|
Endopeptidases
Endopeptidase or endoproteinase are proteolytic peptidases that break peptide bonds of nonterminal amino acids (i.e. within the molecule), in contrast to exopeptidases, which break peptide bonds from end-pieces of terminal amino acids. For this reason, endopeptidases cannot break down peptides into monomers, while exopeptidases can break down proteins into monomers. A particular case of endopeptidase is the oligopeptidase, whose substrates are oligopeptides instead of proteins. They are usually very specific for certain amino acids. Examples of endopeptidases include: * Trypsin - cuts after Arg or Lys, unless followed by Pro. Very strict. Works best at pH 8. * Chymotrypsin - cuts after Phe, Trp, or Tyr, unless followed by Pro. Cuts more slowly after His, Met or Leu. Works best at pH 8. * Elastase - cuts after Ala, Gly, Ser, or Val, unless followed by Pro. * Thermolysin - cuts ''before'' Ile, Met, Phe, Trp, Tyr, or Val, unless ''preceded'' by Pro. Sometimes cuts after Ala, Asp, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Oligopeptidase
An Oligopeptidase is an enzyme that cleaves peptides but not proteins. This property is due to its structure: the active site of this enzyme is located at the end of a narrow cavity which can only be reached by peptides. History Background Proteins are essential macromolecules of living organisms. They are continuously being degraded into their constituent amino acids which can be reused in the synthesis of new proteins. Every cellular protein has its own half-life time. In humans, for instance, 50% of the liver and plasma proteins are replaced in 10 days, whereas in muscles it takes 180 days. In average, every 80 days about 50% of our proteins are totally replaced. Although the regulation of protein degradation is as important as their synthesis to keep each cell protein concentration at the optimum level, research in this area remained until the end of the 1970s. Up to this time, lysosomes, discovered in the 1950s by the Belgian cytologist Christian de Duve, were thought respon ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Glutamyl Endopeptidase I
Glutamyl endopeptidase I is a family of extracellular bacterial serine proteases. The proteases within this family have been identified in species of ''Staphylococcus'', '' Bacillus'', and ''Streptomyces'', among others. The two former are more closely related, while the ''Streptomyces''-type is treated as a separate family, glutamyl endopeptidase II. Identified proteases ''Staphylococcus'' * '' S. aureus'' glutamyl endopeptidase GluV8 : The first discovered enzyme of this family, and the most well characterized, was isolated from the ''Staphylococcus aureus'' strain V8, and hence better known as "V8 protease". Other common references to this protease are staphylococcal serine protease, and SspA from its corresponding gene. * '' S. epidermidis'' glutamyl endopeptidase GluSE : Also called ''S. epidermidis'' serine protease (Esp). * '' S. warneri'' glutamyl endopeptidase GluSW : Also referred to as PROM. ''Bacillus'' * ''B. licheniformis'' glutamyl endopeptidase GluBL * ''B. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Peptidase
A protease (also called a peptidase, proteinase, or proteolytic enzyme) is an enzyme that catalyzes (increases reaction rate or "speeds up") proteolysis, breaking down proteins into smaller polypeptides or single amino acids, and spurring the formation of new protein products. They do this by cleaving the peptide bonds within proteins by hydrolysis, a reaction where water breaks bonds. Proteases are involved in many biological functions, including digestion of ingested proteins, protein catabolism (breakdown of old proteins), and cell signaling. In the absence of functional accelerants, proteolysis would be very slow, taking hundreds of years. Proteases can be found in all forms of life and viruses. They have independently evolved multiple times, and different classes of protease can perform the same reaction by completely different catalytic mechanisms. Hierarchy of proteases Based on catalytic residue Proteases can be classified into seven broad groups: * Serine pr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Trypsin
Trypsin is an enzyme in the first section of the small intestine that starts the digestion of protein molecules by cutting these long chains of amino acids into smaller pieces. It is a serine protease from the PA clan superfamily, found in the digestive system of many vertebrates, where it hydrolyzes proteins. Trypsin is formed in the small intestine when its proenzyme form, the trypsinogen produced by the pancreas, is activated. Trypsin cuts peptide chains mainly at the carboxyl side of the amino acids lysine or arginine. It is used for numerous biotechnological processes. The process is commonly referred to as trypsin proteolysis or trypsinization, and proteins that have been digested/treated with trypsin are said to have been trypsinized. Trypsin was discovered in 1876 by Wilhelm Kühne and was named from the Ancient Greek word for rubbing since it was first isolated by rubbing the pancreas with glycerin. Function In the duodenum, trypsin catalyzes the hydro ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chymotrypsin
Chymotrypsin (, chymotrypsins A and B, alpha-chymar ophth, avazyme, chymar, chymotest, enzeon, quimar, quimotrase, alpha-chymar, alpha-chymotrypsin A, alpha-chymotrypsin) is a digestive enzyme component of pancreatic juice acting in the duodenum, where it performs proteolysis, the breakdown of proteins and polypeptides. Chymotrypsin preferentially cleaves peptide amide bonds where the side chain of the amino acid N-terminal to the scissile amide bond (the P1 position) is a large hydrophobic amino acid ( tyrosine, tryptophan, and phenylalanine). These amino acids contain an aromatic ring in their side chain that fits into a hydrophobic pocket (the S1 position) of the enzyme. It is activated in the presence of trypsin. The hydrophobic and shape complementarity between the peptide substrate P1 side chain and the enzyme S1 binding cavity accounts for the substrate specificity of this enzyme. Chymotrypsin also hydrolyzes other amide bonds in peptides at slower rates, partic ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Elastase
In molecular biology, elastase is an enzyme from the class of ''proteases (peptidases)'' that break down proteins. In particular, it is a serine protease. Forms and classification Eight human genes exist for elastase: Some bacteria (including ''Pseudomonas aeruginosa'') also produce elastase. In bacteria, elastase is considered a virulence factor. Function Elastase breaks down elastin, an elastic fibre that, together with collagen, determines the mechanical properties of connective tissue. The neutrophil form breaks down the ''Outer membrane protein A'' (OmpA) of '' E. coli'' and other Gram-negative bacteria. Elastase also has the important immunological role of breaking down Shigella virulence factors. This is accomplished through the cleavage of peptide bonds in the target proteins. The specific peptide bonds cleaved are those on the carboxyl side of small, hydrophobic amino acids such as glycine, alanine, and valine. For more on how this is accomplished, see serine p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Thermolysin
Thermolysin (, ''Bacillus thermoproteolyticus neutral proteinase'', ''thermoase'', ''thermoase Y10'', ''TLN'') is a thermostable neutral metalloproteinase enzyme produced by the Gram-positive bacteria ''Bacillus thermoproteolyticus''. It requires one zinc ion for enzyme activity and four calcium ions for structural stability. Thermolysin specifically catalyzes the hydrolysis of peptide bonds containing hydrophobic amino acids. However thermolysin is also widely used for peptide bond formation through the reverse reaction of hydrolysis. Thermolysin is the most stable member of a family of metalloproteinases produced by various ''Bacillus'' species. These enzymes are also termed 'neutral' proteinases or thermolysin -like proteinases (TLPs). Synthesis Like all bacterial extracellular proteases thermolysin is first synthesised by the bacterium as a pre-proenzyme. Thermolysin is synthesized as a pre-proenzyme consisting of a signal peptide 28 amino acids long, a pro-peptide 204 a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Pepsin
Pepsin is an endopeptidase that breaks down proteins into smaller peptides. It is produced in the gastric chief cells of the stomach lining and is one of the main digestive enzymes in the digestive systems of humans and many other animals, where it helps digest the proteins in food. Pepsin is an aspartic protease, using a catalytic aspartate in its active site. It is one of three principal endopeptidases (enzymes cutting proteins in the middle) in the human digestive system, the other two being chymotrypsin and trypsin. There are also exopeptidases which remove individual amino acids at both ends of proteins ( carboxypeptidases produced by the pancreas and aminopeptidases secreted by the small intestine). During the process of digestion, these enzymes, each of which is specialized in severing links between particular types of amino acids, collaborate to break down dietary proteins into their components, i.e., peptides and amino acids, which can be readily absorbed by t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Neprilysin
Neprilysin (), also known as membrane metallo-endopeptidase (MME), neutral endopeptidase (NEP), cluster of differentiation 10 (CD10), and common acute lymphoblastic leukemia antigen (CALLA) is an enzyme that in humans is encoded by the ''MME'' gene. Neprilysin is a zinc-dependent metalloprotease that cleaves peptides at the amino side of hydrophobic residues and inactivates several peptide hormones including glucagon, enkephalins, substance P, neurotensin, oxytocin, and bradykinin. It also degrades the amyloid beta peptide whose abnormal folding and aggregation in neural tissue has been implicated as a cause of Alzheimer's disease. Synthesized as a membrane-bound protein, the neprilysin ectodomain is released into the extracellular domain after it has been transported from the Golgi apparatus to the cell surface. Neprilysin is expressed in a wide variety of tissues and is particularly abundant in kidney. It is also a common acute lymphocytic leukemia antigen that is an importa ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Proteolysis
Proteolysis is the breakdown of proteins into smaller polypeptides or amino acids. Uncatalysed, the hydrolysis of peptide bonds is extremely slow, taking hundreds of years. Proteolysis is typically catalysed by cellular enzymes called proteases, but may also occur by intra-molecular digestion. Proteolysis in organisms serves many purposes; for example, digestive enzymes break down proteins in food to provide amino acids for the organism, while proteolytic processing of a polypeptide chain after its synthesis may be necessary for the production of an active protein. It is also important in the regulation of some physiological and cellular processes including apoptosis, as well as preventing the accumulation of unwanted or misfolded proteins in cells. Consequently, abnormality in the regulation of proteolysis can cause disease. Proteolysis can also be used as an analytical tool for studying proteins in the laboratory, and it may also be used in industry, for example in food pr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Peptide Bonds
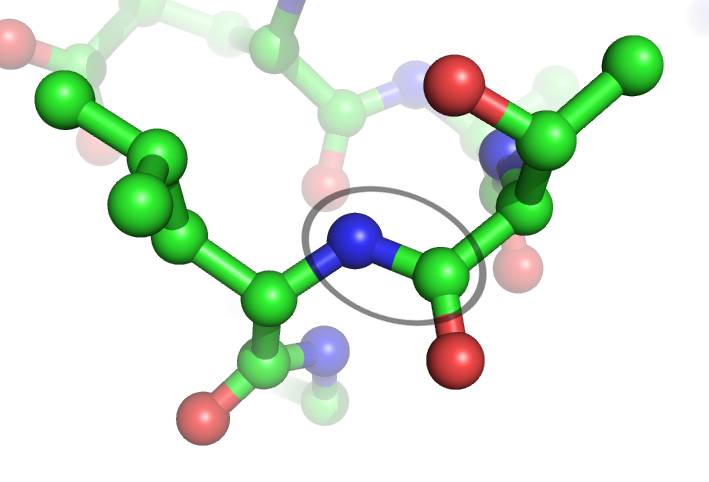
In organic chemistry, a peptide bond is an amide type of covalent chemical bond linking two consecutive alpha-amino acids from C1 (carbon number one) of one alpha-amino acid and N2 (nitrogen number two) of another, along a peptide or protein chain. It can also be called a eupeptide bond to distinguish it from an isopeptide bond, which is another type of amide bond between two amino acids. Synthesis When two amino acids form a ''dipeptide'' through a ''peptide bond'', it is a type of condensation reaction. In this kind of condensation, two amino acids approach each other, with the non-side chain (C1) carboxylic acid moiety of one coming near the non-side chain (N2) amino moiety of the other. One loses a hydrogen and oxygen from its carboxyl group (COOH) and the other loses a hydrogen from its amino group (NH2). This reaction produces a molecule of water (H2O) and two amino acids joined by a peptide bond (−CO−NH−). The two joined amino acids are called a dipeptide. The ami ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Amino Acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although hundreds of amino acids exist in nature, by far the most important are the alpha-amino acids, which comprise proteins. Only 22 alpha amino acids appear in the genetic code. Amino acids can be classified according to the locations of the core structural functional groups, as Alpha and beta carbon, alpha- , beta- , gamma- or delta- amino acids; other categories relate to Chemical polarity, polarity, ionization, and side chain group type (aliphatic, Open-chain compound, acyclic, aromatic, containing hydroxyl or sulfur, etc.). In the form of proteins, amino acid ''residues'' form the second-largest component ( water being the largest) of human muscles and other tissues. Beyond their role as residues in proteins, amino acids participate in a number of processes such as neurotransmitter transport and biosynthesis. It is thought that they played a key role in enabling li ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |