|
Four-gamete Test
In population genetics, the four-gamete test is a method for detecting historical recombination events. Description Given a set of four or more sampled haploid chromosomes, the four-gamete test (FGT) detects recombination events by locating pairs of segregating sites that cannot have arisen without either recombination or a repeat mutation. Under the infinite-sites assumption (i.e. repeat mutations have zero probability), the probability of a repeat mutation is zero, and hence a recombination event is inferred. For example, if the data being studied consists of bi-allelic single-nucleotide polymorphism data, then the following configuration could be generated ''without'' recombination. However, the following configuration cannot be generated without recombination. {, class="wikitable" ! Chromosome !! Site 1 !! Site 2 , - , 1 , , 0 , , 0 , - , 2 , , 1 , , 0 , - , 3 , , 0 , , 1 , - , 4 , , 1 , , 1 The FGT detects a recombination event if the above configurati ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Population Genetics
Population genetics is a subfield of genetics that deals with genetic differences within and between populations, and is a part of evolutionary biology. Studies in this branch of biology examine such phenomena as adaptation, speciation, and population structure. Population genetics was a vital ingredient in the emergence of the modern evolutionary synthesis. Its primary founders were Sewall Wright, J. B. S. Haldane and Ronald Fisher, who also laid the foundations for the related discipline of quantitative genetics. Traditionally a highly mathematical discipline, modern population genetics encompasses theoretical, laboratory, and field work. Population genetic models are used both for statistical inference from DNA sequence data and for proof/disproof of concept. What sets population genetics apart from newer, more phenotypic approaches to modelling evolution, such as evolutionary game theory and adaptive dynamics, is its emphasis on such genetic phenomena as dominance, epi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genetic Recombination
Genetic recombination (also known as genetic reshuffling) is the exchange of genetic material between different organisms which leads to production of offspring with combinations of traits that differ from those found in either parent. In eukaryotes, genetic recombination during meiosis can lead to a novel set of genetic information that can be further passed on from parents to offspring. Most recombination occurs naturally and can be classified into two types: (1) ''interchromosomal'' recombination, occurring through independent assortment of alleles whose loci are on different but homologous chromosomes (random orientation of pairs of homologous chromosomes in meiosis I); & (2) ''intrachromosomal'' recombination, occurring through crossing over. During meiosis in eukaryotes, genetic recombination involves the pairing of homologous chromosomes. This may be followed by information transfer between the chromosomes. The information transfer may occur without physical exchange (a se ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Haploid
Ploidy () is the number of complete sets of chromosomes in a cell, and hence the number of possible alleles for autosomal and pseudoautosomal genes. Sets of chromosomes refer to the number of maternal and paternal chromosome copies, respectively, in each homologous chromosome pair, which chromosomes naturally exist as. Somatic cells, tissues, and individual organisms can be described according to the number of sets of chromosomes present (the "ploidy level"): monoploid (1 set), diploid (2 sets), triploid (3 sets), tetraploid (4 sets), pentaploid (5 sets), hexaploid (6 sets), heptaploid or septaploid (7 sets), etc. The generic term polyploid is often used to describe cells with three or more chromosome sets. Virtually all sexually reproducing organisms are made up of somatic cells that are diploid or greater, but ploidy level may vary widely between different organisms, between different tissues within the same organism, and at different stages in an organism's life cycle. Half ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chromosomes
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins are the histones. These proteins, aided by chaperone proteins, bind to and condense the DNA molecule to maintain its integrity. These chromosomes display a complex three-dimensional structure, which plays a significant role in transcriptional regulation. Chromosomes are normally visible under a light microscope only during the metaphase of cell division (where all chromosomes are aligned in the center of the cell in their condensed form). Before this happens, each chromosome is duplicated (S phase), and both copies are joined by a centromere, resulting either in an X-shaped structure (pictured above), if the centromere is located equatorially, or a two-arm structure, if the centromere is located distally. The joined copies are now called sis ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Segregating Site
Segregating sites are positions which show differences ( polymorphisms) between related genes in a sequence alignment (are not conserved). Segregating sites include conservative, semi-conservative and non-conservative mutations. The proportion of segregating sites within a gene is an important statistic in population genetics since it can be used to estimate mutation rate assuming no selection. For example it is used to calculate the Tajima's D neutral evolution statistic. See also * Conserved sequence * Ultra-conserved element * Sequence alignment * Sequence alignment software * ClustalW Clustal is a series of widely used computer programs used in bioinformatics for multiple sequence alignment. There have been many versions of Clustal over the development of the algorithm that are listed below. The analysis of each tool and its ... References Population genetics {{Genetics-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA or viral replication, mitosis, or meiosis or other types of damage to DNA (such as pyrimidine dimers caused by exposure to ultraviolet radiation), which then may undergo error-prone repair (especially microhomology-mediated end joining), cause an error during other forms of repair, or cause an error during replication (translesion synthesis). Mutations may also result from insertion or deletion of segments of DNA due to mobile genetic elements. Mutations may or may not produce detectable changes in the observable characteristics (phenotype) of an organism. Mutations play a part in both normal and abnormal biological processes including: evolution, cancer, and the development of the immune system, including junctional diversity. Mutation is the ultimate source o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Single-nucleotide Polymorphism
In genetics, a single-nucleotide polymorphism (SNP ; plural SNPs ) is a germline substitution of a single nucleotide at a specific position in the genome. Although certain definitions require the substitution to be present in a sufficiently large fraction of the population (e.g. 1% or more), many publications do not apply such a frequency threshold. For example, at a specific base position in the human genome, the G nucleotide may appear in most individuals, but in a minority of individuals, the position is occupied by an A. This means that there is a SNP at this specific position, and the two possible nucleotide variations – G or A – are said to be the alleles for this specific position. SNPs pinpoint differences in our susceptibility to a wide range of diseases, for example age-related macular degeneration (a common SNP in the CFH gene is associated with increased risk of the disease) or nonalcoholic fatty liver disease (a SNP in the PNPLA3 gene is associated with inc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
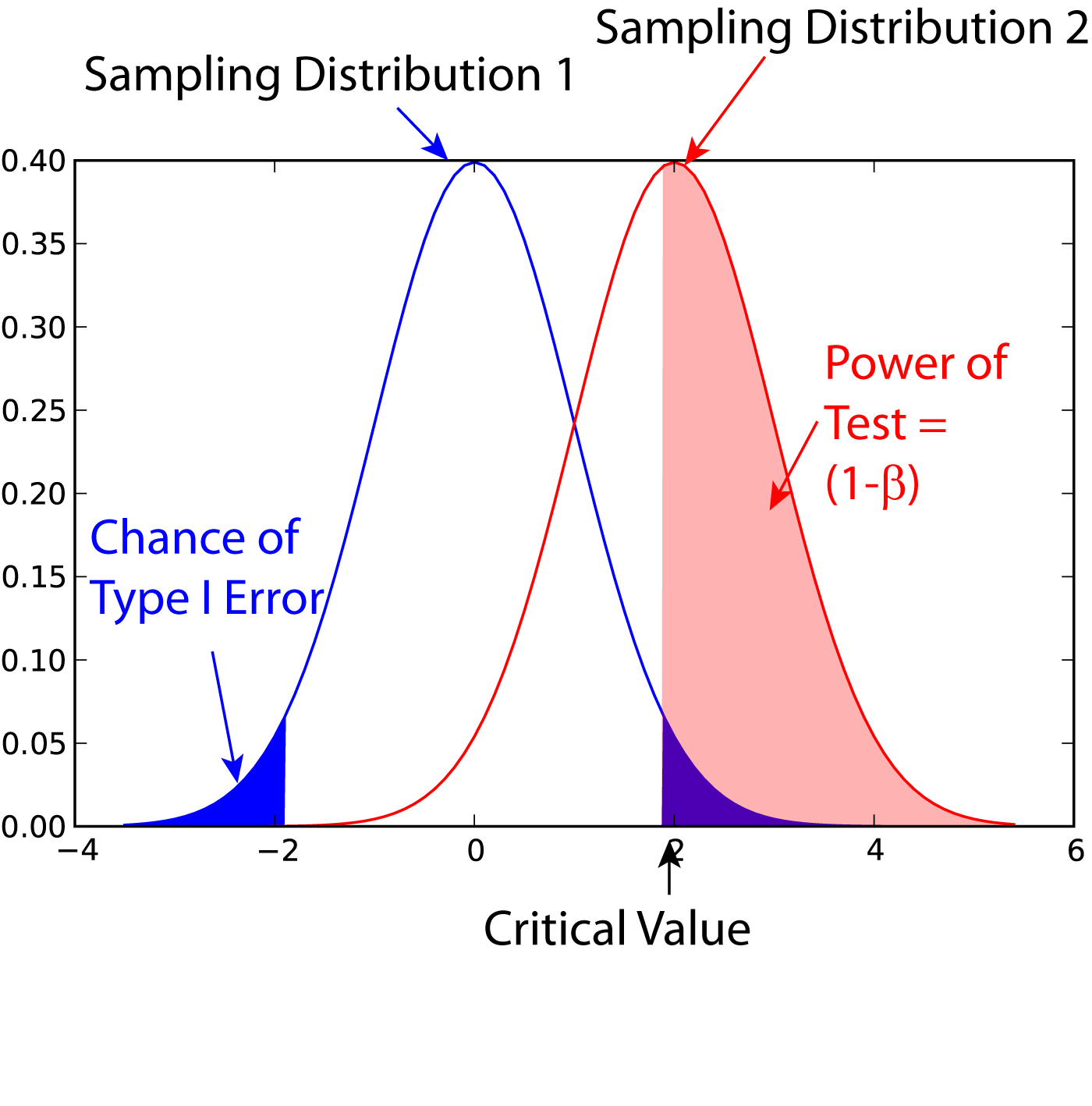
Statistical Power
In statistics, the power of a binary hypothesis test is the probability that the test correctly rejects the null hypothesis (H_0) when a specific alternative hypothesis (H_1) is true. It is commonly denoted by 1-\beta, and represents the chances of a true positive detection conditional on the actual existence of an effect to detect. Statistical power ranges from 0 to 1, and as the power of a test increases, the probability \beta of making a type II error by wrongly failing to reject the null hypothesis decreases. Notation This article uses the following notation: * ''β'' = probability of a Type II error, known as a "false negative" * 1 − ''β'' = probability of a "true positive", i.e., correctly rejecting the null hypothesis. "1 − ''β''" is also known as the power of the test. * ''α'' = probability of a Type I error, known as a "false positive" * 1 − ''α'' = probability of a "true negative", i.e., correctly not rejecting the null hypothesis Description For a ty ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
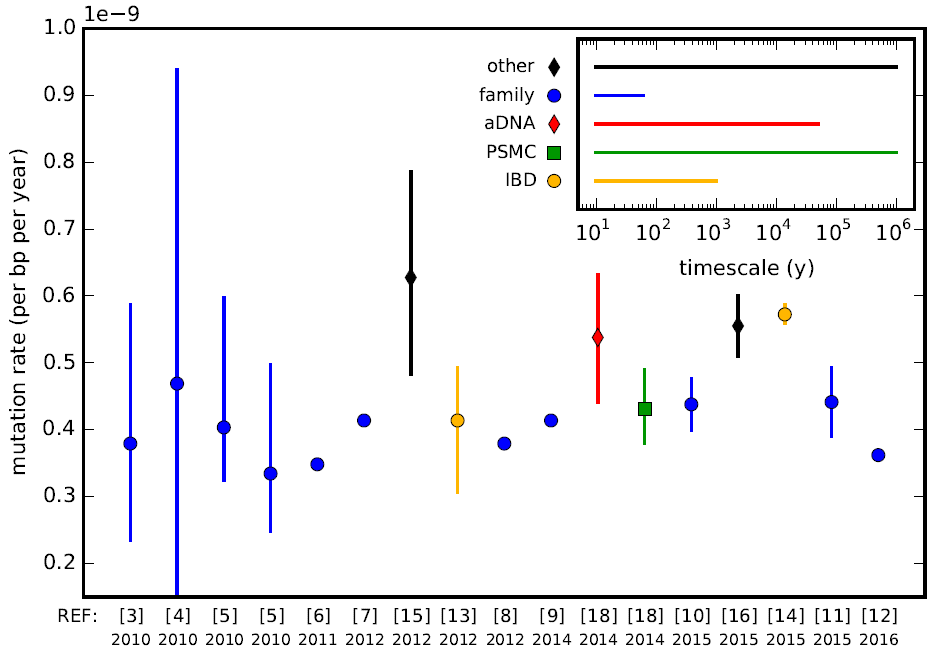
Mutation Rate
In genetics, the mutation rate is the frequency of new mutations in a single gene or organism over time. Mutation rates are not constant and are not limited to a single type of mutation; there are many different types of mutations. Mutation rates are given for specific classes of mutations. Point mutations are a class of mutations which are changes to a single base. Missense and Nonsense mutations are two subtypes of point mutations. The rate of these types of substitutions can be further subdivided into a mutation spectrum which describes the influence of the genetic context on the mutation rate. There are several natural units of time for each of these rates, with rates being characterized either as mutations per base pair per cell division, per gene per generation, or per genome per generation. The mutation rate of an organism is an evolved characteristic and is strongly influenced by the genetics of each organism, in addition to strong influence from the environment. The upper ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Coalescent Theory
Coalescent theory is a model of how alleles sampled from a population may have originated from a common ancestor. In the simplest case, coalescent theory assumes no recombination, no natural selection, and no gene flow or population structure, meaning that each variant is equally likely to have been passed from one generation to the next. The model looks backward in time, merging alleles into a single ancestral copy according to a random process in coalescence events. Under this model, the expected time between successive coalescence events increases almost exponentially back in time (with wide variance). Variance in the model comes from both the random passing of alleles from one generation to the next, and the random occurrence of mutations in these alleles. The mathematical theory of the coalescent was developed independently by several groups in the early 1980s as a natural extension of classical population genetics theory and models, but can be primarily attributed to John K ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |