|
U1 SnRNA
U1 spliceosomal RNA is the small nuclear RNA (snRNA) component of U1 snRNP (''small nuclear ribonucleoprotein''), an RNA-protein complex that combines with other snRNPs, unmodified pre-mRNA, and various other proteins to assemble a spliceosome, a large RNA-protein molecular complex upon which splicing of pre-mRNA occurs. Splicing, or the removal of introns, is a major aspect of post-transcriptional modification, and takes place only in the nucleus of eukaryotes. Structure and function In humans, the U1 spliceosomal RNA is 164 bases long, forms four stem-loops, and possesses a 5'-trimethylguanosine five-prime cap. Bases 3 to 10 are a conserved sequence that base-pairs with the 5' splice site of introns during RNA splicing, and bases 126 to 133 form the Sm site, around which the Sm ring is assembled. Stem-loop I binds to the U1-70K protein, stem-loop II binds to the U1 A protein, stem-loops III and IV bind to the core RNP domain, a heteroheptameric Sm ring consisting of SmB/B', S ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Secondary Structure
Protein secondary structure is the three dimensional conformational isomerism, form of ''local segments'' of proteins. The two most common Protein structure#Secondary structure, secondary structural elements are alpha helix, alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary structure elements typically spontaneously form as an intermediate before the protein protein folding, folds into its three dimensional protein tertiary structure, tertiary structure. Secondary structure is formally defined by the pattern of hydrogen bonds between the Amine, amino hydrogen and carboxyl oxygen atoms in the peptide backbone chain, backbone. Secondary structure may alternatively be defined based on the regular pattern of backbone Dihedral angle#Dihedral angles of proteins, dihedral angles in a particular region of the Ramachandran plot regardless of whether it has the correct hydrogen bonds. The concept of secondary structure was first introduced by Kaj Ulrik ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Intron
An intron is any nucleotide sequence within a gene that is not expressed or operative in the final RNA product. The word ''intron'' is derived from the term ''intragenic region'', i.e. a region inside a gene."The notion of the cistron .e., gene... must be replaced by that of a transcription unit containing regions which will be lost from the mature messenger – which I suggest we call introns (for intragenic regions) – alternating with regions which will be expressed – exons." (Gilbert 1978) The term ''intron'' refers to both the DNA sequence within a gene and the corresponding RNA sequence in RNA transcripts. The non-intron sequences that become joined by this RNA processing to form the mature RNA are called exons. Introns are found in the genes of most organisms and many viruses and they can be located in both protein-coding genes and genes that function as RNA (noncoding genes). There are four main types of introns: tRNA introns, group I introns, group II introns, and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MicroRNA
MicroRNA (miRNA) are small, single-stranded, non-coding RNA molecules containing 21 to 23 nucleotides. Found in plants, animals and some viruses, miRNAs are involved in RNA silencing and post-transcriptional regulation of gene expression. miRNAs base-pair to complementary sequences in mRNA molecules, then gene silence said mRNA molecules by one or more of the following processes: (1) cleavage of mRNA strand into two pieces, (2) destabilization of mRNA by shortening its poly(A) tail, or (3) translation of mRNA into proteins. This last method of gene silencing is the least efficient of the three, and requires the aid of ribosomes. miRNAs resemble the small interfering RNAs (siRNAs) of the RNA interference (RNAi) pathway, except miRNAs derive from regions of RNA transcripts that fold back on themselves to form short hairpins, whereas siRNAs derive from longer regions of double-stranded RNA. The human genome may encode over 1900 miRNAs, although more recent analysis suggests that ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
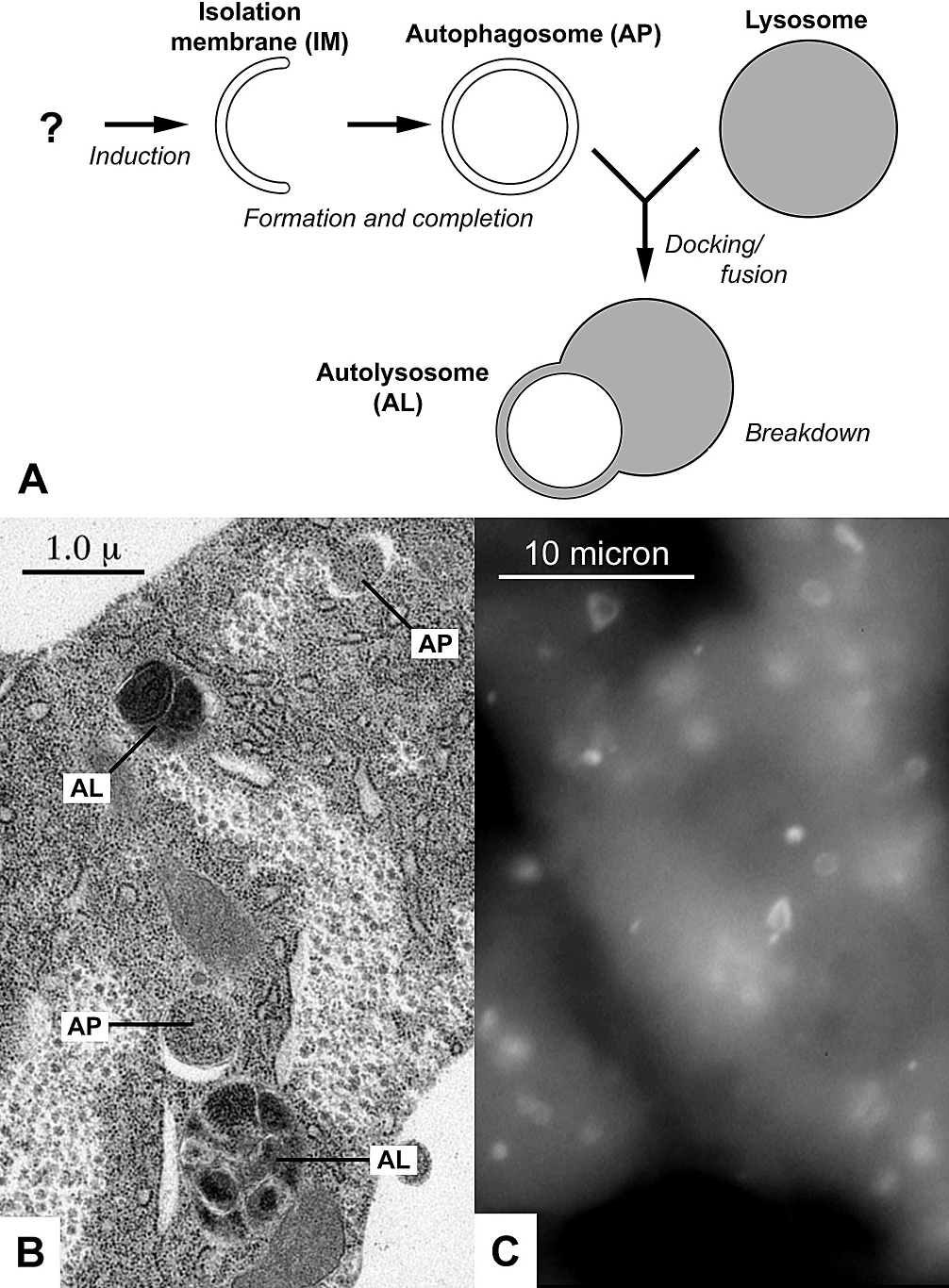
Autophagy
Autophagy (or autophagocytosis; from the Ancient Greek , , meaning "self-devouring" and , , meaning "hollow") is the natural, conserved degradation of the cell that removes unnecessary or dysfunctional components through a lysosome-dependent regulated mechanism. It allows the orderly degradation and recycling of cellular components. Although initially characterized as a primordial degradation pathway induced to protect against starvation, it has become increasingly clear that autophagy also plays a major role in the homeostasis of non-starved cells. Defects in autophagy have been linked to various human diseases, including neurodegeneration and cancer, and interest in modulating autophagy as a potential treatment for these diseases has grown rapidly. Four forms of autophagy have been identified: macroautophagy, microautophagy, chaperone-mediated autophagy (CMA), and crinophagy. In macroautophagy (the most thoroughly researched form of autophagy), cytoplasmic components (like mit ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Morpholino
A Morpholino, also known as a Morpholino oligomer and as a phosphorodiamidate Morpholino oligomer (PMO), is a type of oligomer molecule (colloquially, an oligo) used in molecular biology to modify gene expression. Its molecular structure contains DNA bases attached to a backbone of methylenemorpholine rings linked through phosphorodiamidate groups. Morpholinos block access of other molecules to small (~25 base) specific sequences of the base-pairing surfaces of ribonucleic acid (RNA). Morpholinos are used as research tools for reverse genetics by knocking down gene function. This article discusses only the Morpholino antisense oligomers, which are nucleic acid analogs. The word "Morpholino" can occur in other chemical names, referring to chemicals containing a six-membered morpholine ring. To help avoid confusion with other morpholine-containing molecules, when describing oligos "Morpholino" is often capitalized as a trade name, but this usage is not consistent across scienti ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Antisense
In molecular biology and genetics, the sense of a nucleic acid molecule, particularly of a strand of DNA or RNA, refers to the nature of the roles of the strand and its complement in specifying a sequence of amino acids. Depending on the context, sense may have slightly different meanings. For example, negative-sense strand of DNA is equivalent to the template strand, whereas the positive-sense strand is the non-template strand whose nucleotide sequence is equivalent to the sequence of the mRNA transcript. DNA sense Because of the complementary nature of base-pairing between nucleic acid polymers, a double-stranded DNA molecule will be composed of two strands with sequences that are reverse complements of each other. To help molecular biologists specifically identify each strand individually, the two strands are usually differentiated as the "sense" strand and the "antisense" strand. An individual strand of DNA is referred to as positive-sense (also positive (+) or simply sense) i ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
PolyA
Polyadenylation is the addition of a poly(A) tail to an RNA transcript, typically a messenger RNA (mRNA). The poly(A) tail consists of multiple adenosine monophosphates; in other words, it is a stretch of RNA that has only adenine bases. In eukaryotes, polyadenylation is part of the process that produces mature mRNA for translation. In many bacteria, the poly(A) tail promotes degradation of the mRNA. It, therefore, forms part of the larger process of gene expression. The process of polyadenylation begins as the transcription of a gene terminates. The 3′-most segment of the newly made pre-mRNA is first cleaved off by a set of proteins; these proteins then synthesize the poly(A) tail at the RNA's 3′ end. In some genes these proteins add a poly(A) tail at one of several possible sites. Therefore, polyadenylation can produce more than one transcript from a single gene (alternative polyadenylation), similar to alternative splicing. The poly(A) tail is important for the nuclear ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules within all life-forms on Earth. Nucleotides are obtained in the diet and are also synthesized from common nutrients by the liver. Nucleotides are composed of three subunit molecules: a nucleobase, a five-carbon sugar (ribose or deoxyribose), and a phosphate group consisting of one to three phosphates. The four nucleobases in DNA are guanine, adenine, cytosine and thymine; in RNA, uracil is used in place of thymine. Nucleotides also play a central role in metabolism at a fundamental, cellular level. They provide chemical energy—in the form of the nucleoside triphosphates, adenosine triphosphate (ATP), guanosine triphosphate (GTP), cytidine triphosphate (CTP) and uridine triphosphate (UTP)—throughout the cell for the many cellular func ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Yeast U1 SnRNA
U1 is a small nuclear RNA (snRNA) component of the spliceosome and is involved in pre-mRNA splicing. In the splicing process the 5' end of the U1 snRNA forms complementary base pairing with the 5' splice junction of the intron to be excised, thus defining the 5' donor site of an intron. There are significant differences in sequence and secondary structure between metazoan Animals are multicellular, eukaryotic organisms in the biological kingdom Animalia. With few exceptions, animals Heterotroph, consume organic material, Cellular respiration#Aerobic respiration, breathe oxygen, are Motility, able to move, ca ... and yeast U1 snRNAs, the latter being much longer (568 nucleotides as compared to 164 nucleotides in human). Nevertheless, secondary structure predictions suggest that all U1 snRNAs share a 'common core' consisting of helices I, II, the proximal region of III, and IV. The secondary structure model shows the structure prediction for the larger yeast sequences. R ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Metazoan
Animals are multicellular, eukaryotic organisms in the biological kingdom Animalia. With few exceptions, animals Heterotroph, consume organic material, Cellular respiration#Aerobic respiration, breathe oxygen, are Motility, able to move, can Sexual reproduction, reproduce sexually, and go through an ontogenetic stage in which their body consists of a hollow sphere of Cell (biology), cells, the blastula, during Embryogenesis, embryonic development. Over 1.5 million Extant taxon, living animal species have been Species description, described—of which around 1 million are Insecta, insects—but it has been estimated there are over 7 million animal species in total. Animals range in length from to . They have Ecology, complex interactions with each other and their environments, forming intricate food webs. The scientific study of animals is known as zoology. Most living animal species are in Bilateria, a clade whose members have a Symmetry in biology#Bilateral symmetry, bilat ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
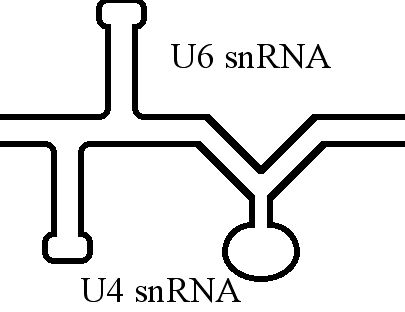
U6 Spliceosomal RNA
U6 snRNA is the non-coding small nuclear RNA (snRNA) component of U6 snRNP (''small nuclear ribonucleoprotein''), an RNA-protein complex that combines with other snRNPs, unmodified pre-mRNA, and various other proteins to assemble a spliceosome, a large RNA-protein molecular complex that catalyzes the excision of introns from pre-mRNA. Splicing, or the removal of introns, is a major aspect of post-transcriptional modification and takes place only in the nucleus of eukaryotes. The RNA sequence of U6 is the most highly conserved across species of all five of the snRNAs involved in the spliceosome, suggesting that the function of the U6 snRNA has remained both crucial and unchanged through evolution. It is common in vertebrate genomes to find many copies of the U6 snRNA gene or U6-derived pseudogenes. This prevalence of "back-ups" of the U6 snRNA gene in vertebrates further implies its evolutionary importance to organism viability. The U6 snRNA gene has been isolated in many organism ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
U2 Spliceosomal RNA
U2 spliceosomal snRNAs are a species of small nuclear RNA ( snRNA) molecules found in the major spliceosomal (Sm) machinery of virtually all eukaryotic organisms. ''In vivo'', U2 snRNA along with its associated polypeptides assemble to produce the U2 small nuclear ribonucleoprotein ( snRNP), an essential component of the major spliceosomal complex. The major spliceosomal-splicing pathway is occasionally referred to as U2 dependent, based on a class of Sm intron—found in mRNA primary transcripts—that are recognized exclusively by the U2 snRNP during early stages of spliceosomal assembly. In addition to U2 dependent intron recognition, U2 snRNA has been theorized to serve a catalytic role in the chemistry of pre-RNA splicing as well. Similar to ribosomal RNAs ( rRNAs), Sm snRNAs must mediate both RNA:RNA and RNA:protein contacts and hence have evolved specialized, highly conserved, primary and secondary structural elements to facilitate these types of interactions. Shortly after ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |