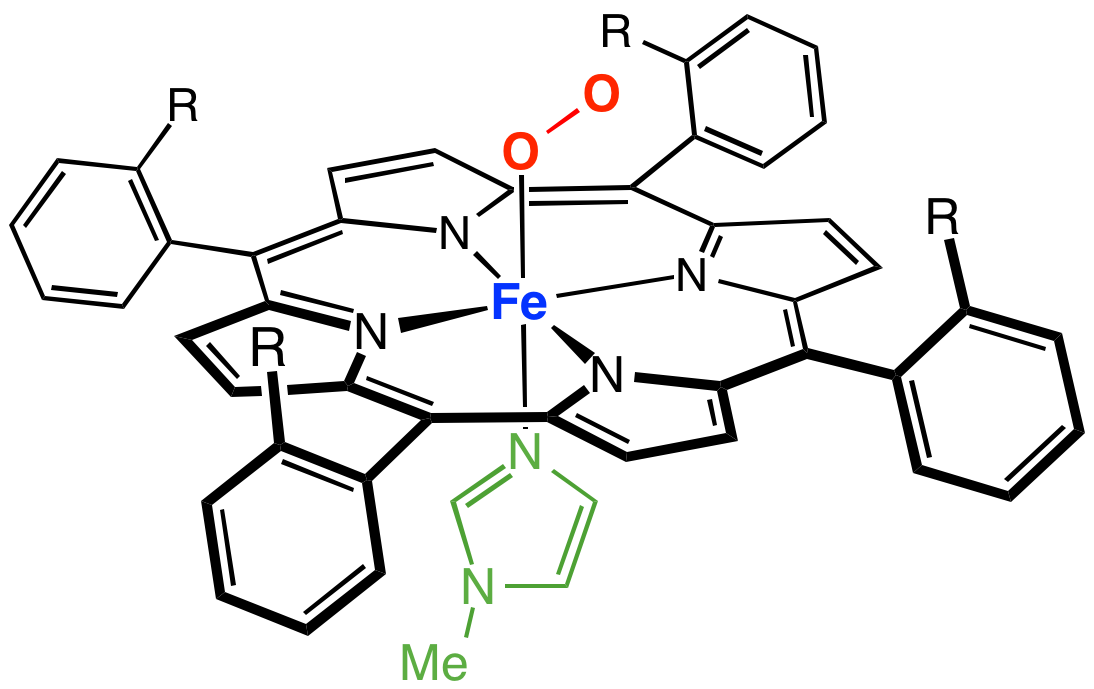
Myoglobin (symbol Mb or MB) is an iron- and oxygen-binding protein found in the cardiac and skeletal muscle tissue of vertebrates in general and in almost all mammals. Myoglobin is distantly related to hemoglobin. Compared to hemoglobin, myoglobin has a higher affinity for oxygen and does not have cooperative binding with oxygen like hemoglobin does. In humans, myoglobin is only found in the bloodstream after muscle injury. (Google books link is the 2008 edition) High concentrations of myoglobin in muscle cells allow organisms to hold their breath for a longer period of time. Diving mammals such as whales and seals have muscles with particularly high abundance of myoglobin. Myoglobin is found in Type I muscle, Type II A, and Type II B; although many texts consider myoglobin not to be found in smooth muscle, this has proved erroneous: there is also myoglobin in smooth muscle cells. Myoglobin was the first protein to have its three-dimensional structure revealed by X-ray crystal ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Covalent
A covalent bond is a chemical bond that involves the sharing of electrons to form electron pairs between atoms. These electron pairs are known as shared pairs or bonding pairs. The stable balance of attractive and repulsive forces between atoms, when they share electrons, is known as covalent bonding. For many molecules, the sharing of electrons allows each atom to attain the equivalent of a full valence shell, corresponding to a stable electronic configuration. In organic chemistry, covalent bonding is much more common than ionic bonding. Covalent bonding also includes many kinds of interactions, including σ-bonding, π-bonding, metal-to-metal bonding, agostic interactions, bent bonds, three-center two-electron bonds and three-center four-electron bonds. The term ''covalent bond'' dates from 1939. The prefix ''co-'' means ''jointly, associated in action, partnered to a lesser degree, '' etc.; thus a "co-valent bond", in essence, means that the atoms share " valence", such a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Host–guest Chemistry
In supramolecular chemistry, host–guest chemistry describes complexes that are composed of two or more molecules or ions that are held together in unique structural relationships by forces other than those of full covalent bonds. Host–guest chemistry encompasses the idea of molecular recognition and interactions through non-covalent bonding. Non-covalent bonding is critical in maintaining the 3D structure of large molecules, such as proteins and is involved in many biological processes in which large molecules bind specifically but transiently to one another. Although non-covalent interactions could be roughly divided into those with more electrostatic or dispersive contributions, there are few commonly mentioned types of non-covalent interactions: ionic bonding, hydrogen bonding, van der Waals forces and hydrophobic interactions. Overview Host–guest chemistry is a branch of supramolecular chemistry in which a host molecule forms a chemical compound with a guest molecul ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Neurotransmitter
A neurotransmitter is a signaling molecule secreted by a neuron to affect another cell across a synapse. The cell receiving the signal, any main body part or target cell, may be another neuron, but could also be a gland or muscle cell. Neurotransmitters are released from synaptic vesicles into the synaptic cleft where they are able to interact with neurotransmitter receptors on the target cell. The neurotransmitter's effect on the target cell is determined by the receptor it binds. Many neurotransmitters are synthesized from simple and plentiful precursors such as amino acids, which are readily available and often require a small number of biosynthetic steps for conversion. Neurotransmitters are essential to the function of complex neural systems. The exact number of unique neurotransmitters in humans is unknown, but more than 100 have been identified. Common neurotransmitters include glutamate, GABA, acetylcholine, glycine and norepinephrine. Mechanism and cycle Synthes ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Lipid-gated Ion Channels
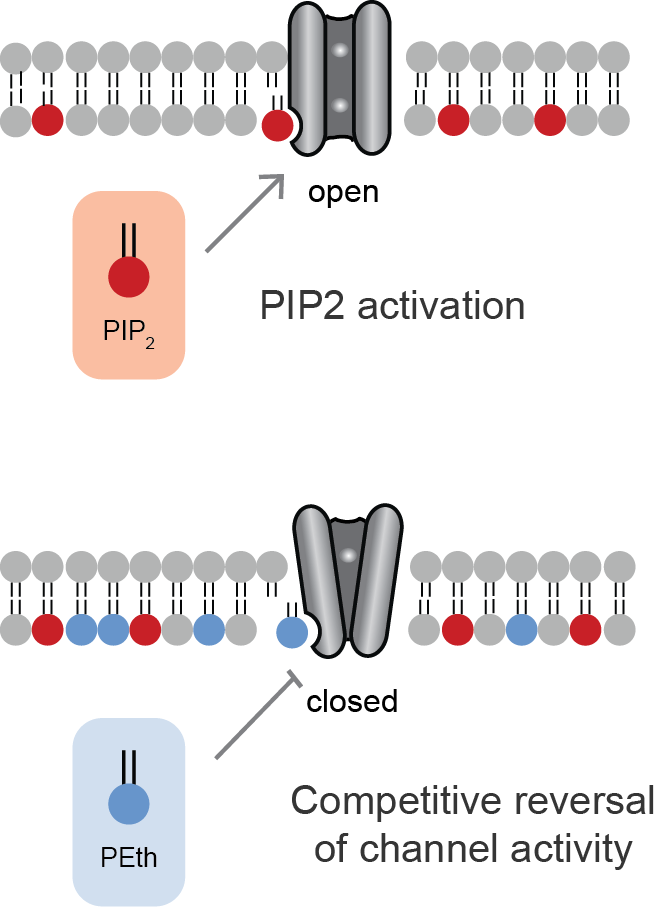
Lipid-gated ion channels are a class of ion channels whose conductance of ions through the membrane depends directly on lipids. Classically the lipids are membrane resident anionic signaling lipids that bind to the transmembrane domain on the inner leaflet of the plasma membrane with properties of a classic ligand. Other classes of lipid-gated channels include the mechanosensitive ion channels that respond to lipid tension, thickness, and hydrophobic mismatch. A lipid ligand differs from a lipid cofactor in that a ligand derives its function by dissociating from the channel while a cofactor typically derives its function by remaining bound. PIP2-gated channels Phosphatidylinositol 4,5-bisphosphate (PIP2) was the first and remains the best studied lipid to gate ion channels. PIP2 is a cell membrane lipid, and its role in gating ion channels represents a novel role for the molecule. Kir channels: PIP2 binds to and directly activates inwardly rectifying potassium channels (Kir) ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Activator (proteomics)
Enzyme activators are molecules that bind to enzymes and increase their activity. They are the opposite of enzyme inhibitors. These molecules are often involved in the allosteric regulation of enzymes in the control of metabolism. An example of an enzyme activator working in this way is fructose 2,6-bisphosphate, which activates phosphofructokinase 1 and increases the rate of glycolysis in response to the hormone glucagon. In some cases, when a substrate binds to one catalytic subunit of an enzyme, this can trigger an increase in the substrate affinity as well as catalytic activity in the enzyme's other subunits, and thus the substrate acts as an activator. Examples Hexokinase-I Hexokinase-I (HK-I) is an enzyme activator because it draws glucose into the glycolysis pathway. Its function is to phosphorylate glucose releasing glucose-6-phosphate (G6P) as the product. HK-I not only signals the activation of glucose into glycolysis but also maintains a low glucose concentration ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Enzyme Inhibitor
An enzyme inhibitor is a molecule that binds to an enzyme and blocks its activity. Enzymes are proteins that speed up chemical reactions necessary for life, in which substrate molecules are converted into products. An enzyme facilitates a specific chemical reaction by binding the substrate to its active site, a specialized area on the enzyme that accelerates the most difficult step of the reaction. An enzyme inhibitor stops ("inhibits") this process, either by binding to the enzyme's active site (thus preventing the substrate itself from binding) or by binding to another site on the enzyme such that the enzyme's catalysis of the reaction is blocked. Enzyme inhibitors may bind reversibly or irreversibly. Irreversible inhibitors form a chemical bond with the enzyme such that the enzyme is inhibited until the chemical bond is broken. By contrast, reversible inhibitors bind non-covalently and may spontaneously leave the enzyme, allowing the enzyme to resume its function. Reve ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Substrate (biochemistry)
In chemistry, the term substrate is highly context-dependent. Broadly speaking, it can refer either to a chemical species being observed in a chemical reaction, or to a surface on which other chemical reactions or microscopy are performed. In the former sense, a reagent is added to the ''substrate'' to generate a product through a chemical reaction. The term is used in a similar sense in synthetic and organic chemistry, where the substrate is the chemical of interest that is being modified. In biochemistry, an enzyme substrate is the material upon which an enzyme acts. When referring to Le Chatelier's principle, the substrate is the reagent whose concentration is changed. ;Spontaneous reaction : :*Where S is substrate and P is product. ;Catalysed reaction : :*Where S is substrate, P is product and C is catalyst. In the latter sense, it may refer to a surface on which other chemical reactions are performed or play a supporting role in a variety of spectroscopic and microscop ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Receptor (biochemistry)
In biochemistry and pharmacology, receptors are chemical structures, composed of protein, that receive and transduce signals that may be integrated into biological systems. These signals are typically chemical messengers which bind to a receptor and cause some form of cellular/tissue response, e.g. a change in the electrical activity of a cell. There are three main ways the action of the receptor can be classified: relay of signal, amplification, or integration. Relaying sends the signal onward, amplification increases the effect of a single ligand, and integration allows the signal to be incorporated into another biochemical pathway. Receptor proteins can be classified by their location. Transmembrane receptors include ligand-gated ion channels, G protein-coupled receptors, and enzyme-linked hormone receptors. Intracellular receptors are those found inside the cell, and include cytoplasmic receptors and nuclear receptors. A molecule that binds to a receptor is called a ligand ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hemoglobin
Hemoglobin (haemoglobin BrE) (from the Greek word αἷμα, ''haîma'' 'blood' + Latin ''globus'' 'ball, sphere' + ''-in'') (), abbreviated Hb or Hgb, is the iron-containing oxygen-transport metalloprotein present in red blood cells (erythrocytes) of almost all vertebrates (the exception being the fish family Channichthyidae) as well as the tissues of some invertebrates. Hemoglobin in blood carries oxygen from the respiratory organs (''e.g.'' lungs or gills) to the rest of the body (''i.e.'' tissues). There it releases the oxygen to permit aerobic respiration to provide energy to power functions of an organism in the process called metabolism. A healthy individual human has 12to 20grams of hemoglobin in every 100mL of blood. In mammals, the chromoprotein makes up about 96% of the red blood cells' dry content (by weight), and around 35% of the total content (including water). Hemoglobin has an oxygen-binding capacity of 1.34mL O2 per gram, which increases the total blood oxygen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Metal
A metal (from Greek μέταλλον ''métallon'', "mine, quarry, metal") is a material that, when freshly prepared, polished, or fractured, shows a lustrous appearance, and conducts electricity and heat relatively well. Metals are typically ductile (can be drawn into wires) and malleable (they can be hammered into thin sheets). These properties are the result of the ''metallic bond'' between the atoms or molecules of the metal. A metal may be a chemical element such as iron; an alloy such as stainless steel; or a molecular compound such as polymeric sulfur nitride. In physics, a metal is generally regarded as any substance capable of conducting electricity at a temperature of absolute zero. Many elements and compounds that are not normally classified as metals become metallic under high pressures. For example, the nonmetal iodine gradually becomes a metal at a pressure of between 40 and 170 thousand times atmospheric pressure. Equally, some materials regarded as metals ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |