|
UBE2M
NEDD8-conjugating enzyme Ubc12 is a protein that in humans is encoded by the ''UBE2M'' gene. The modification of proteins with ubiquitin is an important cellular mechanism for targeting abnormal or short-lived proteins for degradation. Ubiquitination involves at least three classes of enzymes: ubiquitin-activating enzymes, or E1s, ubiquitin-conjugating enzymes, or E2s, and ubiquitin-protein ligases, or E3s. This gene encodes a member of the E2 ubiquitin-conjugating enzyme family. The encoded protein is linked with a ubiquitin-like protein, NEDD8, which can be conjugated to cellular proteins, such as Cdc53/culin. Interactions UBE2M has been shown to interact with NEDD8, PRKAR1A cAMP-dependent protein kinase type I-alpha regulatory subunit is an enzyme that in humans is encoded by the ''PRKAR1A'' gene. Function cAMP is a signaling molecule important for a variety of cellular functions. cAMP exerts its effects by activ ... and UBA3. References Further reading * * * * ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
NEDD8
NEDD8 is a protein that in humans is encoded by the ''NEDD8'' gene. (in ''saccharomyces cerevisiae'' this protein is known as Rub1) This ubiquitin-like (UBL) protein becomes covalently conjugated to a limited number of cellular proteins, in a process called NEDDylation similar to ubiquitination. Human NEDD8 shares 60% amino acid sequence identity to ubiquitin. The primary known substrates of NEDD8 modification are the cullin subunits of cullin-based E3 ubiquitin ligases, which are active only when NEDDylated. Their NEDDylation is critical for the recruitment of E2 to the ligase complex, thus facilitating ubiquitin conjugation. NEDD8 modification has therefore been implicated in cell cycle progression and cytoskeletal regulation. Activation and conjugation As with ubiquitin and SUMO, NEDD8 is conjugated to cellular proteins after its C-terminal tail is processed. The NEDD8 activating E1 enzyme is a heterodimer composed of APPBP1 and UBA3 subunits. The APPBP1/UBA3 enzyme has homolog ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin-conjugating Enzyme
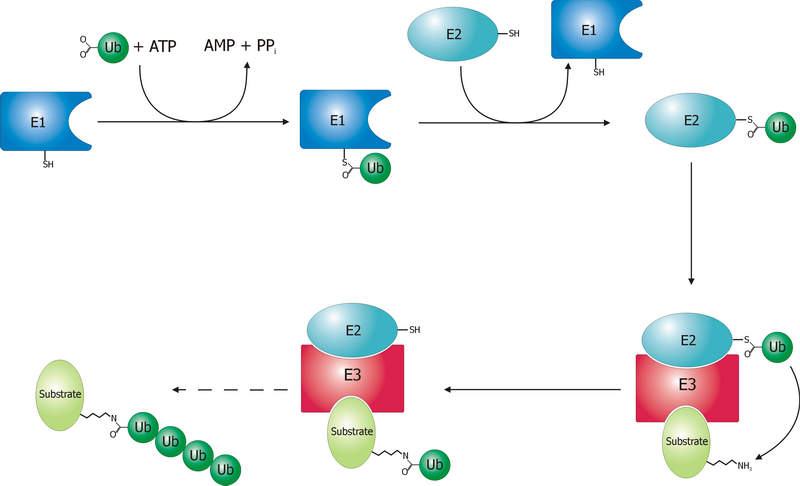
Ubiquitin-conjugating enzymes, also known as E2 enzymes and more rarely as ''ubiquitin-carrier enzymes'', perform the second step in the ubiquitination reaction that targets a protein for degradation via the proteasome. The ubiquitination process covalently attaches ubiquitin, a short protein of 76 amino acids, to a lysine residue on the target protein. Once a protein has been tagged with one ubiquitin molecule, additional rounds of ubiquitination form a polyubiquitin chain that is recognized by the proteasome's 19S regulatory particle, triggering the ATP-dependent unfolding of the target protein that allows passage into the proteasome's 20S core particle, where proteases degrade the target into short peptide fragments for recycling by the cell. Relationships A ubiquitin-activating enzyme, or E1, first activates the ubiquitin by covalently attaching the molecule to its active site cysteine residue. The activated ubiquitin is then transferred to an E2 cysteine. Once conjugated to ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
PRKAR1A
cAMP-dependent protein kinase type I-alpha regulatory subunit is an enzyme that in humans is encoded by the ''PRKAR1A'' gene. Function cAMP is a signaling molecule important for a variety of cellular functions. cAMP exerts its effects by activating the cAMP-dependent protein kinase A ( PKA), which transduces the signal through phosphorylation of different target proteins. The inactive holoenzyme of PKA is a tetramer composed of two regulatory and two catalytic subunits. cAMP causes the dissociation of the inactive holoenzyme into a dimer of regulatory subunits bound to four cAMP and two free monomeric catalytic subunits. Four different regulatory subunits and three catalytic subunits of PKA have been identified in humans. The protein encoded by this gene is one of the regulatory subunits. This protein was found to be a tissue-specific extinguisher that down-regulates the expression of seven liver genes in hepatoma x fibroblast hybrids Three alternatively spliced transcript varia ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residue ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene
In biology, the word gene (from , ; "...Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a basic unit of heredity and the molecular gene is a sequence of nucleotides in DNA that is transcribed to produce a functional RNA. There are two types of molecular genes: protein-coding genes and noncoding genes. During gene expression, the DNA is first copied into RNA. The RNA can be directly functional or be the intermediate template for a protein that performs a function. The transmission of genes to an organism's offspring is the basis of the inheritance of phenotypic traits. These genes make up different DNA sequences called genotypes. Genotypes along with environmental and developmental factors determine what the phenotypes will be. Most biological traits are under the influence of polygenes (many different genes) as well as gen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin
Ubiquitin is a small (8.6 kDa) regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ''ubiquitously''. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 1980s. Four genes in the human genome code for ubiquitin: UBB, UBC, UBA52 and RPS27A. The addition of ubiquitin to a substrate protein is called ubiquitylation (or, alternatively, ubiquitination or ubiquitinylation). Ubiquitylation affects proteins in many ways: it can mark them for degradation via the proteasome, alter their cellular location, affect their activity, and promote or prevent protein interactions. Ubiquitylation involves three main steps: activation, conjugation, and ligation, performed by ubiquitin-activating enzymes (E1s), ubiquitin-conjugating enzymes (E2s), and ubiquitin ligases (E3s), respectively. The result of this sequential cascade is to bind ubiquitin to lysine residues on the protein substrate via an isopeptide bond, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Degradation
Proteolysis is the breakdown of proteins into smaller polypeptides or amino acids. Uncatalysed, the hydrolysis of peptide bonds is extremely slow, taking hundreds of years. Proteolysis is typically catalysed by cellular enzymes called proteases, but may also occur by intra-molecular digestion. Proteolysis in organisms serves many purposes; for example, digestive enzymes break down proteins in food to provide amino acids for the organism, while proteolytic processing of a polypeptide chain after its synthesis may be necessary for the production of an active protein. It is also important in the regulation of some physiological and cellular processes including apoptosis, as well as preventing the accumulation of unwanted or misfolded proteins in cells. Consequently, abnormality in the regulation of proteolysis can cause disease. Proteolysis can also be used as an analytical tool for studying proteins in the laboratory, and it may also be used in industry, for example in food process ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin-activating Enzyme
Ubiquitin-activating enzymes, also known as E1 enzymes, catalyze the first step in the ubiquitination reaction, which (among other things) can target a protein for degradation via a proteasome. This covalent bond of ubiquitin or ubiquitin-like proteins to targeted proteins is a major mechanism for regulating protein function in eukaryotic organisms. Many processes such as cell division, immune responses and embryonic development are also regulated by post-translational modification by ubiquitin and ubiquitin-like proteins. Ubiquitination (ubiquitylation) Ubiquitin-activating enzyme (E1) starts the ubiquitination process (Figure 1). The E1 enzyme, along with ATP, binds to the ubiquitin protein. The E1 enzyme then passes the ubiquitin protein to a second protein, called ubiquitin carrier or conjugation protein (E2). The E2 protein complexes with a ubiquitin protein ligase (E3). This ubiquitin protein ligase recognizes which protein needs to be tagged and catalyzes the transfer ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin-protein Ligase
A ubiquitin ligase (also called an E3 ubiquitin ligase) is a protein that recruits an E2 ubiquitin-conjugating enzyme that has been loaded with ubiquitin, recognizes a protein substrate, and assists or directly catalyzes the transfer of ubiquitin from the E2 to the protein substrate. In simple and more general terms, the ligase enables movement of ubiquitin from a ubiquitin carrier to another thing (the substrate) by some mechanism. The ubiquitin, once it reaches its destination, ends up being attached by an isopeptide bond to a lysine residue, which is part of the target protein. E3 ligases interact with both the target protein and the E2 enzyme, and so impart substrate specificity to the E2. Commonly, E3s polyubiquitinate their substrate with Lys48-linked chains of ubiquitin, targeting the substrate for destruction by the proteasome. However, many other types of linkages are possible and alter a protein's activity, interactions, or localization. Ubiquitination by E3 ligases ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin-like Protein
Ubiquitin-like proteins (UBLs) are a family of small proteins involved in post-translational modification of other proteins in a cell, usually with a regulatory function. The UBL protein family derives its name from the first member of the class to be discovered, ubiquitin (Ub), best known for its role in regulating protein degradation through covalent modification of other proteins. Following the discovery of ubiquitin, many additional evolutionarily related members of the group were described, involving parallel regulatory processes and similar chemistry. UBLs are involved in a widely varying array of cellular functions including autophagy, protein trafficking, inflammation and immune responses, transcription, DNA repair, RNA splicing, and cellular differentiation. Discovery Ubiquitin itself was first discovered in the 1970s and originally named "ubiquitous immunopoietic polypeptide". Subsequently, other proteins with sequence similarity to ubiquitin were occasionally reported i ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |