|
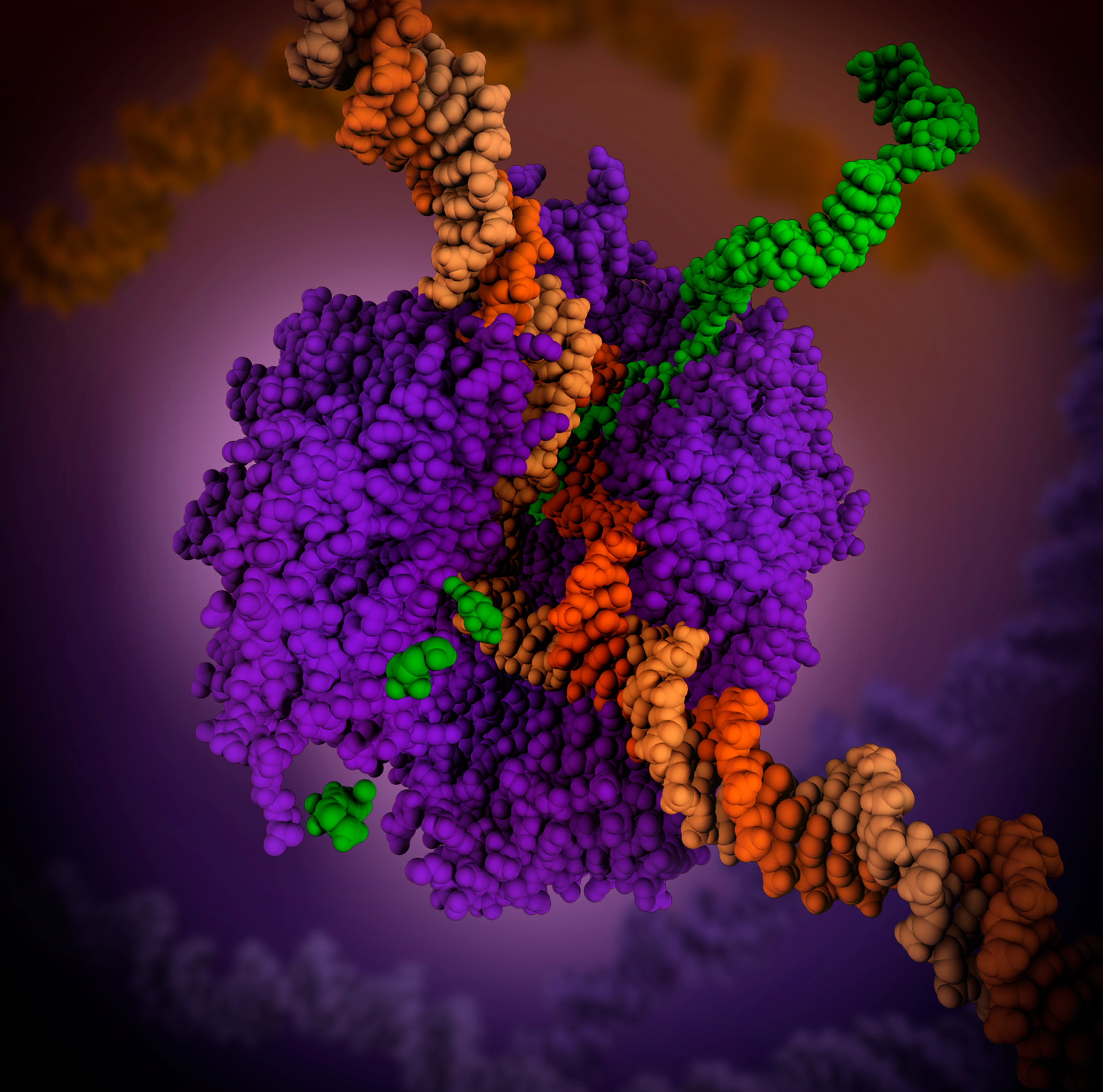
Catabolite Activator Protein
Catabolite activator protein (CAP; also known as cAMP receptor protein, CRP) is a trans-acting transcriptional activator that exists as a homodimer in solution. Each subunit of CAP is composed of a ligand-binding domain at the N-terminus (CAPN, residues 1–138) and a DNA-binding domain at the C-terminus (DBD, residues 139–209). Two cAMP (cyclic AMP) molecules bind dimeric CAP with negative cooperativity. Cyclic AMP functions as an allosteric effector by increasing CAP's affinity for DNA. CAP binds a DNA region upstream from the DNA binding site of RNA Polymerase. CAP activates transcription through protein-protein interactions with the α-subunit of RNA Polymerase. This protein-protein interaction is responsible for (i) catalyzing the formation of the RNAP-promoter closed complex; and (ii) isomerization of the RNAP-promoter complex to the open conformation. CAP's interaction with RNA polymerase causes bending of the DNA near the transcription start site, thus effectively catal ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
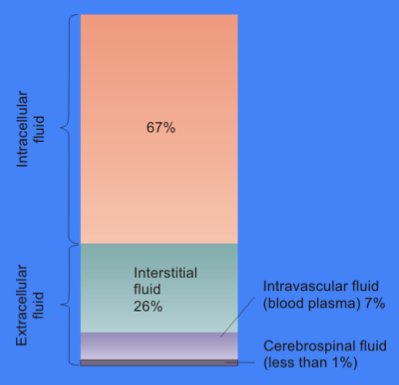
Cytosol
The cytosol, also known as cytoplasmic matrix or groundplasm, is one of the liquids found inside cells (intracellular fluid (ICF)). It is separated into compartments by membranes. For example, the mitochondrial matrix separates the mitochondrion into many compartments. In the eukaryotic cell, the cytosol is surrounded by the cell membrane and is part of the cytoplasm, which also comprises the mitochondria, plastids, and other organelles (but not their internal fluids and structures); the cell nucleus is separate. The cytosol is thus a liquid matrix around the organelles. In prokaryotes, most of the chemical reactions of metabolism take place in the cytosol, while a few take place in membranes or in the periplasmic space. In eukaryotes, while many metabolic pathways still occur in the cytosol, others take place within organelles. The cytosol is a complex mixture of substances dissolved in water. Although water forms the large majority of the cytosol, its structure and prope ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Modulon
In molecular genetics, a regulon is a group of genes that are gene regulation, regulated as a unit, generally controlled by the same regulatory gene that gene expression, expresses a protein acting as a repressor or activator (genetics), activator. This terminology is generally, although not exclusively, used in reference to prokaryotes, whose genomes are often organized into operons; the genes contained within a regulon are usually organized into more than one operon at disparate locations on the chromosome. Applied to eukaryotes, the term refers to any group of non-contiguous genes controlled by the same regulatory gene. A modulon is a set of regulons or operons that are collectively regulated in response to changes in overall conditions or stresses, but may be under the control of different or overlapping regulatory molecules. The term stimulon is sometimes used to refer to the set of genes whose expression responds to specific environmental stimuli. Examples Commonly studied ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Catabolite Repression
Carbon catabolite repression, or simply catabolite repression, is an important part of global control system of various bacteria and other microorganisms. Catabolite repression allows microorganisms to adapt quickly to a preferred (rapidly metabolizable) carbon and energy source first. This is usually achieved through inhibition of synthesis of enzymes involved in catabolism of carbon sources other than the preferred one. The catabolite repression was first shown to be initiated by glucose and therefore sometimes referred to as the glucose effect. However, the term "glucose effect" is actually a misnomer since other carbon sources are known to induce catabolite repression. ''Escherichia coli'' Catabolite repression was extensively studied in ''Escherichia coli''. ''E. coli'' grows faster on glucose than on any other carbon source. For example, if ''E. coli'' is placed on an agar plate containing only glucose and lactose, the bacteria will use glucose first and lactose second. Wh ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Lac Operon
The ''lactose'' operon (''lac'' operon) is an operon required for the transport and metabolism of lactose in ''E. coli'' and many other enteric bacteria. Although glucose is the preferred carbon source for most bacteria, the ''lac'' operon allows for the effective digestion of lactose when glucose is not available through the activity of beta-galactosidase. Gene regulation of the ''lac'' operon was the first genetic regulatory mechanism to be understood clearly, so it has become a foremost example of prokaryotic gene regulation. It is often discussed in introductory molecular and cellular biology classes for this reason. This lactose metabolism system was used by François Jacob and Jacques Monod to determine how a biological cell knows which enzyme to synthesize. Their work on the ''lac'' operon won them the Nobel Prize in Physiology in 1965. Bacterial operons are polycistronic transcripts that are able to produce multiple proteins from one mRNA transcript. In this case, when l ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Catabolism
Catabolism () is the set of metabolic pathways that breaks down molecules into smaller units that are either oxidized to release energy or used in other anabolic reactions. Catabolism breaks down large molecules (such as polysaccharides, lipids, nucleic acids, and proteins) into smaller units (such as monosaccharides, fatty acids, nucleotides, and amino acids, respectively). Catabolism is the breaking-down aspect of metabolism, whereas anabolism is the building-up aspect. Cells use the monomers released from breaking down polymers to either construct new polymer molecules or degrade the monomers further to simple waste products, releasing energy. Cellular wastes include lactic acid, acetic acid, carbon dioxide, ammonia, and urea. The formation of these wastes is usually an oxidation process involving a release of chemical free energy, some of which is lost as heat, but the rest of which is used to drive the synthesis of adenosine triphosphate (ATP). This molecule acts as a way f ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transcription (genetics)
Transcription is the process of copying a segment of DNA into RNA. The segments of DNA transcribed into RNA molecules that can encode proteins are said to produce messenger RNA (mRNA). Other segments of DNA are copied into RNA molecules called non-coding RNAs (ncRNAs). mRNA comprises only 1–3% of total RNA samples. Less than 2% of the human genome can be transcribed into mRNA ( Human genome#Coding vs. noncoding DNA), while at least 80% of mammalian genomic DNA can be actively transcribed (in one or more types of cells), with the majority of this 80% considered to be ncRNA. Both DNA and RNA are nucleic acids, which use base pairs of nucleotides as a complementary language. During transcription, a DNA sequence is read by an RNA polymerase, which produces a complementary, antiparallel RNA strand called a primary transcript. Transcription proceeds in the following general steps: # RNA polymerase, together with one or more general transcription factors, binds to promoter DNA ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA Polymerase
In molecular biology, RNA polymerase (abbreviated RNAP or RNApol), or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme that synthesizes RNA from a DNA template. Using the enzyme helicase, RNAP locally opens the double-stranded DNA so that one strand of the exposed nucleotides can be used as a template for the synthesis of RNA, a process called transcription. A transcription factor and its associated transcription mediator complex must be attached to a DNA binding site called a promoter region before RNAP can initiate the DNA unwinding at that position. RNAP not only initiates RNA transcription, it also guides the nucleotides into position, facilitates attachment and elongation, has intrinsic proofreading and replacement capabilities, and termination recognition capability. In eukaryotes, RNAP can build chains as long as 2.4 million nucleotides. RNAP produces RNA that, functionally, is either for protein coding, i.e. messenger RNA (mRNA); or n ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Major Groove
Major (commandant in certain jurisdictions) is a military rank of commissioned officer status, with corresponding ranks existing in many military forces throughout the world. When used unhyphenated and in conjunction with no other indicators, major is one rank above captain, and one rank below lieutenant colonel. It is considered the most junior of the field officer ranks. Background Majors are typically assigned as specialised executive or operations officers for battalion-sized units of 300 to 1,200 soldiers while in some nations, like Germany, majors are often in command of a company. When used in hyphenated or combined fashion, the term can also imply seniority at other levels of rank, including ''general-major'' or ''major general'', denoting a low-level general officer, and ''sergeant major'', denoting the most senior non-commissioned officer (NCO) of a military unit. The term ''major'' can also be used with a hyphen to denote the leader of a military band such as i ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Helix-turn-helix
Helix-turn-helix is a DNA-binding protein (DBP). The helix-turn-helix (HTH) is a major structural motif capable of binding DNA. Each monomer incorporates two α helices, joined by a short strand of amino acids, that bind to the major groove of DNA. The HTH motif occurs in many proteins that regulate gene expression. It should not be confused with the helix–loop–helix motif. Discovery The discovery of the helix-turn-helix motif was based on similarities between several genes encoding transcription regulatory proteins from bacteriophage lambda and ''Escherichia coli'': Cro, CAP, and λ repressor, which were found to share a common 20–25 amino acid sequence that facilitates DNA recognition. Function The helix-turn-helix motif is a DNA-binding motif. The recognition and binding to DNA by helix-turn-helix proteins is done by the two α helices, one occupying the N-terminal end of the motif, the other at the C-terminus. In most cases, such as in the Cro repressor, the second ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Affinity (pharmacology)
In biochemistry and pharmacology, a ligand is a substance that forms a complex with a biomolecule to serve a biological purpose. The etymology stems from ''ligare'', which means 'to bind'. In protein-ligand binding, the ligand is usually a molecule which produces a signal by binding to a site on a target protein. The binding typically results in a change of conformational isomerism (conformation) of the target protein. In DNA-ligand binding studies, the ligand can be a small molecule, ion, or protein which binds to the DNA double helix. The relationship between ligand and binding partner is a function of charge, hydrophobicity, and molecular structure. Binding occurs by intermolecular forces, such as ionic bonds, hydrogen bonds and Van der Waals forces. The association or docking is actually reversible through dissociation. Measurably irreversible covalent bonding between a ligand and target molecule is atypical in biological systems. In contrast to the definition of liga ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
CAMP Receptor Protein
cAMP receptor protein (CRP; also known as catabolite activator protein, CAP) is a regulatory protein in bacteria. CRP protein binds cAMP, which causes a conformational change that allows CRP to bind tightly to a specific DNA site in the promoters of the genes it controls. CRP then activates transcription through direct protein–protein interactions with RNA polymerase. The genes regulated by CRP are mostly involved in energy metabolism, such as galactose, citrate, or the PEP group translocation system. In ''Escherichia coli'', cyclic AMP receptor protein (CRP) can regulate the transcription of more than 100 genes. The signal to activate CRP is the binding of cyclic AMP. Binding of cAMP to CRP leads to a long-distance signal transduction from the N-terminal cAMP-binding domain to the C-terminal domain of the protein, which is responsible for interaction with specific sequences of DNA. At "Class I" CRP-dependent promoters, CRP binds to a DNA site located upstream of core promo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |