|
Fuzzy Complex
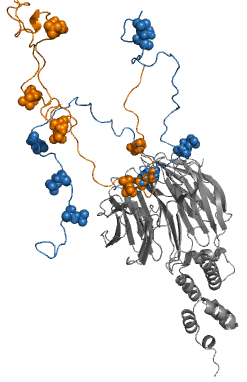
Fuzzy complexes are protein complexes, where structural ambiguity or multiplicity exists and is required for biological function.Fuxreiter, M. & Tompa, P. (2011) Fuzziness: Structural Disorder in Protein Complexes Austin, New York. Alteration, truncation or removal of conformationally ambiguous regions impacts the activity of the corresponding complex. Fuzzy complexes are generally formed by intrinsically disordered proteins. Structural multiplicity usually underlies functional multiplicity of protein complexes following a fuzzy logic. Distinct binding modes of the nucleosome are also regarded as a special case of fuzziness. Historical background For almost 50 years molecular biology was based on two dogmas: (i) equating biological function of the protein with a unique three-dimensional structure and (ii) assuming exquisite specificity in protein complexes. Specificity/selectivity is ensured by unambiguous set of interactions formed between the protein and its ligand (another ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sic1
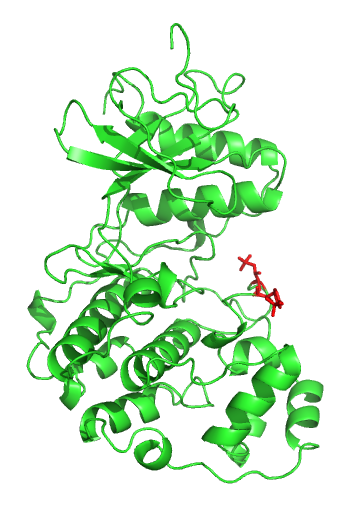
Sic1, a protein, is a stoichiometric inhibitor of Cdk1-Clb (B-type cyclins) complexes in the budding yeast ''Saccharomyces cerevisiae''. Because B-type cyclin-Cdk1 complexes are the drivers of S-phase initiation, Sic1 prevents premature S-phase entry. Multisite phosphorylation of Sic1 is thought to time Sic1 ubiquitination and destruction, and by extension, the timing of S-phase entry. Cell cycle control In the G1 phase of the cell cycle, Sic1 binds tightly to the Cdc28-Clb complex and inhibits it. Low Cdc28-Clb activity leads to the disassembly of the mitotic spindle, the assembly of the prereplicative complex and initiation of bud formation in yeast. At the START point in the yeast cell cycle, the G1- cyclins Cln3, Cln1 and Cln 2 activate Cdc28. The activated complex will phosphorylate Sic1 at multiple sites which leads to its degradation by the SCF complex. When Sic1 is degraded, the Cdc28-Clb complex is no longer inhibited and the cell can enter the S/M-phase. Thus Sic1 i ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Secondary Structures
Secondary may refer to: Science and nature * Secondary emission, of particles ** Secondary electrons, electrons generated as ionization products * The secondary winding, or the electrical or electronic circuit connected to the secondary winding in a transformer * Secondary (chemistry), a term used in organic chemistry to classify various types of compounds * Secondary color, color made from mixing primary colors * Secondary mirror, second mirror element/focusing surface in a reflecting telescope * Secondary craters, often called "secondaries" * Secondary consumer, in ecology * An obsolete name for the Mesozoic in geosciences * Secondary feathers, flight feathers attached to the ulna on the wings of birds Society and culture * Secondary (football), a position in American football and Canadian football * Secondary dominant in music * Secondary education, education which typically takes place after six years of primary education ** Secondary school, the type of school at th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Wnt Signaling Pathway
The Wnt signaling pathways are a group of signal transduction pathways which begin with proteins that pass signals into a cell through cell surface receptors. The name Wnt is a portmanteau created from the names Wingless and Int-1. Wnt signaling pathways use either nearby cell-cell communication (paracrine) or same-cell communication (autocrine). They are highly evolutionarily conserved in animals, which means they are similar across animal species from fruit flies to humans. Three Wnt signaling pathways have been characterized: the canonical Wnt pathway, the noncanonical planar cell polarity pathway, and the noncanonical Wnt/calcium pathway. All three pathways are activated by the binding of a Wnt-protein ligand to a Frizzled family receptor, which passes the biological signal to the Dishevelled protein inside the cell. The canonical Wnt pathway leads to regulation of gene transcription, and is thought to be negatively regulated in part by the SPATS1 gene. The noncanonical plana ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transforming Growth Factor Beta
Transforming growth factor beta (TGF-β) is a multifunctional cytokine belonging to the transforming growth factor superfamily that includes three different mammalian isoforms (TGF-β 1 to 3, HGNC symbols TGFB1, TGFB2, TGFB3) and many other signaling proteins. TGFB proteins are produced by all white blood cell lineages. Activated TGF-β complexes with other factors to form a serine/threonine kinase complex that binds to TGF-β receptors. TGF-β receptors are composed of both type 1 and type 2 receptor subunits. After the binding of TGF-β, the type 2 receptor kinase phosphorylates and activates the type 1 receptor kinase that activates a signaling cascade. This leads to the activation of different downstream substrates and regulatory proteins, inducing transcription of different target genes that function in differentiation, chemotaxis, proliferation, and activation of many immune cells. TGF-β is secreted by many cell types, including macrophages, in a latent form in which it ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mitogen-activated Protein Kinase
A mitogen-activated protein kinase (MAPK or MAP kinase) is a type of protein kinase that is specific to the amino acids serine and threonine (i.e., a serine/threonine-specific protein kinase). MAPKs are involved in directing cellular responses to a diverse array of stimuli, such as mitogens, osmotic stress, heat shock and proinflammatory cytokines. They regulate cell functions including proliferation, gene expression, differentiation, mitosis, cell survival, and apoptosis. MAP kinases are found in eukaryotes only, but they are fairly diverse and encountered in all animals, fungi and plants, and even in an array of unicellular eukaryotes. MAPKs belong to the CMGC (CDK/MAPK/GSK3/CLK) kinase group. The closest relatives of MAPKs are the cyclin-dependent kinases (CDKs). Discovery The first mitogen-activated protein kinase to be discovered was ERK1 (MAPK3) in mammals. Since ERK1 and its close relative ERK2 (MAPK1) are both involved in growth factor signaling, the family was term ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Epidermal Growth Factor
Epidermal growth factor (EGF) is a protein that stimulates cell growth and differentiation by binding to its receptor, EGFR. Human EGF is 6-k Da and has 53 amino acid residues and three intramolecular disulfide bonds. EGF was originally described as a secreted peptide found in the submaxillary glands of mice and in human urine. EGF has since been found in many human tissues, including platelets, submandibular gland (submaxillary gland), and parotid gland. Initially, human EGF was known as urogastrone. Structure In humans, EGF has 53 amino acids (sequence NSDSECPLSHDGYCLHDGVCMYIEALDKYACNCVVGYIGERCQYRDLKWWELR), with a molecular mass of around 6 kDa. It contains three disulfide bridges (Cys6-Cys20, Cys14-Cys31, Cys33-Cys42). Function EGF, via binding to its cognate receptor, results in cellular proliferation, differentiation, and survival. Salivary EGF, which seems to be regulated by dietary inorganic iodine, also plays an important physiological role in the maintenance of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tissue (biology)
In biology, tissue is a biological organizational level between cells and a complete organ. A tissue is an ensemble of similar cells and their extracellular matrix from the same origin that together carry out a specific function. Organs are then formed by the functional grouping together of multiple tissues. The English word "tissue" derives from the French word "tissu", the past participle of the verb tisser, "to weave". The study of tissues is known as histology or, in connection with disease, as histopathology. Xavier Bichat is considered as the "Father of Histology". Plant histology is studied in both plant anatomy and physiology. The classical tools for studying tissues are the paraffin block in which tissue is embedded and then sectioned, the histological stain, and the optical microscope. Developments in electron microscopy, immunofluorescence, and the use of frozen tissue-sections have enhanced the detail that can be observed in tissues. With these tools, the c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alternative Splicing
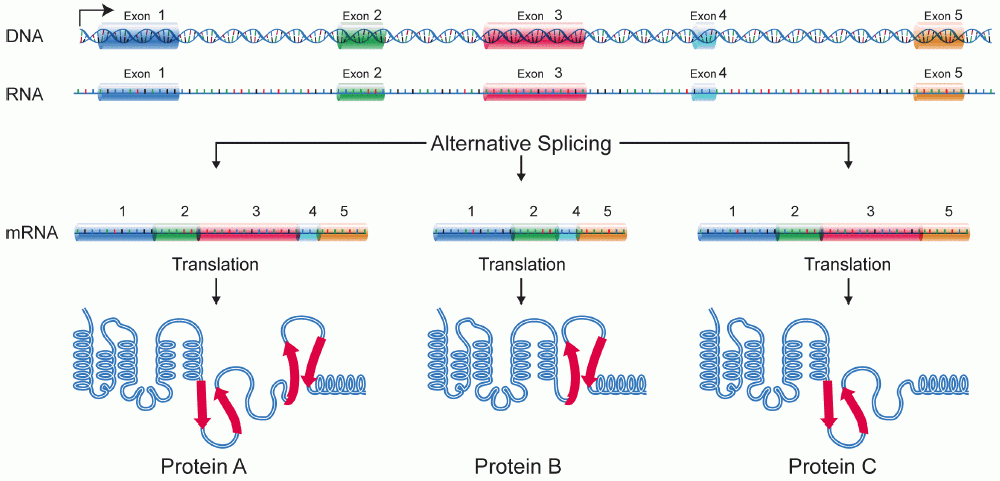
Alternative splicing, or alternative RNA splicing, or differential splicing, is an alternative splicing process during gene expression that allows a single gene to code for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. This means the exons are joined in different combinations, leading to different (alternative) mRNA strands. Consequently, the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions (see Figure). Biologically relevant alternative splicing occurs as a normal phenomenon in eukaryotes, where it increases the number of proteins that can be encoded by the genome. In humans, it is widely believed that ~95% of multi-exonic genes are alternatively spliced to produce functional alternative products from the same gene but many scientists believe that most o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Binding Affinity
In biochemistry and pharmacology, a ligand is a substance that forms a complex with a biomolecule to serve a biological purpose. The etymology stems from ''ligare'', which means 'to bind'. In protein-ligand binding, the ligand is usually a molecule which produces a signal by binding to a site on a target protein. The binding typically results in a change of conformational isomerism (conformation) of the target protein. In DNA-ligand binding studies, the ligand can be a small molecule, ion, or protein which binds to the DNA double helix. The relationship between ligand and binding partner is a function of charge, hydrophobicity, and molecular structure. Binding occurs by intermolecular forces, such as ionic bonds, hydrogen bonds and Van der Waals forces. The association or docking is actually reversible through dissociation. Measurably irreversible covalent bonding between a ligand and target molecule is atypical in biological systems. In contrast to the definition of ligan ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Posttranslational Modifications
Post-translational modification (PTM) is the covalent and generally enzymatic modification of proteins following protein biosynthesis. This process occurs in the endoplasmic reticulum and the golgi apparatus. Proteins are synthesized by ribosomes translating mRNA into polypeptide chains, which may then undergo PTM to form the mature protein product. PTMs are important components in cell signaling, as for example when prohormones are converted to hormones. Post-translational modifications can occur on the amino acid side chains or at the protein's C- or N- termini. They can extend the chemical repertoire of the 20 standard amino acids by modifying an existing functional group or introducing a new one such as phosphate. Phosphorylation is a highly effective mechanism for regulating the activity of enzymes and is the most common post-translational modification. Many eukaryotic and prokaryotic proteins also have carbohydrate molecules attached to them in a process called glycosylati ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Multiprotein Complex
A protein complex or multiprotein complex is a group of two or more associated polypeptide chains. Protein complexes are distinct from multienzyme complexes, in which multiple catalytic domains are found in a single polypeptide chain. Protein complexes are a form of quaternary structure. Proteins in a protein complex are linked by non-covalent protein–protein interactions. These complexes are a cornerstone of many (if not most) biological processes. The cell is seen to be composed of modular supramolecular complexes, each of which performs an independent, discrete biological function. Through proximity, the speed and selectivity of binding interactions between enzymatic complex and substrates can be vastly improved, leading to higher cellular efficiency. Many of the techniques used to enter cells and isolate proteins are inherently disruptive to such large complexes, complicating the task of determining the components of a complex. Examples of protein complexes include the p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Histone
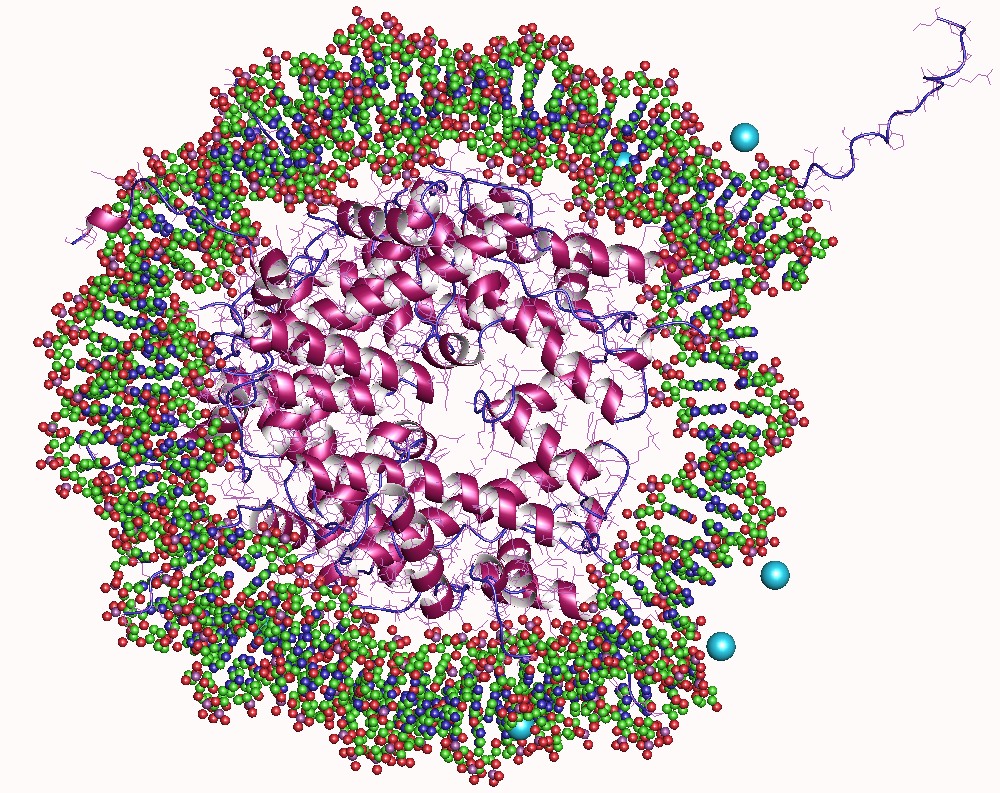
In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei. They act as spools around which DNA winds to create structural units called nucleosomes. Nucleosomes in turn are wrapped into 30-nanometer fibers that form tightly packed chromatin. Histones prevent DNA from becoming tangled and protect it from DNA damage. In addition, histones play important roles in gene regulation and DNA replication. Without histones, unwound DNA in chromosomes would be very long. For example, each human cell has about 1.8 meters of DNA if completely stretched out; however, when wound about histones, this length is reduced to about 90 micrometers (0.09 mm) of 30 nm diameter chromatin fibers. There are five families of histones which are designated H1/H5 (linker histones), H2, H3, and H4 (core histones). The nucleosome core is formed of two H2A-H2B dimers and a H3-H4 tetramer. The tight wrapping of DNA around histones ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |