|
Genetic Analyses
Genetic analysis is the overall process of studying and researching in fields of science that involve genetics and molecular biology. There are a number of applications that are developed from this research, and these are also considered parts of the process. The base system of analysis revolves around general genetics. Basic studies include identification of genes and inherited disorders. This research has been conducted for centuries on both a large-scale physical observation basis and on a more microscopic scale. Genetic analysis can be used generally to describe methods both used in and resulting from the sciences of genetics and molecular biology, or to applications resulting from this research. Genetic analysis may be done to identify genetic/inherited disorders and also to make a differential diagnosis in certain somatic diseases such as cancer. Genetic analyses of cancer include detection of mutations, fusion genes, and DNA copy number changes. History of genetic analysi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
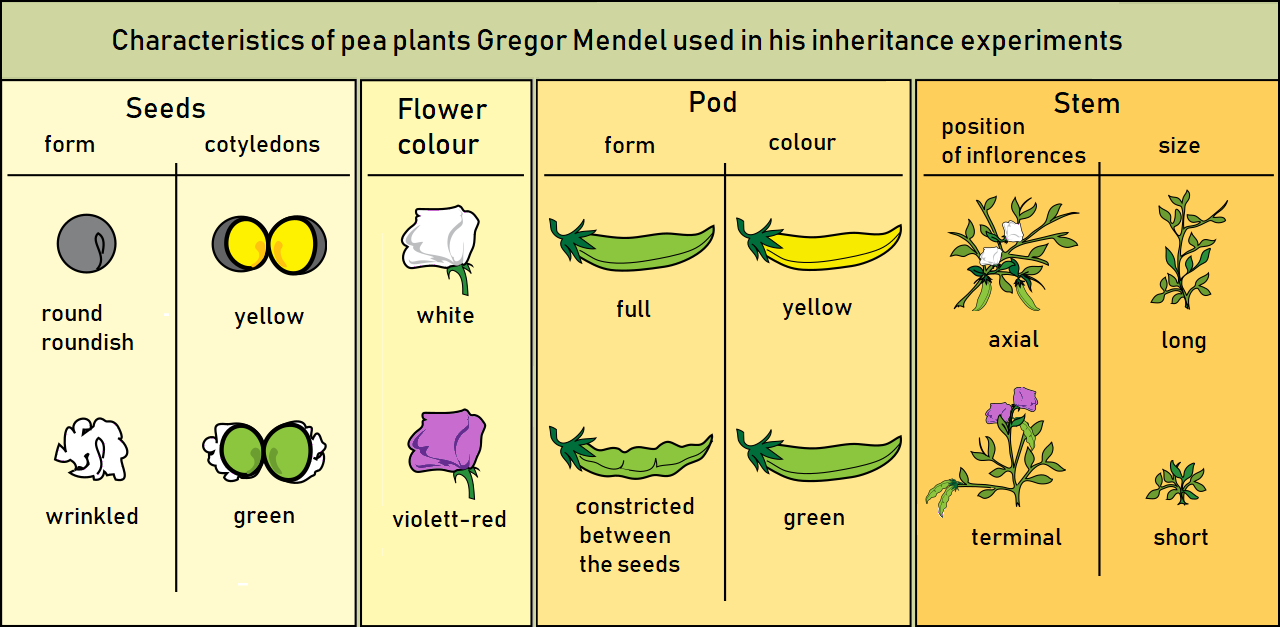
Trait (biology)
A phenotypic trait, simply trait, or character state is a distinct variant of a phenotypic characteristic of an organism; it may be either inherited or determined environmentally, but typically occurs as a combination of the two.Lawrence, Eleanor (2005) ''Henderson's Dictionary of Biology''. Pearson, Prentice Hall. For example, having eye color is a ''character'' of an organism, while blue, brown and hazel versions of eye colour are ''traits''. The term ''trait'' is generally used in genetics, often to describe phenotypic expression of different combinations of alleles in different individual organisms within a single population, such as the famous purple vs. white flower coloration in Gregor Mendel's pea plants. By contrast, in systematics, the term is ''character state'' is employed to describe features that represent fixed diagnostic differences among taxa, such as the absence of tails in great apes, relative to other primate groups. Definition A phenotypic trait is a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Fusion Gene
A fusion gene is a hybrid gene formed from two previously independent genes. It can occur as a result of translocation, interstitial deletion, or chromosomal inversion. Fusion genes have been found to be prevalent in all main types of human neoplasia. The identification of these fusion genes play a prominent role in being a diagnostic and prognostic marker. History The first fusion gene was described in cancer cells in the early 1980s. The finding was based on the discovery in 1960 by Peter Nowell and David Hungerford in Philadelphia of a small abnormal marker chromosome in patients with chronic myeloid leukemia—the first consistent chromosome abnormality detected in a human malignancy, later designated the Philadelphia chromosome. In 1973, Janet Rowley in Chicago showed that the Philadelphia chromosome had originated through a translocation between chromosomes 9 and 22, and not through a simple deletion of chromosome 22 as was previously thought. Several investigators in t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Electrophoresis Apparatus
Electrophoresis, from Ancient Greek ἤλεκτρον (ḗlektron, "amber") and φόρησις (phórēsis, "the act of bearing"), is the motion of dispersed particles relative to a fluid under the influence of a spatially uniform electric field. Electrophoresis of positively charged particles (cations) is sometimes called cataphoresis, while electrophoresis of negatively charged particles (anions) is sometimes called anaphoresis. The electrokinetic phenomenon of electrophoresis was observed for the first time in 1807 by Russian professors Peter Ivanovich Strakhov and Ferdinand Frederic Reuss at Moscow University, who noticed that the application of a constant electric field caused clay particles dispersed in water to migrate. It is ultimately caused by the presence of a charged interface between the particle surface and the surrounding fluid. It is the basis for analytical techniques used in chemistry for separating molecules by size, charge, or binding affinity. Electropho ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Fluorescence In Situ Hybridisation
Fluorescence ''in situ'' hybridization (FISH) is a molecular cytogenetic technique that uses fluorescent probes that bind to only particular parts of a nucleic acid sequence with a high degree of sequence complementarity. It was developed by biomedical researchers in the early 1980s to detect and localize the presence or absence of specific DNA sequences on chromosomes. Fluorescence microscopy can be used to find out where the fluorescent probe is bound to the chromosomes. FISH is often used for finding specific features in DNA for use in genetic counseling, medicine, and species identification. FISH can also be used to detect and localize specific RNA targets (mRNA, lncRNA and miRNA) in cells, circulating tumor cells, and tissue samples. In this context, it can help define the spatial-temporal patterns of gene expression within cells and tissues. Probes – RNA and DNA In biology, a probe is a single strand of DNA or RNA that is complementary to a nucleotide sequence of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Karyotyping
A karyotype is the general appearance of the complete set of metaphase chromosomes in the cells of a species or in an individual organism, mainly including their sizes, numbers, and shapes. Karyotyping is the process by which a karyotype is discerned by determining the chromosome complement of an individual, including the number of chromosomes and any abnormalities. A karyogram or idiogram is a graphical depiction of a karyotype, wherein chromosomes are organized in pairs, ordered by size and position of centromere for chromosomes of the same size. Karyotyping generally combines light microscopy and photography, and results in a photomicrographic (or simply micrographic) karyogram. In contrast, a schematic karyogram is a designed graphic representation of a karyotype. In schematic karyograms, just one of the sister chromatids of each chromosome is generally shown for brevity, and in reality they are generally so close together that they look as one on photomicrographs as well u ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cytogenetics
Cytogenetics is essentially a branch of genetics, but is also a part of cell biology/cytology (a subdivision of human anatomy), that is concerned with how the chromosomes relate to cell behaviour, particularly to their behaviour during mitosis and meiosis. Techniques used include karyotyping, analysis of G-banded chromosomes, other cytogenetic banding techniques, as well as molecular cytogenetics such as fluorescent ''in situ'' hybridization (FISH) and comparative genomic hybridization (CGH). History Beginnings Chromosomes were first observed in plant cells by Carl Nägeli in 1842. Their behavior in animal (salamander) cells was described by Walther Flemming, the discoverer of mitosis, in 1882. The name was coined by another German anatomist, von Waldeyer in 1888. The next stage took place after the development of genetics in the early 20th century, when it was appreciated that the set of chromosomes (the karyotype) was the carrier of the genes. Levitsky seems to have been t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Microarrays
A DNA microarray (also commonly known as DNA chip or biochip) is a collection of microscopic DNA spots attached to a solid surface. Scientists use DNA microarrays to measure the expression levels of large numbers of genes simultaneously or to genotype multiple regions of a genome. Each DNA spot contains picomoles (10−12 moles) of a specific DNA sequence, known as '' probes'' (or ''reporters'' or '' oligos''). These can be a short section of a gene or other DNA element that are used to hybridize a cDNA or cRNA (also called anti-sense RNA) sample (called ''target'') under high-stringency conditions. Probe-target hybridization is usually detected and quantified by detection of fluorophore-, silver-, or chemiluminescence-labeled targets to determine relative abundance of nucleic acid sequences in the target. The original nucleic acid arrays were macro arrays approximately 9 cm × 12 cm and the first computerized image based analysis was published in 1981. It was inv ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
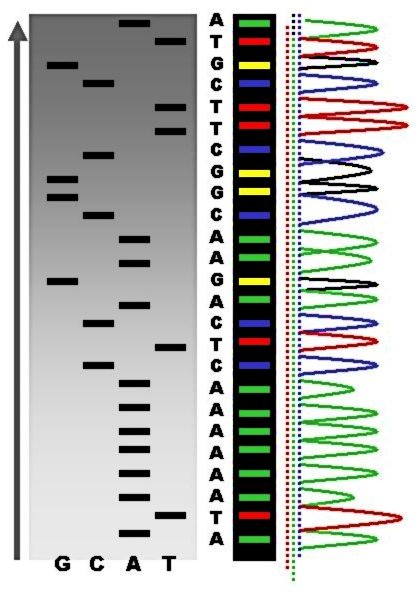
DNA Sequencing
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery. Knowledge of DNA sequences has become indispensable for basic biological research, DNA Genographic Projects and in numerous applied fields such as medical diagnosis, biotechnology, forensic biology, virology and biological systematics. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers, characterize antibody repertoire, and can be used to guide patient treatment. Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged. The rapid speed of sequencing attained with modern D ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RT-PCR
Reverse transcription polymerase chain reaction (RT-PCR) is a laboratory technique combining reverse transcription of RNA into DNA (in this context called complementary DNA or cDNA) and amplification of specific DNA targets using polymerase chain reaction (PCR). It is primarily used to measure the amount of a specific RNA. This is achieved by monitoring the amplification reaction using fluorescence, a technique called real-time PCR or quantitative PCR (qPCR). Combined RT-PCR and qPCR are routinely used for analysis of gene expression and quantification of viral RNA in research and clinical settings. The close association between RT-PCR and qPCR has led to metonymic use of the term qPCR to mean RT-PCR. Such use may be confusing, as RT-PCR can be used without qPCR, for example to enable molecular cloning, sequencing or simple detection of RNA. Conversely, qPCR may be used without RT-PCR, for example to quantify the copy number of a specific piece of DNA. Nomenclature The combine ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polymerase Chain Reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) to a large enough amount to study in detail. PCR was invented in 1983 by the American biochemist Kary Mullis at Cetus Corporation; Mullis and biochemist Michael Smith (chemist), Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993. PCR is fundamental to many of the procedures used in genetic testing and research, including analysis of Ancient DNA, ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory research for a broad variety of applications ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Law Of Independent Assortment
Mendelian inheritance (also known as Mendelism) is a type of biological inheritance following the principles originally proposed by Gregor Mendel in 1865 and 1866, re-discovered in 1900 by Hugo de Vries and Carl Correns, and later popularized by William Bateson. These principles were initially controversial. When Mendel's theories were integrated with the Boveri–Sutton chromosome theory of inheritance by Thomas Hunt Morgan in 1915, they became the core of classical genetics. Ronald Fisher combined these ideas with the theory of natural selection in his 1930 book ''The Genetical Theory of Natural Selection'', putting evolution onto a mathematical footing and forming the basis for population genetics within the modern evolutionary synthesis. History The principles of Mendelian inheritance were named for and first derived by Gregor Johann Mendel, a nineteenth-century Moravian monk who formulated his ideas after conducting simple hybridisation experiments with pea plants (''P ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Law Of Segregation
Mendelian inheritance (also known as Mendelism) is a type of biological inheritance following the principles originally proposed by Gregor Mendel in 1865 and 1866, re-discovered in 1900 by Hugo de Vries and Carl Correns, and later popularized by William Bateson. These principles were initially controversial. When Mendel's theories were integrated with the Boveri–Sutton chromosome theory of inheritance by Thomas Hunt Morgan in 1915, they became the core of classical genetics. Ronald Fisher combined these ideas with the theory of natural selection in his 1930 book ''The Genetical Theory of Natural Selection'', putting evolution onto a mathematical footing and forming the basis for population genetics within the modern evolutionary synthesis. History The principles of Mendelian inheritance were named for and first derived by Gregor Johann Mendel, a nineteenth-century Moravian monk who formulated his ideas after conducting simple hybridisation experiments with pea plants (''P ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |